Abstract
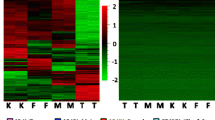
Abnormal patterns of DNA methylation are observed in many diseases such as tumors and imprinting disorders. Little is known about inter-individual and gender specific variations. Here, we report on accurate and sensitive quantitative measurements of methylation in DNA from total blood in 96 healthy human males and 96 healthy human females. Global methylation was estimated by studying two repetitive DNA elements, namely Line-1 and Alu repeats, while single loci were investigated for three differentially methylated regions (DMRs) at PEG3, NESP55 and H19 imprinted genes and two additional loci at Xq28 (F8 gene) and at 19q13.4 (locus between PEG3 and ubiquitin specific protease 29). We observed inter-individual correlations in the degree of methylation between Alu and Line-1 repeats. Moreover, all studied CpGs showed slightly higher methylation in males (P < 0.0003–0.0381), with the exception of DMRs at imprinted genes (P = 0.0342–0.9616) which were almost equally methylated in both sexes with only a small tendency towards higher methylation in males. This observed difference could be due to the process of X chromosome inactivation or merely to the presence of an additional X chromosome in female cells or could be a result of downstream effects of sex determination.


Similar content being viewed by others
References
Abrusan G, Krambeck HJ (2006) The distribution of L1 and Alu retroelements in relation to GC content on human sex chromosomes is consistent with the ectopic recombination model. J Mol Evol 63:484–492
Blewitt ME, Vickaryous NK, Hemley SJ, Ashe A, Bruxner TJ, Preis JI, Arkell R, Whitelaw E (2005) An N-ethyl-N-nitrosourea screen for genes involved in variegation in the mouse. Proc Natl Acad Sci USA 102:7629–7634
Bollati V, Baccarelli A, Hou L, Bonzini M, Fustinoni S, Cavallo D, Byun HM, Jiang J, Marinelli B, Pesatori AC, Bertazzi PA, Yang AS (2007) Changes in DNA methylation patterns in subjects exposed to low-dose benzene. Cancer Res 67:876–880
Djuric U, El-Maarri O, Lamb B, Kuick R, Seoud M, Coullin P, Oldenburg J, Hanash S, Slim R (2006) Familial molar tissues due to mutations in the inflammatory gene, NALP7, have normal postzygotic DNA methylation. Hum Genet 120:390–395
Durcova-Hills G, Burgoyne P, McLaren A (2004) Analysis of sex differences in EGC imprinting. Dev Biol 268:105–110
Durcova-Hills G, Hajkova P, Sullivan S, Barton S, Surani MA, McLaren A (2006) Influence of sex chromosome constitution on the genomic imprinting of germ cells. Proc Natl Acad Sci USA 103:11184–11188
Eads CA, Danenberg KD, Kawakami K, Saltz LB, Blake C, Shibata D, Danenberg PV, Laird PW (2000) MethyLight: a high-throughput assay to measure DNA methylation. Nucleic Acids Res 28:e32
Eckhardt F, Lewin J, Cortese R, Rakyan VK, Attwood J, Burger M, Burton J, Cox TV, Davies R, Down TA, Haefliger C, Horton R, Howe K, Jackson DK, Kunde J, Koenig C, Liddle J, Niblett D, Otto T, Pettett R, Seemann S, Thompson C, West T, Rogers J, Olek A, Berlin K, Beck S (2006) DNA methylation profiling of human chromosomes 6, 20 and 22. Nat Genet 38:1378–1385
El-Maarri O, Herbiniaux U, Walter J, Oldenburg J (2002) A rapid, quantitative, non-radioactive bisulfite-SNuPE- IP RP HPLC assay for methylation analysis at specific CpG sites. Nucleic Acids Res 30:e25
El-Maarri O (2004) SIRPH analysis: SNuPE with IP-RP-HPLC for quantitative measurements of DNA methylation at specific CpG sites. Methods Mol Biol 287:195–205
El-Maarri O, Kueper M, Oldenburg J, Walter J (2004) Quantitative DNA-methylation analysis by the bisulfite conversion method. In: Weissensteiner T, Griffin HG, Griffin A (eds) PCR technology: current innovations, 2nd edn. CRC press, London, pp 175–185
Ehrlich M (2006) Cancer-linked DNA hypomethylation and its relationship to hypermethylation. Curr Top Microbiol Immunol 310:251–274
Flanagan JM, Popendikyte V, Pozdniakovaite N, Sobolev M, Assadzadeh A, Schumacher A, Zangeneh M, Lau L, Virtanen C, Wang SC, Petronis A (2006) Intra- and interindividual epigenetic variation in human germ cells. Am J Hum Genet 79:67–84
Fraga MF, Ballestar E, Paz MF, Ropero S, Setien F, Ballestar ML, Heine-Suner D, Cigudosa JC, Urioste M, Benitez J, Boix-Chornet M, Sanchez-Aguilera A, Ling C, Carlsson E, Poulsen P, Vaag A, Stephan Z, Spector TD, Wu YZ, Plass C, Esteller M (2005) Epigenetic differences arise during the lifetime of monozygotic twins. Proc Natl Acad Sci USA 102:10604–10609
