Abstract
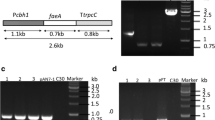
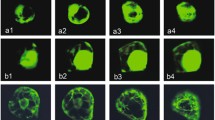
Feruloyl esterase (FAE)-encoding genes AnfaeA and AnfaeB were isolated from Aspergillus niger 0913. For overexpression of the two genes in Trichoderma reesei, constitutive and inductive expression plasmids were constructed based on parental plasmid pAg1-H3. The constructed plasmids contained AnfaeA or AnfaeB gene under the control of glyceraldehyde-3-phosphate dehydrogenase A gene (gpdA) promoter (from A. nidulans) or cellobiohydrolases I (cbh I) gene promoter (from T. reesei), and cbh I terminator from T. reesei. The target plasmids were transferred into T. reesei D-86271 (Rut-C30) by Agrobacterium tumefaciens-mediated transformation (ATMT), respectively. A high level of feruloyl esterase was produced by the recombinant fungal strains under solid-state fermentation, and the cbh I promoter was more efficient than the gpdA promoter in the expression of AnfaeA. The optimum temperatures and pH values were 50 °C and 5.0 for AnFAEA, and 35 °C and 6.0 for AnFAEB. The maximum production levels were 20.69 U/gsd for AnFAEA and 15.08 U/gsd for AnFAEB. The recombinant fungal enzyme systems could release 62.9% (for AnFAEA) and 52.2% (for AnFAEB) of total ferulic acids from de-starched wheat bran, which was higher than the 46.3% releasing efficiency of A. niger 0913. The supplement of xylanase from T. longibrachiatum in the enzymatic hydrolysis led to a small increment of the ferulic acids release.






Similar content being viewed by others
References
Perez-Rodriguez N, Moreira CD, Agrasar AT, Dominguez JM (2016) Feruloyl esterase production by Aspergillus terreus CECT 2808 and subsequent application to enzymatic hydrolysis. Enzyme Microb Tech 91:52–58
Gopalan N, Rodriguez-Duran LV, Saucedo-Castaneda G, Nampoothiri KM (2015) Review on technological and scientific aspects of feruloyl esterases: a versatile enzyme for biorefining of biomass. Bioresource Technol 193:534–544
Kumar N, Pruthi V (2014) Potential applications of ferulic acid from natural sources. Biotechnol Rep 4:86–93
Shirai A, Watanabe T, Matsuki H (2016) Inactivation of foodborne pathogenic and spoilage micro-organisms using ultraviolet—a light in combination with ferulic acid. Lett Appl Microbiol 64:96–102
Mathew S, Abraham TE (2006) Bioconversions of ferulic acid, an hydroxycinnamic acid. Crit Rev Microbiol 32:115–125
Abokitse K, Wu MQ, Bergeron H, Grosse S, Lau PCK (2010) Thermostable feruloyl esterase for the bioproduction of ferulic acid from triticale bran. Appl Microbiol Biot 87:195–203
Mathew S, Abraham TE (2004) Ferulic acid: an antioxidant found naturally in plant cell walls and feruloyl esterases involved in its release and their applications. Crit Rev Biotechnol 24:59–83
Gong YY, Yin X, Zhang HM, Wu MC, Tang CD, Wang JQ, Pang QF (2013) Cloning, expression of a feruloyl esterase from Aspergillus usamii E001 and its applicability in generating ferulic acid from wheat bran. J Ind Microbiol Biot 40:1433–1441
Topakas E, Vafiadi C, Christakopoulos P (2007) Microbial production, characterization and applications of feruloyl esterases. Process Biochem 42:497–509
Uraji M, Arima J, Inoue Y, Harazono K, Hatanaka T (2014) Application of two newly identified and characterized feruloyl esterases from Streptomyces sp. in the enzymatic production of ferulic acid from agricultural biomass. Plos One 9:1–7
Dilokpimol A, Makela MR, Aguilar-Pontes MV, Benoit-Gelber I, Hilden KS, de Vries RP (2016) Diversity of fungal feruloyl esterases: updated phylogenetic classification, properties, and industrial applications. Biotechnol Biofuels 9:231
Crepin VF, Faulds CB, Connerton IF (2004) Functional classification of the microbial feruloyl esterases. Appl Microbiol Biot 63:647–652
Wu H, Li H, Xue Y, Luo G, Gan L, Liu J, Mao L, Long M (2017) High efficiency co-production of ferulic acid and xylooligosaccharides from wheat bran by recombinant xylanase and feruloyl esterase. Biochem Eng J 120:41–48
Moukouli M, Topakas E, Christakopoulos P (2008) Cloning, characterization and functional expression of an alkalitolerant type C feruloyl esterase from Fusarium oxysporum. Appl Microbiol Biot 79:245–254
Wong DWS, Chan VJ, Liao H, Zidwick MJ (2013) Cloning of a novel feruloyl esterase gene from rumen microbial metagenome and enzyme characterization in synergism with endoxylanases. J Ind Microbiol Biot 40:287–295
Martinez D, Berka RM, Henrissat B, Saloheimo M, Arvas M, Baker SE, Chapman J, Chertkov O, Coutinho PM, Cullen D et al (2008) Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nat Biotechnol 26:1193–1193
Bischof RH, Ramoni J, Seiboth B (2016) Cellulases and beyond: the first 70 years of the enzyme producer Trichoderma reesei. Microb Cell Fact 15:106
Peterson R, Nevalainen H (2012) Trichoderma reesei RUT-C30–thirty years of strain improvement. Microbiology 158:58–68
Ramoni J, Marchetti-Deschmann M, Seidl-Seiboth V, Seiboth B (2017) Trichoderma reesei xylanase 5 is defective in the reference strain QM6a but functional alleles are present in other wild-type strains. Appl Microbiol Biotechnol 101:4139–4149
