Abstract
Pneumonia is counted among the leading causes of death worldwide. Viruses, bacteria and pathogen-related molecules interact with cells present in the human alveolus by numerous, yet poorly understood ways. Traditional cell culture models little reflect the cellular composition, matrix complexity and three-dimensional architecture of the human lung. Integrative animal models suffer from species differences, which are of particular importance for the investigation of zoonotic lung diseases. The use of cultured ex vivo infected human lung tissue may overcome some of these limitations and complement traditional models. The present review gives an overview of common bacterial lung infections, such as pneumococcal infection and of widely neglected pathogens modeled in ex vivo infected lung tissue. The role of ex vivo infected lung tissue for the investigation of emerging viral zoonosis including influenza A virus and Middle East respiratory syndrome coronavirus is discussed. Finally, further directions for the elaboration of such models are revealed. Overall, the introduced models represent meaningful and robust methods to investigate principles of pathogen-host interaction in original human lung tissue.
Similar content being viewed by others
Introduction
Pneumonia is counted among the group of widespread diseases and lower respiratory tract infections belong to the five most common causes of death worldwide (Global Burden of Disease Study 2015). The high burden of disease consequently entails a great economic burden to the general public (Welte et al. 2012). Pneumonia is a severe inflammatory condition of the lung affecting primarily the peripheral alveolar compartment. Clinically, patients suffer from a productive or dry cough, chest pain, fever and compromised respiration. A simple pneumonia may progress to a life-threating condition with subsequent respiratory failure and systemic inflammation (Bauer et al. 2006). Usually, infection with viruses or bacteria and less commonly with other microorganisms causes pneumonia. Despite applicable vaccination strategies against pathogens commonly causing pneumonia, such as Streptococcus pneumoniae (Mehr and Wood 2012; Scott et al. 2012) and influenza A virus (IAV; Ortiz et al. 2016), being available, these pathogens still cause tremendous morbidity and mortality worldwide. Furthermore, antibiotic resistance is an emerging problem in infectious diseases per se (Brown and Wright 2016). Risk factors for pneumonia comprise further lung diseases such as cystic fibrosis, chronic obstructive pulmonary disease (COPD) and asthma and include other problematic conditions such as diabetes, heart failure, a history of smoking and alcohol abuse (Torres et al. 2015). In addition, children younger than 5 years of age (Walker et al. 2013) and people older than 65 years (Torres et al. 2013) are at a higher risk for developing pneumonia.
Why is the investigation of host-pathogen interaction, especially in human lung tissue, the key for developing new intervention strategies in pneumonia? Classical cell culture models are useful for the mechanistic and functional analysis of the one-to-one interaction between cell and microbe. In an expanded version, a second implemented cell type could, for example, reflect the simple influence of additional host factors. However, these basic in vitro systems neither reflect tissue diversity at a cellular level, nor recapitulate the typical alveolar cell-cell interaction in the natural matrix (e.g., alveolar epithelial cell [AEC]-I interaction with AEC-II and the capillary endothelium) of the three-dimensional architecture. Therefore, if researchers would like to investigate the cellular tropism of a virus causing pneumonia in a realistic model, this is only possible if all cell types forming the potential habitat are present in the dish at once. Frequently passaged primary immortalized cells (such as bronchial epithelial BEAS-2B; Ke et al. 1988) or tumor-derived cells (such as alveolar A549 cells; Smith 1977), by using proteases such as trypsin for cell splitting, may lack central biological characteristics important for host-pathogen interactions. These include, for example, the capability to form tight junctions (Rothen-Rutishauser et al. 2008) or to express receptors involved in pathogen-binding or recognition. In addition, the use of wild-type encapsulated bacteria is limited in most cell cultures because of the cell toxic effects of the capsule (N’Guessan et al. 2005; Schmeck et al. 2004) motivating researchers to use unencapsulated laboratory-derived strains. Since, for the pathogenicity of, for example, the over 90 serotypes of S. pneumoniae (the major cause of pneumonia; Bauer et al. 2006; Bogaert et al. 2004; Drijkoningen and Rohde 2014; Jefferson et al. 2006), capsule-related effects are of enormous significance for virulence and tissue invasion (Geno et al. 2015), the use of original encapsulated patient-derived strains is important for the realistic modeling of pathogen-host interaction.
Animals such as mice represent integrative models and modern genetic animal manipulation allows the sophisticated analysis of host-pathogen interaction, including that of the lung (Baron et al. 2012; Hraiech et al. 2015; Thangavel and Bouvier 2014). Nevertheless, significant differences in anatomy, innate and adaptive immunity have long been known to exist between humans and, in particular, small rodents (Mestas and Hughes 2004). For many results obtained in such models, their translation to humans remains unclear (Mak et al. 2014). With respect to the investigation of infectious diseases in rodents, including pneumonia, a major problem is the host specificity of most pathogens (Bean et al. 2013; Bouvier 2015; Gretebeck and Subbarao 2015; Sutton and Subbarao 2015; Ware 2008; Wolfe et al. 2007). For example, the human nasopharynx seems to represent the natural reservoir for S. pneumoniae (Bogaert et al. 2004; Kadioglu et al. 2008) and only rare observations of this pathogen in wildlife species without contact to humans are documented (Chi et al. 2007). Some animal models, such as ferrets and guinea pigs, are naturally susceptible to infection by human influenza A strains (IAV); others, such as mice, require adaptation of the virus (Bouvier 2015). In particular, the majority of IAV research in mice employs either BALB/C or C57BL/6 strains in conjunction with the laboratory adapted A/Puerto Rico/8/1934 (H1N1) (PR8) or A/WSN/1933 (H1N1) (WSN). Such adaptation of pathogens to animal model species involves serial passaging to increase virulence, a procedure that inevitably alters pathogen behavior. Emerging viral lung diseases, such as the severe acute respiratory syndrome coronavirus (SARS-CoV; Peiris et al. 2004; Poon et al. 2004), Middle East respiratory syndrome coronavirus (MERS-CoV; Fehr et al. 2016; Mohd et al. 2016; Zumla et al. 2015) and new IAV strains infecting humans (e.g., H5N1, H1N1, H7N9; Lai et al. 2016; Novel Swine-Origin Influenza et al. 2009; Peiris et al. 2009; Zhu et al. 2016) represent classical zoonotic diseases, indicating that the species used for research is of great importance. Noteworthy, although, for example, IAV infects a broad variety of species, it does not infect rodents such as mice in nature. For some diseases, such as MERS-CoV, the available animal models do not represent the human clinical disease (Gretebeck and Subbarao 2015; Sutton and Subbarao 2015) and the lack of receptors for viral binding (Raj et al. 2013) has forced researchers to create complex transgenic humanized models (Gretebeck and Subbarao 2015; Sutton and Subbarao 2015).
Overall, in order to investigate the biology and to estimate the virulence potential of such pathogens, we need to complement in vitro cell culture studies and animal studies with suitable human-derived models. In the following, the use of ex vivo cultured and infected human lungs for the investigation of bacterial and viral infections will be discussed and further directions for model development will be revealed.
Ex vivo bacterial infection of human lung tissue
Streptococcus pneumoniae
Pneumococci frequently colonize the human nasopharynx (Bogaert et al. 2004; Scott et al. 2012), which seems to be indeed the primary natural habitat of this important human pathogen. S. pneumoniae infections of animals such as race horses, rhesus monkeys, or chimpanzees occur mostly in animals held in human captivity and are suspected to be attributable to human-animal transmission (Chi et al. 2007). In all studies searching for the causative agent of pneumonia, pneumococci are the most frequently isolated pathogen, in both out-patient and in-inpatient settings (including severe pneumonia treated at intensive care units; Drijkoningen and Rohde 2014). S. pneumoniae strains differ considerably in their capacity to cause a disease in humans in general and some strains may primarily cause invasive disease, whereas others predominately induce otitis media (Jefferson et al. 2006; Mehr and Wood 2012). Since pneumococci are a true human-specific pathogen of high clinical importance with a high diversity of diseases caused by the various clinical isolates, we urgently need to study the biology of this pathogen in human tissue.
The ex vivo infection of human lungs with pneumococci induces the expression of immunomodulatory molecules such as tumor necrosis factor α (TNFα), interleukin-1β (IL-1β), granulocyte-macrophage colony-stimulating factor, platelet-derived growth factor (PDGF), IL-6, IL-8, IL-10, IL-15, IL-17, or prostaglandin-E2 (PGE2; Fatykhova et al. 2015; Szymanski et al. 2012; Xu et al. 2008). Experiments performed with the clodronate-induced depletion of alveolar lung macrophages (AM) indicated that these cells might, in particular, contribute to the induction of IL-6 and TNFα (Xu et al. 2008). Fatykhova et al. (2015) involved the use of various clinical isolates to investigate their capacity to induce the potent pro-inflammatory cytokine IL-1β. These clinical strains differed in the expression of the cholesterol-dependent cytolysin pneumolysin (PLY), a major virulence factor of pneumococci (Kadioglu et al. 2008): Whereas serotypes 2, 3, 6B and 9N pneumococci express fully hemolytic PLY, serotype 1 and 8 strains express non-hemolytic PLY (Fatykhova et al. 2015). The authors demonstrated that only strains expressing lytic PLY induce IL-1β in a NLRP3-inflammasome-dependent manner. Experiments with purified allele 1 PLY (capable of causing pores) and allele 5 PLY (not causing pores) verified that pore formation is a pre-requisite for PLY-related IL1β induction in human lung tissue. In addition to the mechanistic information about IL-1β regulation in human lungs, this study highlighted the suitability of the model to analyze the biology of patient-derived bacterial isolates and showed the meaningfulness of the results obtained.