Fuke C, Shimabukuro M, Petronis A, Sugimoto J, Oda T, Miura K, Miyazaki T, Ogura C, Okazaki Y, Jinno Y (2004) Age related changes in 5-methylcytosine content in human peripheral leukocytes and placentas: an HPLC-based study. Ann Hum Genet 68:196–204
Gama-Sosa MA, Wang RY, Kuo KC, Gehrke CW, Ehrlich M (1983) The 5-methylcytosine content of highly repeated sequences in human DNA. Nucleic Acids Res 11:3087–3095
Handa V, Jeltsch A (2005) Profound flanking sequence preference of Dnmt3a and Dnmt3b mammalian DNA methyltransferases shape the human epigenome. J Mol Biol 348:1103–1112
Hansen RS (2003) X inactivation-specific methylation of LINE-1 elements by DNMT3B: implications for the Lyon repeat hypothesis. Hum Mol Genet 12:2559–2567
Hellman A, Chess A (2007) Gene body-specific methylation on the active X chromosome. Science 315:1141–1143
Karimi M, Johansson S, Stach D, Corcoran M, Grander D, Schalling M, Bakalkin G, Lyko F, Larsson C, Ekstrom TJ (2006) LUMA (LUminometric methylation assay)–a high throughput method to the analysis of genomic DNA methylation. Exp Cell Res 312:1989–1995
Kuo KC, McCune RA, Gehrke CW, Midgett R, Ehrlich M (1980) Quantitative reversed-phase high performance liquid chromatographic determination of major and modified deoxyribonucleosides in DNA. Nucleic Acids Res 8:4763–4776
Norris DP, Brockdorff N, Rastan S (1991) Methylation status of CpG-rich islands on active and inactive mouse X chromosomes. Mamm Genome 1:78–83
Ramsahoye BH (2002) Measurement of genome wide DNA methylation by reversed-phase high-performance liquid chromatography. Methods 27:156–161
Rakyan VK, Hildmann T, Novik KL, Lewin J, Tost J, Cox AV, Andrews TD, Howe KL, Otto T, Olek A, Fischer J, Gut IG, Berlin K, Beck S (2004) DNA methylation profiling of the human major histocompatibility complex: a pilot study for the human epigenome project. PLoS Biol 2:e405
Sandovici I, Kassovska-Bratinova S, Loredo-Osti JC, Leppert M, Suarez A, Stewart R, Bautista FD, Schiraldi M, Sapienza C (2005) Interindividual variability and parent of origin DNA methylation differences at specific human Alu elements. Hum Mol Genet 14:2135–2143
Sarter B, Long TI, Tsong WH, Koh WP, Yu MC, Laird PW (2005) Sex differential in methylation patterns of selected genes in Singapore Chinese. Hum Genet 117:402–403
Shimabukuro M, Sasaki T, Imamura A, Tsujita T, Fuke C, Umekage T, Tochigi M, Hiramatsu K, Miyazaki T, Oda T, Sugimoto J, Jinno Y, Okazaki Y (2006) Global hypomethylation of peripheral leukocyte DNA in male patients with schizophrenia: a potential link between epigenetics and schizophrenia J Psychiatr Res 41(12):1042–1046
Stach D, Schmitz OJ, Stilgenbauer S, Benner A, Dohner H, Wiessler M, Lyko F (2003) Capillary electrophoretic analysis of genomic DNA methylation levels. Nucleic Acids Res 31:e2
Tost J, Dunker J, Gut IG (2003) Analysis and quantification of multiple methylation variable positions in CpG islands by pyrosequencing. Biotechniques 35:152–156
Waterland RA, Dolinoy DC, Lin JR, Smith CA, Shi X, Tahiliani KG (2006) Maternal methyl supplements increase offspring DNA methylation at axin fused. Genesis 44:401–406
Weber M, Davies JJ, Wittig D, Oakeley EJ, Haase M, Lam WL, Schübeler D (2005) Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nat Genet 37:853–862
Weisenberger DJ, Campan M, Long TI, Kim M, Woods C, Fiala E, Ehrlich M, Laird PW (2005) Analysis of repetitive element DNA methylation by MethyLight. Nucleic Acids Res 33:6823–6836
Wilson IM, Davies JJ, Weber M, Brown CJ, Alvarez CE, MacAulay C, Schubeler D, Lam WL (2006) Epigenomics: mapping the methylome. Cell Cycle 5:155–158
Wolff GL, Kodell RL, Moore SR, Cooney CA (1998) Maternal epigenetics and methyl supplements affect agouti gene expression in Avy/a mice. FASEB J 12:949–957
Zvetkova I, Apedaile A, Ramsahoye B, Mermoud JE, Crompton LA, John R, Feil R, Brockdorff N (2005) Global hypomethylation of the genome in XX embryonic stem cells. Nat Genet 37:1274–1279
Acknowledgments
We would like to thank the blood donors for participating in this study and Doris Schmidt for her technical support with the pyrosequencing analysis.
Conflict of interest
The authors declare no competing of interests.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
El-Maarri, O., Becker, T., Junen, J. et al. Gender specific differences in levels of DNA methylation at selected loci from human total blood: a tendency toward higher methylation levels in males. Hum Genet 122, 505–514 (2007). https://doi.org/10.1007/s00439-007-0430-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00439-007-0430-3