Gong WL, Zhang HQ, Liu SJ, Zhang LL, Gao PJ, Chen GJ, Wang LS (2015) Comparative secretome analysis of Aspergillus niger, Trichoderma reesei, and Penicillium oxalicum during solid-state fermentation. Appl Biochem Biotech 177:1252–1271
Mandels M, Andreotti RE (1978) Problems and challenges in the cellulose to cellulase fermentation. Process Biochem 13:6–13
Long L, Ding D, Han Z, Zhao H, Lin Q, Ding S (2016) Thermotolerant hemicellulolytic and cellulolytic enzymes from Eupenicillium parvum 4–14 display high efficiency upon release of ferulic acid from wheat bran. J Appl Microbiol 121:422–434
Khang CH, Park SY, Rho HS, Lee YH, Kang S (2006) Filamentous fungi (Magnaporthe grisea and Fusarium oxysporum). Methods Mol Biol 344:403–420. Agrobacterium protocols, 2/e, volume 2. Edited by Wang K, Humana, Totowa
Ullán RV, Godio RP, Teijeira F, Vaca I, García-Estrada C, Feltrer R, Kosalkova K, Martín JF (2008) RNA-silencing in Penicillium chrysogenum and Acremonium chrysogenum: validation studies using beta-lactam genes expression. J Microbiol Meth 75:209–218
Mullins ED, Chen X, Romaine P, Raina R, Geiser DM, Kang S (2001) Agrobacterium-mediated transformation of Fusarium oxysporum: an efficient tool for insertional mutagenesis and gene transfer. Phytopathology 91:173–180
Long LK, Yang J, An Y, Liu G (2012) Disruption of a glutathione reductase encoding gene in Acremonium chrysogenum leads to reduction of its growth, cephalosporin production and antioxidative ability which is recovered by exogenous methionine. Fungal Genet Biol 49:114–122
de Cassia Pereira J, Marques NP, Rodrigues A, de Oliveira TB, Boscolo M, da Silva R, Gomes E, Martins DAB (2015) Thermophilic fungi as new sources for production of cellulases and xylanases with potential use in sugarcane bagasse saccharification. J Appl Microbiol 118:928–939
Turner BL (2010) Variation in pH optima of hydrolytic enzyme activities in tropical rain forest soils. Appl Environ Microbiol 76:6485–6493
Levasseur A, Navarro D, Punt PJ, Belaich JP, Asther M, Record E (2005) Construction of engineered bifunctional enzymes and their overproduction in Aspergillus niger for improved enzymatic tools to degrade agricultural by-products. Appl Environ Microb 71:8132–8140
de Vries RP, VanKuyk PA, Kester HCM, Visser J (2002) The Aspergillus niger faeB gene encodes a second feruloyl esterase involved in pectin and xylan degradation and is specifically induced in the presence of aromatic compounds. Biochem J 363:377–386
Faulds CB (2010) What can feruloyl esterases do for us? Phytochem Rev 9:121–132
Levasseur A, Saloheimo M, Navarro D, Andberg M, Monot F, Nakari-Setala T, Asther M, Record E (2006) Production of a chimeric enzyme tool associating the Trichoderma reesei swollenin with the Aspergillus niger feruloyl esterase A for release of ferulic acid. Appl Microbiol Biot 73:872–880
Levasseur A, Benoit I, Asther M, Asther M, Record E (2004) Homologous expression of the feruloyl esterase B gene from Aspergillus niger and characterization of the recombinant enzyme. Protein Expres Purif 37:126–133
Kroon PA, Faulds CB, Williamson G (1996) Purification and characterization of a novel esterase induced by growth of Aspergillus niger on sugar-beet pulp. Biotechnol Appl Biochem 23:255–262
Faulds CB, Williamson G (1995) Release of ferulic acid from wheat bran by a ferulic acid esterase (FAE-III) from Aspergillus niger. Appl Microbiol Biotechnol 43:1082–1087
de Vries RP, Michelsen B, Poulsen CH, Kroon PA, van den Heuvel RH, Faulds CB, Williamson G, van den Hombergh JP, Visser J (1997) The faeA genes from Aspergillus niger and Aspergillus tubingensis encode ferulic acid esterases involved in degradation of complex cell wall polysaccharides. Appl Environ Microbiol 63:4638–4644
Bartolomé B, Faulds CB, Tuohy M, Hazlewood GP, Gilbert HJ, Williamson G (1995) Influence of different xylanases on the activity of ferulic acid esterase on wheat bran. Biotechnol Appl Biochem 22:65–73
de Vries RP, Visser J (1999) Regulation of the feruloyl esterase (faeA) gene from Aspergillus niger. Appl Environ Microbiol 65:5500–5503
Acknowledgements
We are grateful to Prof. Gang Liu (Institute of Microbiology, CAS) for providing the plasmid pAg1-H3 and Agrobacterium tumefaciens AGL-1. This work was supported by grants from a 948 Research Project (No. 2013-4-16) from the State Forestry Administration of China, the Natural Science Fund for Provincial Colleges and University of Jiangsu Province, China (15KJB220003), the Research Fund for the Advanced Talents, Nanjing Forestry University (GXL201311) and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Long, L., Zhao, H., Ding, D. et al. Heterologous expression of two Aspergillus niger feruloyl esterases in Trichoderma reesei for the production of ferulic acid from wheat bran. Bioprocess Biosyst Eng 41, 593–601 (2018). https://doi.org/10.1007/s00449-018-1894-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-018-1894-3