Cyclooxygenases (COX) produce fatty acid mediators, including prostaglandins such as PGE2, which play an important role in the regulation of lung immunity (Claar et al. 2015; Zhou et al. 2016). In human lungs, pneumococci induce the strong up-regulation of the inducible form of COX, namely COX-2, in particular, in AM, AEC-II (but not AEC-I) and the vascular endothelium. Notably, same results have been seen in the tissue of patients suffering from pneumonia (Szymanski et al. 2012). Inhibition of p38 MAPK (mitogen-activated protein kinase) or ERK1/2 (extracellular signal-regulated kinase 1/2) blocked both the induction of COX-2 and the release of PGE2. In addition to PGE2, the authors showed the release of 6-keto PGF1α and thromboxane B2 into the infected tissue. Tissue expresses predominately the E prostanoid receptor 4 (EP4) and EP4 stimulation results in an increased cAMP production in lung tissue. Such PGE2 production by lung cells may contribute to the control of inflammatory mediator production, such as that of IL-1β (Mortimer et al. 2016) in pneumococcal pneumonia.
Furthermore, Xu et al. (2008) noted the increased expression of Toll-like receptor 2 (TLR2) and TLR4 mRNA in pneumococci-infected tissue; however, neither the way that this translates into protein expression nor the effected cells are known. In accordance, the means by which the observed activation of signaling pathways such as MAPKs (Szymanski et al. 2012; Xu et al. 2008) is made cell-specific is unknown.
In addition to PLY, pneumococci liberate significant amounts of hydrogen peroxide and thus we can reasonably suggest that this causes oxidative stress to the lungs (Kadioglu et al. 2008). However, although pneumococci cause oxidative stress in human lungs, as evidenced by a decreased ratio of glutathione to glutathione disulfide in infected tissue (Zahlten et al. 2015b), this seems not to depend on oxygen radicals. Unexpectedly, a pneumococcal autolysin A (LytA)-dependent process turns out to induce oxidant stress. LytA is the major autolysin of pneumococci (Lopez and Garcia 2004) and causes the release of intrabacterial components such as PLY and bacterial DNA. Since the pneumococci-related expression of the immunomodulatory transcription factor Krueppel-like factor (KLF) 4 (McConnell and Yang 2010) in human lungs seems also to depend on LytA-related pneumococcal autolysis (Zahlten et al. 2015b) and as KLFs have an impact on lung cell activation in pneumococcal infection (Zahlten et al. 2010, 2013, 2015a, b), further investigation of LytA-related activation of lung tissue is highly recommended.
The above-mentioned studies indicate strong pro-inflammatory mediator release in pneumococci-infected lung tissue (Fatykhova et al. 2015; Szymanski et al. 2012; Xu et al. 2008) and massive inflammation, in particular during severe pneumonia, is suggested to foster the progression of the disease to acute respiratory failure, sepsis and multiorgan dysfunction (Bauer et al. 2006). Quinolones, such as moxifloxacin, have been hypothesized to exert anti-inflammatory (beneficial) effects, in addition to their well-established antimicrobial properties (Dalhoff and Shalit 2003). However, when investigating the effect of moxifloxacin on pneumococci- or TNF-α-stimulated IL-6 and IL-8 release in human lung tissue, Müller-Redetzky et al. (2015) found that only TNF-α-related IL-6 release was reduced by moxifloxacin. Accompanying investigations in a pneumococcal mouse pneumonia model showed similar results and, thus, did not support the hypothesis that moxifloxacin exhibits potent anti-inflammatory potency in pneumococcal pneumonia.
Bacillus anthracis
Although Bacillus anthracis does not cause pneumonia, the lung is the entry site for B. anthracis in inhalation anthrax, which is the most deadly form of the disease. It seems that inhaled spores escape from the alveolus into regional lymph nodes. Therein, spores may germinate and induce disease after having reached the circulatory system (Moayeri et al. 2015). Important virulence factors of B. anthracis include two toxins: both lethal toxin (LT) and edema toxin (ET) share the protective antigen (PA) as a common receptor-binding component. PA allows the transport of the catalytic components LF and EF into the cytosol of target cells (Friebe et al. 2016; Moayeri et al. 2015). Chakrabarty et al. (2007) tested B. anthracis spore-related inflammatory tissue activation by utilizing spores prepared from the B. anthracis Sterne strain 7702(pX01+, pX02-). A significant increase in IL-6, TNFα, IL-8, monocyte chemoattractant protein-1 (MCP-1) and macrophage inflammatory protein 1 (MIP)-1 α/β was noted in the lung tissue supernatants; however, although a tremendous up-regulation of IL-1β mRNA was documented, the authors noted no increase in IL-1β protein. Nevertheless, supernatants of spore-exposed lung tissue stimulated neutrophil and monocyte chemotaxis. B. anthracis spores caused strong activation of the MAPKs ERK, JNK and p38 and chemical inhibition of the kinases reduced chemo- and cytokine liberation. Finally, immunohistochemistry revealed the presence of IL-6 and IL-8 in epithelial cells and AM. Overall, the study (Chakrabarty et al. 2007) revealed that B. anthracis spores initiated a prominent pro-inflammatory response.
An important unresolved question is: how do B. anthracis spores escape from the alveolar airspace into the systemic circulation (Friebe et al. 2016; Moayeri et al. 2015)? In principle, transport might occur via host cells used as Trojan horses (AM, dendritic cells [DC], or a hitherto unidentified “carrier” cell) or spores may cross the alveolar epithelial wall without the help of migratory cells. Finally, a sequence of spore clustering, germination and production of B. anthracis virulence toxins has been proposed to cause epithelial damage that allows free spore passage (Friebe et al. 2016; Goossens and Tournier 2015; Moayeri et al. 2015), the so-called “jailbreak” model (Weiner and Glomski 2012). By using the above-mentioned model (Chakrabarty et al. 2007), the same group (Booth et al. 2016) aimed to identify the role of carrier cells and of B. anthracis toxins in this process. Around 5 % of spores were internalized in APC and around 13 % in AEC, whereas around 80 % of the spores were free after 2 h post-infection and these numbers did not change over time. Importantly, the clustering of spores occurred only in infected cells. The clear identification of the AEC cells that take up spores (AEC I, AEC II), by means of appropriate imaging techniques, would be of interest. Interestingly, the addition of PA or LT neither significantly influences spore uptake nor causes any cytotoxicity (as spore treatment alone also does not). Overall, the data suggest that B. anthracis spores migrate through the lung soon after exposure. The primary initial phase of spore movement from the alveolar space across the alveolar epithelial barrier may not essentially require a cellular carrier. However, the above study showed no translocation of spores into lung blood vessels (e.g., alveolar capillaries) and further investigations are required to demonstrate that the presence of spores in alveolar cells finally results in spore movement into the circulation.
Mycobacterium tuberculosis
Mycobacterial lung infections are still a major cause of morbidity and mortality worldwide. In 2012, around 9.0 million people developed tuberculosis (TB) and 1.5 million people died of this chronic infectious lung disease (Zumla et al. 2013). Although the innate immune system may clear early infection of M. tuberculosis in a significant number of cases (Khan et al. 2016; Morrison et al. 2008), little is known about the initial interaction of this important pathogen with human lung tissue. Ganbat et al. (2016) have now started to explore such interactions by establishing an ex vivo human lung infection model with various mycobacterial strains (M. tuberculosis, M. abscessus, M. avium). The authors note that AM, monocytes, neutrophils and AEC-II cells are infected by the mycobacterial strains. Interestingly, AEC-II seem to be infected in a significantly higher frequency by M. tuberculosis than by M. abscessus or M. avium. The presented results indicate the occurrence of cell death in all infected cell types but this also differs among strains. Although the study is limited by the relatively short time period post-infection (16 h post-infection), this model in principle will allow the detailed analysis of the initial mycobacterium-alveolar interaction, including the activation of inflammation-regulating mediators by, for example, clinical isolates. Thus, the model adds an important step forward for the investigation of these important events in early mycobacterial infection.
Nontypeable Haemophilus influenzae
Nontypeable Haemophilus influenza (NTHI) infections, in particular newly acquired strains, may trigger infection-related exacerbation in COPD and recurrent infections may induce overall disease progression (Duell et al. 2016). On using the NTHI strain Rd KW20 isolated from a COPD patient suffering from invasive pneumonia, Wagner et al. (2015) noticed strong IL-8 induction in ex vivo infected human lungs; this was reduced in tissue pre-treated with the anti-inflammatory drug budenoside. Interestingly, the steroid reduced not only the induction of IL-8; it furthermore reduced the presence of intracellular bacteria in the tissue by a hitherto unexplored mechanism. How the C-type lectin receptor Dectin-1, which has recently been found to be expressed apically on human bronchial and alveolar epithelium (Heyl et al. 2014), participates in the pro-inflammatory activation of human lung tissue, in addition to the “classic” innate immune receptors such as TLRs, needs to be determined. In a subsequent study, the group of Drömann (Dromann et al. 2010) used two clinical isolates: one isolated from a COPD patient with invasive disease and one from a patient without COPD and invasion. Both strains infected AM and lung epithelial cells; however, AEC subtypes were not identified in this study. Notably, the authors observed a moderate induction of TGF-β but a strong upregulation of the TGF-β-pseudoreceptor BMP and activin membrane-bound inhibitor (BAMBI) on both the alveolar wall epithelium and AM. This was accompanied by p38 MAPK-triggered induction of the pro-inflammatory mediators IL-8 and TNF-α. TGF-β is an important mediator of inflammatory and remodeling events in the lung and elevated levels of BAMBI protein in plasma were observed in COPD patients (Zhang et al. 2016). In these patients, increased BAMBI expression on human CD4+ T cell membranes was noted and the enhanced plasma BAMBI levels in COPD positively correlated with increased plasma TGF-β1 levels and the Th17/Treg ratio indicating that an impaired TGF-β/BAMBI pathway can promote inflammation in COPD (Zhang et al. 2016). Furthermore, studies in the lungs of idiopathic pulmonary fibrosis patients and the mouse model of bleomycin-related lung fibrosis (Murphy et al. 2016) have demonstrated increased levels of BAMBI (and related BMP accessory proteins noggin and FSTL1) in the lungs suggesting a role of these molecules in inflammatory and fibrotic lung injury. Since other studies have suggested a role of TGF-β and related molecules in acute lung infection, such as IAV-related pneumonia (Furuya et al. 2015; Woods et al. 2015) and IAV-bacterial coinfection (Li et al. 2015), a more detailed analysis of the function of these molecules in human lungs is highly desirable.
Legionella pneumophila
Legionella pneumophila (L. pneumophila) is the causative agent of Legionnaires’ disease. The inhalation of contaminated water droplets causes disease outbreaks of public health significance (Phin et al. 2014). The bacterium interacts with the human host by the release of extracellular proinflammatory outer membrane vesicles (OMV) and uses the Dot/Icm type IVB translocation system to inject over 300 effector proteins into the infected cell (Ensminger 2016). It creates a replicative niche by avoiding fusion of phagosomes with the lysosome, interacts with endoplasmic reticulum–Golgi traffic (Prashar and Terebiznik 2015) and induces the massive pro-inflammatory activation of human lung epithelium in vitro (Schmeck et al. 2007, 2008). Jäger et al. (2014) started their investigation of Legionella infection in human lung tissue by using wild-type strain Corby and a DotA-negative mutant (defect for the Dot/Icm type IVB translocation system). Furthermore, they explored the role of OMV, suspected of inducing massive lung cell activation (Galka et al. 2008). They showed that extracellular adhesion to the alveolar epithelial barrier took place before Legionella entered macrophages. Whereas wild-type bacteria multiplied more than 10-fold within 48 h, DotA-negative bacteria could not replicate within the tissue. Of note, infection caused damage to both the infected AM and the alveolar wall. Bacteria deficient for DotA induced less damage than wild-type bacteria highlighting the importance of this virulence system. The AM surface and cytoplasm were decorated by OMV suggesting that these cells are particularly targeted by pro-inflammatory Legionella vesicles. OMV induced damage to the lung comparable with that caused by wild-type bacteria, shedding light on the possible important role of OMV in Legionella pathogenesis. By using transcriptome analysis, the authors showed the differentiated expression of more than 2400 genes in Legionella-infected human lung tissue, including genes related to extracellular proteins, components of the immune response and lipoprotein transport proteins. The pathophysiological role of, for example, the observed down-regulation of the protein content of immunoregulatory uteroglobin (a member of the secretoglobin superfamily; Mukherjee et al. 2007) and the downregulation of MARCO, a class A scavenger receptor, which seems to be involved in, for example, the pathogenesis of the pneumococcus (Dorrington et al. 2013), remains unknown. In addition, the further characterization of the way that vacuole formation (Naujoks et al. 2016) and the activation of innate immune receptors (Cunha and Zamboni 2014) take place in original human lung tissue would be of great interest.
Coxiella burnetii
The obligate intracellular pathogen Coxiella burnetii causes Q fever, a disease starting with flu-like symptoms, which, in cases of prolonged infections, may proceed to severe endocarditis. Similar to Legionnaires’ disease, the public health system noticed Q fever mostly in the form of localized outbreaks resulting from the inhalation of contaminated aerosols from farm and domestic animals (in particular sheep and goats; Cilloniz et al. 2016). Coxiella replicates within an acidic lysosome-like parasitophorous vacuole (PV), mostly in macrophages and uses a Dot/Icm type IV secretion-system-based molecule delivery into host cells to hijack host cell signaling cascades, thereby creating its replicate niche (Moffatt et al. 2015). Graham et al. (2016) compared the behavior of avirulent C. burnetii NMII (RSA439, clone 4, a frequently used laboratory strain) with virulent C. burnetii in human precision-cut lung slices. Although single bacteria were detected in the alveolar epithelium, bacteria replicated sufficiently only in AM. Of note, only live avirulent bacteria induced the liberation of IL-1β and IL-18, whereas virulent bacteria did not. Subsequently, on using human AM, the authors found that a human-specific noncanonical inflammasome dependent on caspase-4/-5 might induce IL-1β release without the induction of pyroptosis. Since caspase-4 and caspase-5 are human-specific proteins, caspase-5 is undetectable in the THP-1 human macrophage-like cell line model and mouse macrophages do not respond with IL-1β liberation to C. burnettii infection (Cunha et al. 2015); this discovery was essentially based on the use of original human tissue. Notably, since numerous studies involving the use of avirulent and virulent C. burnettii indicated widely identical intracellular behavior of the bacteria (Moffatt et al. 2015), further investigations in primary human tissue (and cells) are needed in order to discover the way that virulent C. burnetii suppress IL-1β production, thereby inhibiting one of the most powerful pro-inflammatory mediators in the host.
Ex vivo viral infection of human lung tissue
Influenza virus
Because of the enormous clinical importance and continuous emergence of new influenza A strains (IAV; Lai et al. 2016; Trombetta et al. 2015), various workers have investigated IAV infection of human lung tissue. Research has focused, in most reports, on viral replication, tissue tropism and tissue activation in the sense of cytokine and chemokine liberation. Knowledge of the level of efficiency of an emerging IAV replicate in human lung tissue is of enormous importance to estimate its potential to spread in the community and to gauge the potential of the virus to cause harm to the lungs. For example, the avian IAV H5N1 has caused typically severe human infections with a high fatality rate over many years (Lai et al. 2016). By using human lung tissue ex vivo infected with IAV H5N1, several groups have demonstrated strong viral replication (R.W. Chan et al. 2009; Hocke et al. 2013a; Nicholls et al. 2007; Weinheimer et al. 2012) comparable with or even higher than that observed with classic pandemic strains such as IAV H3N2 (Weinheimer et al. 2012). In 2009, the first pandemic of the 21st century started with a novel swine-originated IAV H1N1 virus (Novel Swine-Origin Influenza et al. 2009). In most cases, this virus caused a disease of moderate severity as observed in most seasonal influenza (Peiris et al. 2009). In accordance, compared with highly virulent IAV strains such as H5N1 or classic pandemic strains such as IAV H3N2, this virus showed intermediate replication in ex vivo infected human lung tissue (M.C. Chan et al. 2010; Weinheimer et al. 2012; Wu et al. 2010b, 2012; Zhang et al. 2010) thus reflecting the moderate disease caused in humans. A new IAV H7N9 virus resulting from sequential reassortments in ducks and chickens has been shown to infect human beings since 2013. Originated from eastern China, the virus has caused repeated waves of outbreaks and has thus raised concerns of a pandemic threat (Zhou et al. 2016). Two reports have revealed that IAV H7N9 efficiently replicates in human lung tissue (M.C. Chan et al. 2013; Knepper et al. 2013). Noticeably, whereas virus isolated from a fatal human infection (A/Anhui/1/2013 (H7N9)) replicated comparably with seasonal IAV, classic avian H7 subtype viruses propagated poorly indicating that this new H7N9 virus is well adapted to replicate in the human host (Knepper et al. 2013). In agreement, several studies have demonstrated that IAV strains not adapted to humans (such as classic swine or avian strains) do not propagate efficiently in human material (M.C. Chan et al. 2010, 2013; Knepper et al. 2013; Weinheimer et al. 2012; Wu et al. 2010b; Zhang et al. 2010). Overall, the ex vivo models used in these studies seem to robustly reflect the capacity to infect and propagate in humans.
Infection of a host cell with IAV results in numerous alterations in cell function, ranging from the release of immune-regulating mediators to changes in sodium pump activity and finally the killing of IAV-infected cells (Herold et al. 2012; Short et al. 2014). Thus, the identification of the primary target cells of IAV in the human lung is of primary importance if we are to estimate the possible effects of cell damage for organ function and to investigate possible cell-based interventions. Irrespective of the virus strain used in the ex vivo infection models, most reports identify AEC II cells as the primary replicative niche in the peripheral human lung (Fig. 1; M.C. Chan et al. 2010, 2013; Hocke et al. 2013b; Knepper et al. 2013; Weinheimer et al. 2012; Zhang et al. 2010). Patients with lung fibrosis typically show AEC II hyperplasia (Fernandez and Eickelberg 2012). Hocke et al. (2013b) demonstrated massive infection of AEC II cells of fibrotic lungs and Fujino et al. (2013) noticed increased IAV replication in AEC II cells derived from patients with pulmonary fibrosis thus indicating that the AEC II cell is the primary target cell of IAV in the human lung. In contrast, the presence of viral antigen is detected much less frequently in human alveolar macrophages (M.C. Chan et al. 2010, 2013; Weinheimer et al. 2012) but nevertheless these cells are of importance for the immune response to IAV (Halstead and Chroneos 2015). Although (1) IAV H1N1 particles have been identified in endothelial cells of fatal cases (Ru et al. 2011), (2) in vitro experiments indicate IAV propagation in endothelial cells (Wang et al. 2015) and (3) the endothelium might significantly contribute to the course of the disease (Teijaro et al. 2011), IAV antigen has not been detected in the ex vivo models discussed here.
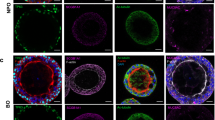
Influenza A virus (IAV) targets AEC-II in ex vivo infected human lung tissue. Cross-sections from infected lung explants were stained for IAV antigen (green), for prosurfactant protein C (blue) to detect alveolar epithelial cell II (AEC-II, T II; white arrowheads infected cells [cyan], open arrowheads uninfected cells [blue]) and for CD68 (red) to detect alveolar lung macrophages (asterisk infected cell [yellow]). Nuclear staining with 4,6-diamidino-2-phenylindole is shown in dark orange and lung structure is visualized with differential interference contrast. The stains were visualized by confocal microscopy and tissue autofluorescence was separated from specific fluorescence by spectral unmixing. From Weinheimer et al. (2012). Reproduction by permission of Oxford University Press
The tissue responded to IAV infection with the release of multiple immunomodulatory mediators including IL-1β, IL-6, IL-8, MCP-1, MIP-1α/β, interferon-gamma inducible protein 10 kDa (IP-10) and interferon-beta (IFN-β) in a somewhat strain-specific manner (Knepper et al. 2013; Weinheimer et al. 2012; Wu et al. 2010b, 2012). However, a subsequent strain-related analysis of mediator expression is still lacking and limited information is available about the mediator origin (Wu et al. 2010b). Experiments with chemical inhibitors have suggested an important role of MAPKs for the regulation of inflammatory mediator expression (Wu et al. 2010b) and IAV seem to interfere with the expression of the pattern-recognition receptor retinoic acid-inducible gene I (RIG-I; Wu et al. 2012). Patients with COPD are prone to IAV infections, and IAV vaccination is highly warranted in those suffering from COPD (Sehatzadeh 2012). By using human lung explants exposed to cigarette smoke extract before IAV infection Wu et al. (2011) demonstrated altered IFN and IP-10 expression and IAV-mediated RIG-I upregulation, suggesting that smoking impairs the host response to IAV.
That these models might be useful for the testing of new therapeutic approaches is indicated by a report of Chan et al. (R.W. 2009). The binding to sialic acids (alpha2-6-linked and alpha2-3-linked depending on the viral strain) on the host cell membrane is an integral step of the IAV infection process. R.W. Chan et al. (2009) demonstrated that DAS181 induces de-sialylation of both sialic acids in ex vivo human lung tissue and that two doses of DAS181 treatment given 12 h post-infection are sufficient to block H5N1 infection in ex vivo lung tissue culture.
Adenovirus 7
The human adenovirus 7 belongs to the Adenoviridae subgroup B (HAdV-B7), which causes pneumonia and systemic disease in both immuno-compromised and non-immuno-compromised hosts, mainly in regional outbreaks (Ng et al. 2015; Scott et al. 2016). In addition to being human pathogens, recombinant adenoviruses attracted the interest of researchers, several years ago, as promising tools for gene therapy but the lack of suitable animal (Ginsberg et al. 1990) and cell culture (Jogler et al. 2006) models supporting human HAdV replication hampered research progress. Booth et al. (2004) thus used a human ex vivo infection model with human HAdV-B7 and showed efficient viral replication. HAdV-B7 provoked activation of ERK kinases and ERK inhibition blocked the release of IL-8. In a subsequent study (Wu et al. 2010a), the same group showed the liberation of IL-6, IL-8, IP-10, MIP-1α/β and MCP-1 in infected tissue. Multicolor immunostaining documented the infection of AEC-I and AEC-II cells. The authors identified lung AEC as the primary source for IL-8, whereas IP-10 was found in AM and epithelial cells. Overall, this model now gives a solid basis for further assessment of HAdV viral replication and pathology.
Coronaviruses
Infections with coronaviruses (CoVs) in humans primarily target the upper respiratory tract and, in most cases, induce a rather mild, self-limiting disease, such as the common cold (Su et al. 2016). However, SARS-CoV and MERS-CoV differ from the other CoV. In 2002/2003, SARS-CoV caused a global outbreak of a severe respiratory disease killing over 700 people and illustrating the potential worldwide impact of a new interspecies transmission of a highly pathogenic zoonotic virus (Peiris et al. 2004). An important initial step for the understanding of such emerging viral infections of the lung is the identification of host cell receptors and primary target cells. By using human SARS-CoV-infected lung tissue slices, the function of angiotensin-converting enzyme 2 as a human receptor for SARS-CoV could be substantiated (V.S. Chan et al. 2006). Furthermore, results indicate that a subpopulation of lung cells expressing stem/progenitor cell markers CD34 and Oct-4 (while being negative for cytokeratin or surfactant) may be important target cells of SARS-CoV in human lungs (Chen et al. 2007). However, the way in which SARS-CoV interacts with the human lung alveolus is far away from being understood and research efforts to elucidate its pathobiology have been paralyzed with the temporal distance of the outbreak.
In 2012, MERS-CoV (originally named human coronavirus-EMC) emerged as a new CoV causing an acute respiratory syndrome in humans (Fehr et al. 2016; Mohd et al. 2016; Zumla et al. 2015). Probably originating in bats, the MERS-CoV infection is endemic in dromedary camel populations of East Africa and the Middle East (Mohd et al. 2016). Most human cases are based on dromedary camel to human transmission, although, under some circumstances, significant human-to-human transmission also occur (Cho et al. 2016; Drosten et al. 2014, 2015). MERS-CoV uses, as a receptor, the human Dipeptidyl peptidase 4 (DPP4; Raj et al. 2013), which is not present in the most frequently used animal models (Gretebeck and Subbarao 2015; Sutton and Subbarao 2015), thus impeding its analysis in complex lung tissue.
Hocke et al. (2013a) and R.W. Chan et al. (2014) used ex vivo infection of human lungs to assess viral replication and cellular tropism of MERS-CoV. Both groups infected the tissue with the original human MERS-CoV Erasmus Medical Center strain and the group of R.W. Chan (2014) expanded the observations to strains isolated from dromedaries. Both groups showed strong viral replication in lung tissue. Bronchial epithelial cells, AEC-I, AEC-II cells and endothelial cells were shown to be infected (R.W. Chan et al. 2014; Hocke et al. 2013a). Indeed, electron microscopy showed the presence of MERS-CoV in the AEC-I and AEC-II cells (Hocke et al. 2013a). The presence of virus particles in the basal lamina below intact AEC suggested the basolateral release of MERS-CoV (Hocke et al. 2013a). Notably, both reports showed no evidence for the infection of AM. DPP4 was present in all cell types infected (Hocke et al. 2013a) and this broad receptor expression might be one of the major factors for the observed widespread cellular tropism of MERS-CoV. To gain insight into the mechanistic cause of lung failure in MERS-CoV, Hocke et al. (2013a) assessed the cell death of epithelial cells and the integrity of the alveolar tight junction protein occludin. Detachment of apoptotic MERS-CoV–infected AEC-II from the alveolar base membrane with disruption of alveolar tight junctions (indicated by the disintegration of the occludin protein band) indicated structural lung damage caused by MERS-CoV (Fig. 2). Notably, the first (and only) published autopsy performed on a fatal case of MERS-CoV essentially confirmed the results of the ex vivo models with respect to viral tissue tropism and damage (Ng et al. 2016). Because of the persistent reintroduction of the virus into the human population and the lack of specific therapies, an ongoing need exists to investigate MERS-CoV interaction with the human host.
MERS coronavirus (MERS-CoV) causes structural damage in ex vivo infected human lungs. Detachment of MERS-CoV-infected cell (green) from the alveolar epithelial layer disrupts epithelial continuity. An annular formation of tight junction protein occludin (red, white arrowheads) still surrounds the detached cell and is dissolved from the alveolar junctional band (white arrow). Stain visualization by confocal microscopy and separation of tissue autofluorescence from specific fluorescence by spectral unmixing. From Hocke et al. (2013a). Reprinted with permission of the American Thoracic Society. Copyright © 2016 American Thoracic Society
Summary and outlook
The presented studies indicate that ex vivo infected human lung tissue is useful for the investigation of basic principles of pathogen-host interaction in the lung. A great advantage of this model is the possibility of using wild-type patient isolates of bacteria and viruses for the investigation of general pathogenicity and risk assessment of emergent pathogens (e.g., new zoonotic viruses) in original three-dimensional tissue. Many studies focus on pathogen replication, cellular tropism and tissue activation in the sense of the release of immunomodulatory factors. Despite this obvious value, many areas of human tissue culture per se can be improved and, in particular, many further directions can be taken in the investigation of infectious lung diseases.
One important issue is that we need to know, in more detail and on several levels, “what is in the box”. Most studies involve the use of peripheral (alveolar) lung material obtained from surgery because of lung cancer; some groups have investigated lung tissue not used for transplantation. Although tumor-free tissue is used, we cannot rule out that the very presence of the tumor in the lung has altered tissue responses or that, for example, mechanical ventilation before explantation has affected sample responses. Donors of lung tissue might have a long history of smoking and might suffer from additional diseases. However, if sample numbers in the laboratories increase, a detailed stratification of patient history and an analysis of tissue responses may help us to understand the way that smoking, COPD, asthma, or diabetes mellitus and chronic heart disease lead to increased susceptibility to pneumonia (Torres et al. 2015). Lung tissue contains a huge variety of cellular components and even nowadays, new cell types are being identified (Franks et al. 2008). We need to use robust technologies such as advanced cell sorting (Fujino et al. 2012; Gross et al. 2015) and microscopy to identify cells present in our samples and to investigate any changes in these populations over culture time and during lung-pathogen interaction. This step is a prerequisite for the in-depth analysis of the role of those hitherto neglected cells for the responses to lung infection. For example, Piet et al. (2011) used human lung tissue-derived T cells to investigate the specificity and function of CD103+CD8+ T cells. Their study indicated that the human lung might harbor local virus-specific epithelial CD8+ T cells that might protect the lung against recurring IAV infection (Piet et al. 2011). Techniques such as laser-assisted cell picking for the extraction of single cells out of infected lung tissue (Fink et al. 1998) combined with recently developed assays for single-cell transcriptome analysis (Bell and Eberwine 2015; Grun et al. 2015; Hodne and Weltzien 2015; Macaulay et al. 2016) might help to expand our knowledge of cell-specific responses in the human lung. Indeed, the combination of such cutting-edge technologies with complex human samples is a challenge and needs strong interdisciplinary collaboration. Microscopy-based documentation of live-dead staining of cells and subsequent three-dimensional reconstruction will help us to gain insight into tissue viability over time and into the specific effects of pathogens and their virulence factors (e.g., PLY release of pneumocooci).
The preservation of the complex three-dimensional structure of the lung is one of the great advantages of these models and high-end microscopy is a key to obtaining information about pathogen-host interaction in this complex organ architecture. Unfortunately, human lung tissue contains very strong autofluorescent structures (e.g., collagen, elastin) causing significant overlap with fluorophore emission spectra. Spectral confocal microscopy can be used to achieve higher signal-to-noise ratios as demonstrated in some studies (Hocke et al. 2013a, b; Szymanski et al. 2012; Weinheimer et al. 2012). On the other hand, autofluorescence itself may be useful for the visualization of tissue morphology and cellular dynamics in human lung tissue by performing autofluorescence multiphoton microscopy (Kretschmer et al. 2016). No technical reason exists as to why the real-time imaging of pulmonary reactions combined with micropuncture techniques, which have previously been successfully used in animal experiments (Islam et al. 2012; Kreisel et al. 2010; Looney and Bhattacharya 2014; Westphalen et al. 2014) should not also be used for the dissection of intra-alveolar host-pathogen interactions in ex vivo human lung models.
A next step is to advance from phenomenological studies to mechanistic investigations in order to improve the analytical potency of the model. Beyond the use of chemical inhibitors or the testing of innovative anti-infective drugs, viral transformation (McBride et al. 2000) allowing cell-specific functional analysis is a possible strategy. This can be combined with gene editing approaches such as the CRISPR-Cas9 system (Chen and Goncalves 2016; Wang and Qi 2016), thereby permitting mechanistic studies.
In general, we need to be able to establish longer durations of tissue cultivation. For example, viral transformation and subsequent gene editing, together with studies addressing tissue injury and repair, would profit from expanded observation times. The rapid emergence of sophisticated microfluidic systems (Esch et al. 2015) better mimicking organ supply and possibly including physical forces might lead to longer periods of lung tissue cultivation. Finally, studies are hampered by limited tissue availability, a limitation that could, at least in part, be overcome by the rigorous improvement of human lung-specific cryopreservation methods (Baatz et al. 2014; Bai et al. 2016; Rosner et al. 2014).
Beyond the analytical power of these models, we wish to stress that the use of human lung tissue reduces the burden of animal experiments by contributing to the 3-R principle to replace, reduce, or refine animal experiments (Russell 1995).
Overall, ex vivo infection models of human lung tissue are today of great value for the investigation of pneumonia-related host-pathogen interaction. The combination of these models with the now available cutting-edge technologies will booster the mechanistic understanding of pneumonia in humans.
References
Baatz JE, Newton DA, Riemer EC, Denlinger CE, Jones EE, Drake RR, Spyropoulos DD (2014) Cryopreservation of viable human lung tissue for versatile post-thaw analyses and culture. In vivo 28:411–423
Bai Y, Krishnamoorthy N, Patel KR, Rosas I, Sanderson MJ, Ai X (2016) Cryopreserved human precision-Cut lung slices as a bioassay for live tissue banking. A viability study of bronchodilation with bitter-taste receptor agonists. Am J Respir Cell Mol Biol 54:656–663
Baron RM, Choi AJ, Owen CA, Choi AM (2012) Genetically manipulated mouse models of lung disease: potential and pitfalls. Am J Physiol Lung Cell Mol Physiol 302:L485–L497
Bauer TT, Ewig S, Rodloff AC, Muller EE (2006) Acute respiratory distress syndrome and pneumonia: a comprehensive review of clinical data. Clin Infect Dis 43:748–756
Bean AG, Baker ML, Stewart CR, Cowled C, Deffrasnes C, Wang LF, Lowenthal JW (2013) Studying immunity to zoonotic diseases in the natural host—keeping it real. Nat Rev Immunol 13:851–861
Bell TJ, Eberwine J (2015) Live cell genomics: cell-specific transcriptome capture in live tissues and cells. Methods Mol Biol 1324:447–456
Bogaert D, De Groot R, Hermans PW (2004) Streptococcus pneumoniae colonisation: the key to pneumococcal disease. Lancet Infect Dis 4:144–154
Booth JL, Coggeshall KM, Gordon BE, Metcalf JP (2004) Adenovirus type 7 induces interleukin-8 in a lung slice model and requires activation of Erk. J Virol 78:4156–4164
Booth JL, Duggan ES, Patel VI, Langer M, Wu W, Braun A, Coggeshall KM, Metcalf JP (2016) Bacillus anthracis spore movement does not require a carrier cell and is not affected by lethal toxin in human lung models. Microbes Infect 18:615–626
Bouvier NM (2015) Animal models for influenza virus transmission studies: a historical perspective. Curr Opin Virol 13:101–108
Brown ED, Wright GD (2016) Antibacterial drug discovery in the resistance era. Nature 529:336–343
Chakrabarty K, Wu W, Booth JL, Duggan ES, Nagle NN, Coggeshall KM, Metcalf JP (2007) Human lung innate immune response to Bacillus anthracis spore infection. Infect Immun 75:3729–3738
Chan MC, Chan RW, Yu WC, Ho CC, Yuen KM, Fong JH, Tang LL, Lai WW, Lo AC, Chui WH, Sihoe AD, Kwong DL, Wong DS, Tsao GS, Poon LL, Guan Y, Nicholls JM, Peiris JS (2010) Tropism and innate host responses of the 2009 pandemic H1N1 influenza virus in ex vivo and in vitro cultures of human conjunctiva and respiratory tract. Am J Pathol 176:1828–1840
Chan MC, Chan RW, Chan LL, Mok CK, Hui KP, Fong JH, Tao KP, Poon LL, Nicholls JM, Guan Y, Peiris JS (2013) Tropism and innate host responses of a novel avian influenza A H7N9 virus: an analysis of ex-vivo and in-vitro cultures of the human respiratory tract. Lancet Respir Med 1:534–542
Chan RW, Chan MC, Wong AC, Karamanska R, Dell A, Haslam SM, Sihoe AD, Chui WH, Triana-Baltzer G, Li Q, Peiris JS, Fang F, Nicholls JM (2009) DAS181 inhibits H5N1 influenza virus infection of human lung tissues. Antimicrob Agents Chemother 53:3935–3941
Chan RW, Hemida MG, Kayali G, Chu DK, Poon LL, Alnaeem A, Ali MA, Tao KP, Ng HY, Chan MC, Guan Y, Nicholls JM, Peiris JS (2014) Tropism and replication of Middle East respiratory syndrome coronavirus from dromedary camels in the human respiratory tract: an in-vitro and ex-vivo study. Lancet Respir Med 2:813–822
Chan VS, Chan KY, Chen Y, Poon LL, Cheung AN, Zheng B, Chan KH, Mak W, Ngan HY, Xu X, Screaton G, Tam PK, Austyn JM, Chan LC, Yip SP, Peiris M, Khoo US, Lin CL (2006) Homozygous L-SIGN (CLEC4M) plays a protective role in SARS coronavirus infection. Nat Genet 38:38–46
Chen X, Goncalves MA (2016) Engineered viruses as genome editing devices. Mol Ther 24:447–457
Chen Y, Chan VS, Zheng B, Chan KY, Xu X, To LY, Huang FP, Khoo US, Lin CL (2007) A novel subset of putative stem/progenitor CD34+ Oct-4+ cells is the major target for SARS coronavirus in human lung. J Exp Med 204:2529–2536
Chi F, Leider M, Leendertz F, Bergmann C, Boesch C, Schenk S, Pauli G, Ellerbrok H, Hakenbeck R (2007) New Streptococcus pneumoniae clones in deceased wild chimpanzees. J Bacteriol 189:6085–6088
Cho SY, Kang JM, Ha YE, Park GE, Lee JY, Ko JH, Lee JY, Kim JM, Kang CI, Jo IJ, Ryu JG, Choi JR, Kim S, Huh HJ, Ki CS, Kang ES, Peck KR, Dhong HJ, Song JH, Chung DR, Kim YJ (2016) MERS-CoV outbreak following a single patient exposure in an emergency room in South Korea: an epidemiological outbreak study. Lancet 388:994–1001
Cilloniz C, Torres A, Niederman M, Eerden M van der, Chalmers J, Welte T, Blasi F (2016) Community-acquired pneumonia related to intracellular pathogens. Intensive Care Med 42:1374–1386
Claar D, Hartert TV, Peebles RS Jr (2015) The role of prostaglandins in allergic lung inflammation and asthma. Expert Rev Respir Med 9:55–72
Cunha LD, Zamboni DS (2014) Recognition of Legionella pneumophila nucleic acids by innate immune receptors. Microbes Infect 16:985–990
Cunha LD, Ribeiro JM, Fernandes TD, Massis LM, Khoo CA, Moffatt JH, Newton HJ, Roy CR, Zamboni DS (2015) Inhibition of inflammasome activation by Coxiella burnetii type IV secretion system effector IcaA. Nat Commun 6:10205
Dalhoff A, Shalit I (2003) Immunomodulatory effects of quinolones. Lancet Infect Dis 3:359–371
Dorrington MG, Roche AM, Chauvin SE, Tu Z, Mossman KL, Weiser JN, Bowdish DM (2013) MARCO is required for TLR2- and Nod2-mediated responses to Streptococcus pneumoniae and clearance of pneumococcal colonization in the murine nasopharynx. J Immunol 190:250–258
Drijkoningen JJ, Rohde GG (2014) Pneumococcal infection in adults: burden of disease. Clin Microbiol Infect 20(Suppl 5):45–51
Dromann D, Rupp J, Rohmann K, Osbahr S, Ulmer AJ, Marwitz S, Roschmann K, Abdullah M, Schultz H, Vollmer E, Zabel P, Dalhoff K, Goldmann T (2010) The TGF-beta-pseudoreceptor BAMBI is strongly expressed in COPD lungs and regulated by nontypeable Haemophilus influenzae. Respir Res 11:67
Drosten C, Meyer B, Muller MA, Corman VM, Al-Masri M, Hossain R, Madani H, Sieberg A, Bosch BJ, Lattwein E, Alhakeem RF, Assiri AM, Hajomar W, Albarrak AM, Al-Tawfiq JA, Zumla AI, Memish ZA (2014) Transmission of MERS-coronavirus in household contacts. N Engl J Med 371:828–835
Drosten C, Muth D, Corman VM, Hussain R, Al Masri M, HajOmar W, Landt O, Assiri A, Eckerle I, Al Shangiti A, Al-Tawfiq JA, Albarrak A, Zumla A, Rambaut A, Memish ZA (2015) An observational, laboratory-based study of outbreaks of middle East respiratory syndrome coronavirus in Jeddah and Riyadh, Kingdom of Saudi Arabia, 2014. Clin Infect Dis 60:369–377
Duell BL, Su YC, Riesbeck K (2016) Host-pathogen interactions of nontypeable Haemophilus influenzae: from commensal to pathogen. FEBS Lett 590:3840–3853
Ensminger AW (2016) Legionella pneumophila, armed to the hilt: justifying the largest arsenal of effectors in the bacterial world. Curr Opin Microbiol 29:74–80
Esch EW, Bahinski A, Huh D (2015) Organs-on-chips at the frontiers of drug discovery. Nat Rev Drug Discov 14:248–260
Fatykhova D, Rabes A, Machnik C, Guruprasad K, Pache F, Berg J, Toennies M, Bauer TT, Schneider P, Schimek M, Eggeling S, Mitchell TJ, Mitchell AM, Hilker R, Hain T, Suttorp N, Hippenstiel S, Hocke AC, Opitz B (2015) Serotype 1 and 8 pneumococci evade sensing by inflammasomes in human lung tissue. PLoS One 10:e0137108
Fehr AR, Channappanavar R, Perlman S (2016) Middle east respiratory syndrome: emergence of a pathogenic human coronavirus. Annu Rev Med (in press)
Fernandez IE, Eickelberg O (2012) New cellular and molecular mechanisms of lung injury and fibrosis in idiopathic pulmonary fibrosis. Lancet 380:680–688
Fink L, Seeger W, Ermert L, Hanze J, Stahl U, Grimminger F, Kummer W, Bohle RM (1998) Real-time quantitative RT-PCR after laser-assisted cell picking. Nat Med 4:1329–1333
Franks TJ, Colby TV, Travis WD, Tuder RM, Reynolds HY, Brody AR, Cardoso WV, Crystal RG, Drake CJ, Engelhardt J, Frid M, Herzog E, Mason R, Phan SH, Randell SH, Rose MC, Stevens T, Serge J, Sunday ME, Voynow JA, Weinstein BM, Whitsett J, Williams MC (2008) Resident cellular components of the human lung: current knowledge and goals for research on cell phenotyping and function. Proc Am Thorac Soc 5:763–766
Friebe S, Goot FG van der, Burgi J (2016) The ins and outs of anthrax toxin. Toxins (Basel) 8:69
Fujino N, Kubo H, Ota C, Suzuki T, Suzuki S, Yamada M, Takahashi T, He M, Suzuki T, Kondo T, Yamaya M (2012) A novel method for isolating individual cellular components from the adult human distal lung. Am J Respir Cell Mol Biol 46:422–430
Fujino N, Kubo H, Ota C, Suzuki T, Takahashi T, Yamada M, Suzuki S, Kondo T, Nagatomi R, Tando Y, Yamaya M (2013) Increased severity of 2009 pandemic influenza A virus subtype H1N1 infection in alveolar type II cells from patients with pulmonary fibrosis. J Infect Dis 207:692–693
Furuya Y, Furuya AK, Roberts S, Sanfilippo AM, Salmon SL, Metzger DW (2015) Prevention of influenza virus-induced immunopathology by TGF-beta produced during allergic asthma. PLoS Pathog 11:e1005180
Galka F, Wai SN, Kusch H, Engelmann S, Hecker M, Schmeck B, Hippenstiel S, Uhlin BE, Steinert M (2008) Proteomic characterization of the whole secretome of Legionella pneumophila and functional analysis of outer membrane vesicles. Infect Immun 76:1825–1836
Ganbat D, Seehase S, Richter E, Vollmer E, Reiling N, Fellenberg K, Gaede KI, Kugler C, Goldmann T (2016) Mycobacteria infect different cell types in the human lung and cause species dependent cellular changes in infected cells. BMC Pulm Med 16:19
Geno KA, Gilbert GL, Song JY, Skovsted IC, Klugman KP, Jones C, Konradsen HB, Nahm MH (2015) Pneumococcal capsules and their types: past, present, and future. Clin Microbiol Rev 28:871–899
Ginsberg HS, Horswood RL, Chanock RM, Prince GA (1990) Role of early genes in pathogenesis of adenovirus pneumonia. Proc Natl Acad Sci U S A 87:6191–6195
Global Burden of Disease Study C (2015) Global, regional, and national incidence, prevalence, and years lived with disability for 301 acute and chronic diseases and injuries in 188 countries, 1990-2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet 386:743–800
Goossens PL, Tournier JN (2015) Crossing of the epithelial barriers by Bacillus anthracis: the known and the unknown. Front Microbiol 6:1122
Graham JG, Winchell CG, Kurten RC, Voth DE (2016) Development of an ex vivo tissue platform to study the human lung response to Coxiella burnetii. Infect Immun 84:1438–1445
Gretebeck LM, Subbarao K (2015) Animal models for SARS and MERS coronaviruses. Curr Opin Virol 13:123–129
Gross A, Schoendube J, Zimmermann S, Steeb M, Zengerle R, Koltay P (2015) Technologies for single-cell isolation. Int J Mol Sci 16:16897–16919
Grun D, Lyubimova A, Kester L, Wiebrands K, Basak O, Sasaki N, Clevers H, Oudenaarden A van (2015) Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525:251–255
Halstead ES, Chroneos ZC (2015) Lethal influenza infection: is a macrophage to blame? Expert Rev Anti Infect Ther 13:1425–1428
Herold S, Ludwig S, Pleschka S, Wolff T (2012) Apoptosis signaling in influenza virus propagation, innate host defense, and lung injury. J Leukoc Biol 92:75–82
Heyl KA, Klassert TE, Heinrich A, Muller MM, Klaile E, Dienemann H, Grunewald C, Bals R, Singer BB, Slevogt H (2014) Dectin-1 is expressed in human lung and mediates the proinflammatory immune response to nontypeable Haemophilus influenzae. MBio 5:e01492–e01414
Hocke AC, Becher A, Knepper J, Peter A, Holland G, Tonnies M, Bauer TT, Schneider P, Neudecker J, Muth D, Wendtner CM, Ruckert JC, Drosten C, Gruber AD, Laue M, Suttorp N, Hippenstiel S, Wolff T (2013a) Emerging human middle East respiratory syndrome coronavirus causes widespread infection and alveolar damage in human lungs. Am J Respir Crit Care Med 188:882–886
Hocke AC, Berg J, Becher A, Knepper J, Klauschen F, Tonnies M, Bauer TT, Schneider P, Neudecker J, Ruckert JC, Gruber AD, Suttorp N, Hippenstiel S, Wolff T (2013b) Reply to Fujino et al. J Infect Dis 207:693–695
Hodne K, Weltzien FA (2015) Single-cell isolation and gene analysis: pitfalls and possibilities. Int J Mol Sci 16:26832–26849
Hraiech S, Papazian L, Rolain JM, Bregeon F (2015) Animal models of polymicrobial pneumonia. Drug Des Devel Ther 9:3279–3292
Islam MN, Das SR, Emin MT, Wei M, Sun L, Westphalen K, Rowlands DJ, Quadri SK, Bhattacharya S, Bhattacharya J (2012) Mitochondrial transfer from bone-marrow-derived stromal cells to pulmonary alveoli protects against acute lung injury. Nat Med 18:759–765
Jager J, Marwitz S, Tiefenau J, Rasch J, Shevchuk O, Kugler C, Goldmann T, Steinert M (2014) Human lung tissue explants reveal novel interactions during Legionella pneumophila infections. Infect Immun 82:275–285
Jefferson T, Ferroni E, Curtale F, Giorgi Rossi P, Borgia P (2006) Streptococcus pneumoniae in western Europe: serotype distribution and incidence in children less than 2 years old. Lancet Infect Dis 6:405–410
Jogler C, Hoffmann D, Theegarten D, Grunwald T, Uberla K, Wildner O (2006) Replication properties of human adenovirus in vivo and in cultures of primary cells from different animal species. J Virol 80:3549–3558
Kadioglu A, Weiser JN, Paton JC, Andrew PW (2008) The role of Streptococcus pneumoniae virulence factors in host respiratory colonization and disease. Nat Rev Microbiol 6:288–301
Ke Y, Reddel RR, Gerwin BI, Miyashita M, McMenamin M, Lechner JF, Harris CC (1988) Human bronchial epithelial cells with integrated SV40 virus T antigen genes retain the ability to undergo squamous differentiation. Differentiation 38:60–66
Khan N, Vidyarthi A, Javed S, Agrewala JN (2016) Innate immunity holding the flanks until reinforced by adaptive immunity against Mycobacterium tuberculosis infection. Front Microbiol 7:328
Knepper J, Schierhorn KL, Becher A, Budt M, Tonnies M, Bauer TT, Schneider P, Neudecker J, Ruckert JC, Gruber AD, Suttorp N, Schweiger B, Hippenstiel S, Hocke AC, Wolff T (2013) The novel human influenza A(H7N9) virus is naturally adapted to efficient growth in human lung tissue. MBio 4:e00601–e00613
Kreisel D, Nava RG, Li W, Zinselmeyer BH, Wang B, Lai J, Pless R, Gelman AE, Krupnick AS, Miller MJ (2010) In vivo two-photon imaging reveals monocyte-dependent neutrophil extravasation during pulmonary inflammation. Proc Natl Acad Sci U S A 107:18073–18078
Kretschmer S, Pieper M, Huttmann G, Bolke T, Wollenberg B, Marsh LM, Garn H, Konig P (2016) Autofluorescence multiphoton microscopy for visualization of tissue morphology and cellular dynamics in murine and human airways. Lab Invest 96:918–931
Lai S, Qin Y, Cowling BJ, Ren X, Wardrop NA, Gilbert M, Tsang TK, Wu P, Feng L, Jiang H, Peng Z, Zheng J, Liao Q, Li S, Horby PW, Farrar JJ, Gao GF, Tatem AJ, Yu H (2016) Global epidemiology of avian influenza A H5N1 virus infection in humans, 1997-2015: a systematic review of individual case data. Lancet Infect Dis 16:e108–e118
Li N, Ren AH, Wang XS, Fan X, Zhao Y, Gao GF, Cleary P, Wang BN (2015) Influenza viral neuraminidase primes bacterial coinfection through TGF-beta-mediated expression of host cell receptors. Proc Natl Acad Sci U S A 112:238–243
Looney MR, Bhattacharya J (2014) Live imaging of the lung. Annu Rev Physiol 76:431–445
Lopez R, Garcia E (2004) Recent trends on the molecular biology of pneumococcal capsules, lytic enzymes, and bacteriophage. FEMS Microbiol Rev 28:553–580
Macaulay IC, Teng MJ, Haerty W, Kumar P, Ponting CP, Voet T (2016) Separation and parallel sequencing of the genomes and transcriptomes of single cells using G&T-seq. Nat Protoc 11:2081–2103
Mak IW, Evaniew N, Ghert M (2014) Lost in translation: animal models and clinical trials in cancer treatment. Am J Transl Res 6:114–118
McBride S, Rannie D, Harrison DJ (2000) Gene transfer to adult human lung tissue ex vivo. Gene Ther 7:675–678
McConnell BB, Yang VW (2010) Mammalian Kruppel-like factors in health and diseases. Physiol Rev 90:1337–1381
Mehr S, Wood N (2012) Streptococcus pneumoniae—a review of carriage, infection, serotype replacement and vaccination. Paediatr Respir Rev 13:258–264
Mestas J, Hughes CC (2004) Of mice and not men: differences between mouse and human immunology. J Immunol 172:2731–2738
Moayeri M, Leppla SH, Vrentas C, Pomerantsev AP, Liu S (2015) Anthrax pathogenesis. Annu Rev Microbiol 69:185–208
Moffatt JH, Newton P, Newton HJ (2015) Coxiella burnetii: turning hostility into a home. Cell Microbiol 17:621–631
Mohd HA, Al-Tawfiq JA, Memish ZA (2016) Middle east respiratory syndrome coronavirus (MERS-CoV) origin and animal reservoir. Virol J 13:87
Morrison J, Pai M, Hopewell PC (2008) Tuberculosis and latent tuberculosis infection in close contacts of people with pulmonary tuberculosis in low-income and middle-income countries: a systematic review and meta-analysis. Lancet Infect Dis 8:359–368
Mortimer L, Moreau F, MacDonald JA, Chadee K (2016) NLRP3 inflammasome inhibition is disrupted in a group of auto-inflammatory disease CAPS mutations. Nat Immunol 17:1176–1186
Mukherjee AB, Zhang Z, Chilton BS (2007) Uteroglobin: a steroid-inducible immunomodulatory protein that founded the secretoglobin superfamily. Endocr Rev 28:707–725
Muller-Redetzky HC, Wienhold SM, Berg J, Hocke AC, Hippenstiel S, Hellwig K, Gutbier B, Opitz B, Neudecker J, Ruckert J, Gruber AD, Kershaw O, Mayer K, Suttorp N, Witzenrath M (2015) Moxifloxacin is not anti-inflammatory in experimental pneumococcal pneumonia. J Antimicrob Chemother 70:830–840
Murphy N, Gaynor KU, Rowan SC, Walsh SM, Fabre A, Boylan J, Keane MP, McLoughlin P (2016) Altered expression of bone morphogenetic protein accessory proteins in murine and human pulmonary fibrosis. Am J Pathol 186:600–615
Naujoks J, Tabeling C, Dill BD, Hoffmann C, Brown AS, Kunze M, Kempa S, Peter A, Mollenkopf HJ, Dorhoi A, Kershaw O, Gruber AD, Sander LE, Witzenrath M, Herold S, Nerlich A, Hocke AC, van Driel I, Suttorp N, Bedoui S, Hilbi H, Trost M, Opitz B (2016) IFNs modify the proteome of legionella-containing vacuoles and restrict infection via IRG1-derived itaconic acid. PLoS Pathog 12:e1005408
Ng DL, Al Hosani F, Keating MK, Gerber SI, Jones TL, Metcalfe MG, Tong S, Tao Y, Alami NN, Haynes LM, Mutei MA, Abdel-Wareth L, Uyeki TM, Swerdlow DL, Barakat M, Zaki SR (2016) Clinicopathologic, immunohistochemical, and ultrastructural findings of a fatal case of middle east respiratory syndrome coronavirus infection in the United Arab Emirates, April 2014. Am J Pathol 186:652–658
Ng OT, Thoon KC, Chua HY, Tan NW, Chong CY, Tee NW, Lin RT, Cui L, Venkatachalam I, Tambyah PA, Chew J, Fong RK, Oh HM, Krishnan PU, Lee VJ, Tan BH, Ng SH, Ting PJ, Maurer-Stroh S, Gunalan V, Khong WX (2015) Severe pediatric adenovirus 7 disease in Singapore linked to recent outbreaks across Asia. Emerg Infect Dis 21:1192–1196
N’Guessan PD, Schmeck B, Ayim A, Hocke AC, Brell B, Hammerschmidt S, Rosseau S, Suttorp N, Hippenstiel S (2005) Streptococcus pneumoniae R6x induced p38 MAPK and JNK-mediated caspase-dependent apoptosis in human endothelial cells. Thromb Haemost 94:295–303
Nicholls JM, Chan MC, Chan WY, Wong HK, Cheung CY, Kwong DL, Wong MP, Chui WH, Poon LL, Tsao SW, Guan Y, Peiris JS (2007) Tropism of avian influenza A (H5N1) in the upper and lower respiratory tract. Nat Med 13:147–149
Novel Swine-Origin Influenza AVIT, Dawood FS, Jain S, Finelli L, Shaw MW, Lindstrom S, Garten RJ, Gubareva LV, Xu X, Bridges CB, Uyeki TM (2009) Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N Engl J Med 360:2605–2615
Ortiz JR, Perut M, Dumolard L, Wijesinghe PR, Jorgensen P, Ropero AM, Danovaro-Holliday MC, Heffelfinger JD, Tevi-Benissan C, Teleb NA, Lambach P, Hombach J (2016) A global review of national influenza immunization policies: analysis of the 2014 WHO/UNICEF Joint Reporting Form on immunization. Vaccine 34:5400–5405
Peiris JS, Guan Y, Yuen KY (2004) Severe acute respiratory syndrome. Nat Med 10:S88–S97
Peiris JS, Tu WW, Yen HL (2009) A novel H1N1 virus causes the first pandemic of the 21st century. Eur J Immunol 39:2946–2954
Phin N, Parry-Ford F, Harrison T, Stagg HR, Zhang N, Kumar K, Lortholary O, Zumla A, Abubakar I (2014) Epidemiology and clinical management of Legionnaires’ disease. Lancet Infect Dis 14:1011–1021
Piet B, Bree GJ de, Smids-Dierdorp BS, Loos CM van der, Remmerswaal EB, Thusen JH von der, Haarst JM van, Eerenberg JP, Brinke A ten, Bij W van der, Timens W, Lier RA van, Jonkers RE (2011) CD8(+) T cells with an intraepithelial phenotype upregulate cytotoxic function upon influenza infection in human lung. J Clin Invest 121:2254–2263
Poon LL, Guan Y, Nicholls JM, Yuen KY, Peiris JS (2004) The aetiology, origins, and diagnosis of severe acute respiratory syndrome. Lancet Infect Dis 4:663–671
Prashar A, Terebiznik MR (2015) Legionella pneumophila: homeward bound away from the phagosome. Curr Opin Microbiol 23:86–93
Raj VS, Mou H, Smits SL, Dekkers DH, Muller MA, Dijkman R, Muth D, Demmers JA, Zaki A, Fouchier RA, Thiel V, Drosten C, Rottier PJ, Osterhaus AD, Bosch BJ, Haagmans BL (2013) Dipeptidyl peptidase 4 is a functional receptor for the emerging human coronavirus-EMC. Nature 495:251–254
Rosner SR, Ram-Mohan S, Paez-Cortez JR, Lavoie TL, Dowell ML, Yuan L, Ai X, Fine A, Aird WC, Solway J, Fredberg JJ, Krishnan R (2014) Airway contractility in the precision-cut lung slice after cryopreservation. Am J Respir Cell Mol Biol 50:876–881
Rothen-Rutishauser B, Blank F, Muhlfeld C, Gehr P (2008) In vitro models of the human epithelial airway barrier to study the toxic potential of particulate matter. Expert Opin Drug Metab Toxicol 4:1075–1089
Ru YX, Li YC, Zhao Y, Zhao SX, Yang JP, Zhang HM, Pang TX (2011) Multiple organ invasion by viruses: pathological characteristics in three fatal cases of the 2009 pandemic influenza A/H1N1. Ultrastruct Pathol 35:155–161
Russell WM (1995) The development of the three Rs concept. Altern Lab Anim 23:298–304
Schmeck B, Gross R, N’Guessan PD, Hocke AC, Hammerschmidt S, Mitchell TJ, Rosseau S, Suttorp N, Hippenstiel S (2004) Streptococcus pneumoniae-induced caspase 6-dependent apoptosis in lung epithelium. Infect Immun 72:4940–4947
Schmeck B, N’Guessan PD, Ollomang M, Lorenz J, Zahlten J, Opitz B, Flieger A, Suttorp N, Hippenstiel S (2007) Legionella pneumophila-induced NF-kappaB- and MAPK-dependent cytokine release by lung epithelial cells. Eur Respir J 29:25–33
Schmeck B, Lorenz J, N’Guessan PD, Opitz B, Laak V van, Zahlten J, Slevogt H, Witzenrath M, Flieger A, Suttorp N, Hippenstiel S (2008) Histone acetylation and flagellin are essential for Legionella pneumophila-induced cytokine expression. J Immunol 181:940–947
Scott JR, Millar EV, Lipsitch M, Moulton LH, Weatherholtz R, Perilla MJ, Jackson DM, Beall B, Craig MJ, Reid R, Santosham M, O’Brien KL (2012) Impact of more than a decade of pneumococcal conjugate vaccine use on carriage and invasive potential in native American communities. J Infect Dis 205:280–288
Scott MK, Chommanard C, Lu X, Appelgate D, Grenz L, Schneider E, Gerber SI, Erdman DD, Thomas A (2016) Human adenovirus associated with severe respiratory infection, Oregon, USA, 2013-2014. Emerg Infect Dis 22:1044–1051
Sehatzadeh S (2012) Influenza and pneumococcal vaccinations for patients with chronic obstructive pulmonary disease (COPD): an evidence-based review. Ont Health Technol Assess Ser 12:1–64
Short KR, Kroeze EJ, Fouchier RA, Kuiken T (2014) Pathogenesis of influenza-induced acute respiratory distress syndrome. Lancet Infect Dis 14:57–69
Smith BT (1977) Cell line A549: a model system for the study of alveolar type II cell function. Am Rev Respir Dis 115:285–293
Su S, Wong G, Shi W, Liu J, Lai AC, Zhou J, Liu W, Bi Y, Gao GF (2016) Epidemiology, genetic recombination, and pathogenesis of coronaviruses. Trends Microbiol 24:490–502
Sutton TC, Subbarao K (2015) Development of animal models against emerging coronaviruses: from SARS to MERS coronavirus. Virology 479–480:247–258
Szymanski KV, Toennies M, Becher A, Fatykhova D, N’Guessan PD, Gutbier B, Klauschen F, Neuschaefer-Rube F, Schneider P, Rueckert J, Neudecker J, Bauer TT, Dalhoff K, Dromann D, Gruber AD, Kershaw O, Temmesfeld-Wollbrueck B, Suttorp N, Hippenstiel S, Hocke AC (2012) Streptococcus pneumoniae-induced regulation of cyclooxygenase-2 in human lung tissue. Eur Respir J 40:1458–1467
Teijaro JR, Walsh KB, Cahalan S, Fremgen DM, Roberts E, Scott F, Martinborough E, Peach R, Oldstone MB, Rosen H (2011) Endothelial cells are central orchestrators of cytokine amplification during influenza virus infection. Cell 146:980–991
Thangavel RR, Bouvier NM (2014) Animal models for influenza virus pathogenesis, transmission, and immunology. J Immunol Methods 410:60–79
Torres A, Peetermans WE, Viegi G, Blasi F (2013) Risk factors for community-acquired pneumonia in adults in Europe: a literature review. Thorax 68:1057–1065
Torres A, Blasi F, Dartois N, Akova M (2015) Which individuals are at increased risk of pneumococcal disease and why? Impact of COPD, asthma, smoking, diabetes, and/or chronic heart disease on community-acquired pneumonia and invasive pneumococcal disease. Thorax 70:984–989
Trombetta C, Piccirella S, Perini D, Kistner O, Montomoli E (2015) Emerging influenza strains in the last two decades: a threat of a new pandemic? Vaccines (Basel) 3:172–185
Wagner C, Goldmann T, Rohmann K, Rupp J, Marwitz S, Rotta Detto Loria J, Limmer S, Zabel P, Dalhoff K, Dromann D (2015) Budesonide inhibits intracellular infection with non-typeable Haemophilus influenzae despite its anti-inflammatory effects in respiratory cells and human lung tissue: a role for p38 MAP kinase. Respiration 90:416–425
Walker CL, Rudan I, Liu L, Nair H, Theodoratou E, Bhutta ZA, O’Brien KL, Campbell H, Black RE (2013) Global burden of childhood pneumonia and diarrhoea. Lancet 381:1405–1416
Wang F, Qi LS (2016) Applications of CRISPR genome engineering in cell biology. Trends Cell Biol 26:875–888
Wang W, Mu X, Zhao L, Wang J, Chu Y, Feng X, Feng B, Wang X, Zhang J, Qiao J (2015) Transcriptional response of human umbilical vein endothelial cell to H9N2 influenza virus infection. Virology 482:117–127
Ware LB (2008) Modeling human lung disease in animals. Am J Physiol Lung Cell Mol Physiol 294:L149–L150
Weiner ZP, Glomski IJ (2012) Updating perspectives on the initiation of Bacillus anthracis growth and dissemination through its host. Infect Immun 80:1626–1633
Weinheimer VK, Becher A, Tonnies M, Holland G, Knepper J, Bauer TT, Schneider P, Neudecker J, Ruckert JC, Szymanski K, Temmesfeld-Wollbrueck B, Gruber AD, Bannert N, Suttorp N, Hippenstiel S, Wolff T, Hocke AC (2012) Influenza A viruses target type II pneumocytes in the human lung. J Infect Dis 206:1685–1694
Welte T, Torres A, Nathwani D (2012) Clinical and economic burden of community-acquired pneumonia among adults in Europe. Thorax 67:71–79
Westphalen K, Gusarova GA, Islam MN, Subramanian M, Cohen TS, Prince AS, Bhattacharya J (2014) Sessile alveolar macrophages communicate with alveolar epithelium to modulate immunity. Nature 506:503–506
Wolfe ND, Dunavan CP, Diamond J (2007) Origins of major human infectious diseases. Nature 447:279–283
Woods PS, Tazi MF, Chesarino NM, Amer AO, Davis IC (2015) TGF-beta-induced IL-6 prevents development of acute lung injury in influenza A virus-infected F508del CFTR-heterozygous mice. Am J Physiol Lung Cell Mol Physiol 308:L1136–L1144
Wu W, Booth JL, Duggan ES, Patel KB, Coggeshall KM, Metcalf JP (2010a) Human lung innate immune cytokine response to adenovirus type 7. J Gen Virol 91:1155–1163
Wu W, Booth JL, Duggan ES, Wu S, Patel KB, Coggeshall KM, Metcalf JP (2010b) Innate immune response to H3N2 and H1N1 influenza virus infection in a human lung organ culture model. Virology 396:178–188
Wu W, Patel KB, Booth JL, Zhang W, Metcalf JP (2011) Cigarette smoke extract suppresses the RIG-I-initiated innate immune response to influenza virus in the human lung. Am J Physiol Lung Cell Mol Physiol 300:L821–L830
Wu W, Zhang W, Booth JL, Metcalf JP (2012) Influenza A(H1N1)pdm09 virus suppresses RIG-I initiated innate antiviral responses in the human lung. PLoS One 7:e49856
Xu F, Droemann D, Rupp J, Shen H, Wu X, Goldmann T, Hippenstiel S, Zabel P, Dalhoff K (2008) Modulation of the inflammatory response to Streptococcus pneumoniae in a model of acute lung tissue infection. Am J Respir Cell Mol Biol 39:522–529
Zahlten J, Steinicke R, Opitz B, Eitel J, N’Guessan PD, Vinzing M, Witzenrath M, Schmeck B, Hammerschmidt S, Suttorp N, Hippenstiel S (2010) TLR2- and nucleotide-binding oligomerization domain 2-dependent Kruppel-like factor 2 expression downregulates NF-kappa B-related gene expression. J Immunol 185:597–604
Zahlten J, Steinicke R, Bertrams W, Hocke AC, Scharf S, Schmeck B, Witzenrath M, Hammerschmidt S, Suttorp N, Hippenstiel S (2013) TLR9- and Src-dependent expression of Krueppel-like factor 4 controls interleukin-10 expression in pneumonia. Eur Respir J 41:384–391
Zahlten J, Herta T, Kabus C, Steinfeldt M, Kershaw O, Garcia P, Hocke AC, Gruber AD, Hubner RH, Steinicke R, Doehn JM, Suttorp N, Hippenstiel S (2015a) Role of pneumococcal autolysin for KLF4 expression and chemokine secretion in lung epithelium. Am J Respir Cell Mol Biol 53:544–554
Zahlten J, Kim YJ, Doehn JM, Pribyl T, Hocke AC, Garcia P, Hammerschmidt S, Suttorp N, Hippenstiel S, Hubner RH (2015b) Streptococcus pneumoniae-induced oxidative stress in lung epithelial cells depends on pneumococcal autolysis and is reversible by resveratrol. J Infect Dis 211:1822–1830
Zhang J, Zhang Z, Fan X, Liu Y, Wang J, Zheng Z, Chen R, Wang P, Song W, Chen H, Guan Y (2010) 2009 pandemic H1N1 influenza virus replicates in human lung tissues. J Infect Dis 201:1522–1526
Zhang JC, Chen G, Chen L, Meng ZJ, Xiong XZ, Liu HJ, Jin Y, Tao XN, Wu JH, Sun SW (2016) TGF-beta/BAMBI pathway dysfunction contributes to peripheral Th17/Treg imbalance in chronic obstructive pulmonary disease. Sci Rep 6:31911
Zhou W, Toki S, Zhang J, Goleniewksa K, Newcomb DC, Cephus JY, Dulek DE, Bloodworth MH, Stier MT, Polosuhkin V, Gangula RD, Mallal SA, Broide DH, Peebles RS Jr (2016) Prostaglandin I2 signaling and inhibition of group 2 innate lymphoid cell responses. Am J Respir Crit Care Med 193:31–42
Zhu H, Lam TT, Smith DK, Guan Y (2016) Emergence and development of H7N9 influenza viruses in China. Curr Opin Virol 16:106–113
Zumla A, George A, Sharma V, Herbert N, Cunliffe-Lister (Baroness Masham of Ilton) SLP (2013) WHO’s 2013 global report on tuberculosis: successes, threats, and opportunities. Lancet 382:1765–1767
Zumla A, Hui DS, Perlman S (2015) Middle East respiratory syndrome. Lancet 386:995–1007
Acknowledgments and funding information
This work is supported by the German Research Foundation (DFG; SFB-TR84, project B6, Z1a, to A.C.H., project B1 to N.S and project B6 to S.H.) and DFG-SFB-TR84 project TF1 to A.C.H and S.H. We thank all the organ material donors and the clinical partners for their generous support in making human-material-based research possible. The authors are grateful to the current and former members of our laboratory for their contributions and apologize to those authors whose relevant contributions could not be cited because of space limitations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Hocke, A.C., Suttorp, N. & Hippenstiel, S. Human lung ex vivo infection models. Cell Tissue Res 367, 511–524 (2017). https://doi.org/10.1007/s00441-016-2546-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-016-2546-z