Abstract
Main conclusion
OsNAC103 negatively regulates rice plant height by influencing the cell cycle and crosstalk of phytohormones.
Abstract
Plant height is an important characteristic of rice farming and is directly related to agricultural yield. Although there has been great progress in research on plant growth regulation, numerous genes remain to be elucidated. NAC transcription factors are widespread in plants and have a vital function in plant growth. Here, we observed that the overexpression of OsNAC103 resulted in a dwarf phenotype, whereas RNA interference (RNAi) plants and osnac103 mutants showed no significant difference. Further investigation revealed that the cell length did not change, indicating that the dwarfing of plants was caused by a decrease in cell number due to cell cycle arrest. The content of the bioactive cytokinin N6-Δ2-isopentenyladenine (iP) decreased as a result of the cytokinin synthesis gene being downregulated and the enhanced degradation of cytokinin oxidase. OsNAC103 overexpression also inhibited cell cycle progression and regulated the activity of the cell cyclin OsCYCP2;1 to arrest the cell cycle. We propose that OsNAC103 may further influence rice development and gibberellin–cytokinin crosstalk by regulating the Oryza sativa homeobox 71 (OSH71). Collectively, these results offer novel perspectives on the role of OsNAC103 in controlling plant architecture.
Similar content being viewed by others
Introduction
The plant height of rice plants is regulated by various factors. In addition to environmental conditions, numerous genes regulate plant height, and most of the regulatory pathways involve phytohormones, such as gibberellins, cytokinins, brassinosteroids, and auxins, which affect stem elongation and thickness (Margis-Pinheiro et al. 2005; Tanabe et al. 2005; Peng et al. 2014; Wang et al. 2018). Cytokinins influence plant height, development, and senescence, and have a crucial function in the cell cycle (Hwang et al. 2012; Liu et al. 2023). Cytokinins exist as free bases, ribosides, and ribotides (Miyawaki et al. 2006). Free-base cytokinins represent the biologically active forms, such as N6-Δ2-isopentenyladenine (iP), cis-zeation (cZ), and trans-zeation (tZ). However, ribosides, such as isopentenyladenine riboside (iPR), zeatin riboside (ZR), and dihydrozeatin (DHZ), are less active (Lomin et al. 2015). Recent findings suggested that ribotides can be directly converted into free-base forms by enzymes (Kudo et al. 2010). Active cytokinin levels are finely controlled by the enzymes that are involved in cytokinin biosynthesis and degradation. The homeostatic balance of cytokinin biosynthesis and catabolism mainly involves several enzymes, including isopentenyl transferase (IPT), cytochrome P450 monooxygenases (CYP75A), a cytokinin-activating enzyme (LONELY GUY, LOG), and a catabolic enzyme cytokinin oxidase/dehydrogenase (CKX) (Duan et al. 2019). In cytokinin synthesis, the initiation of iP and tZ biosynthesis catalyzed by IPT is also the rate-limiting enzyme (Kudo et al. 2010). Overexpression of CKXs reduces cytokinin levels and inhibits stem development (Ha et al. 2012). Overexpression of AtCKX2 significantly reduces the levels of different intermediate metabolites of cytokinins (Werner et al. 2001). Moreover, the overexpression of AtCKX5 may produce a more intense phenotype than other CKX genes (Ha et al. 2012). In rice, overexpression of OsCKX4 or OsCKX9 leads to shorter plant height and fewer grains. However, CRISPR/Cas9-generated osckx9 was also shown to have a similar phenotype (Duan et al. 2019). Gene alterations in the cytokinin signaling pathway also affect plant height. For instance, the overexpression of the type-A response regulator 6 (OsARR6) leads to reduced sensitivity to cytokinins and the development of dwarf phenotypes (Hirose et al. 2007; Gao et al. 2014). In tomato, Solanum lycopersicum response regulator 6 (SlRR6) is an important component of the cytokinin, gibberellin, and indole-3-acetic acid (IAA) signaling networks that influence plant height (Liu et al. 2023).
Cytokinins are key factors in controlling cell division and cell cycle in plants. Changes in the concentration of cytokinins affect cell division rate and cell cycle (Riou-Khamlichi et al. 1999; Schaller et al. 2014). The cell cycle in plants is divided into G1, S, G2, and M phases, with G1/S and G2/M serving as important progression nodes (Qi and Zhang 2019). The regulation of cell division depends on cyclins, cyclin-dependent kinases (CDKs), and their related complexes. CYCBs are specifically expressed during the G2/M phase (Haga et al. 2011; Schaller et al. 2014). OsCYCB2;2 interacts with CDKA1 to regulate cell cycle (Peng et al. 2014). In addition to the classic CYCA, CYCB, and CYCD families, a new type of cyclin has been identified, the P-type cyclin (CYCP, also known as CYCU) (Torres Acosta et al. 2004; La et al. 2006; Chevalier 2008; Deng et al. 2014; Chen et al. 2020). OsCYCP4 integrates phosphate starvation signal with cell division (Xu et al. 2020). Under low-phosphorous conditions, OsCYCP1;1 negatively regulates root growth (Deng et al. 2014). CYCP2;1 is a target of WOX9, which is required to activate meristem growth during the germination of Arabidopsis thaliana. Overexpression of CYCP2;1 rescues the short-stalk phenotype of wox9 (Polyn et al. 2015). Brassinosteroids promote mesocotyl elongation via CYCU2-regulated cell division (Sun et al. 2018).
Gibberellins are a class of diterpenoids, that are biosynthesized via complex pathways. According to recent studies, gibberellins are believed to have a significant impact on the height of rice plants (Itoh et al. 2002; Sasaki et al. 2002), panicle development (Su et al. 2021), and nitrogen fertilizer efficiency (Camut et al. 2021). GA1, GA3, GA4, and GA7 are the main bioactive gibberellins. The first two stages of gibberellin synthesis are catalyzed by CPS, KS, KO2, and KAO. Subsequently, the branches form different gibberellin intermediates (Yamaguchi 2008). Research has found that genetic variations associated with gibberellin-synthesis result in stunted plant growth (Sakamoto et al. 2004). Gibberellin contents can vary after the overexpression or mutation of various genes, leading to either shortening or increased cell length (Lo et al. 2008; Chen et al. 2015; Zheng et al. 2018). In pea plants with strigolactone (SL)-related mutations, SL may stimulate cell division to increase stem elongation, in contrast to increasing cell length. SL appears to act independently of gibberellins to stimulate stem growth (de Saint Germain et al. 2013).
Different phytohormones function together rather than in isolation to control growth and development. Studies have shown that knotted1-like homeobox (KNOX) genes mediate gibberellin–cytokinin crosstalk (Jasinski et al. 2005; Wu et al. 2016; Su et al. 2021). KNOX gene expression promotes cytokinin signaling and inhibits gibberellin signaling (Jasinski et al. 2005). Overexpression of knotted1 can increase cytokinin content and delay senescence (Ori et al. 1999). In potatoes, overexpression of potato homeobox 1 (POTH1) inhibits gibberellin synthesis and alters vegetative development (Rosin et al. 2003). Furthermore, the KNOX gene family plays an essential role in the regulation of organogenesis and plant morphogenesis. Overexpression of Oryza sativa homeobox 71 (OSH71) or Oryza sativa homeobox 15 (OSH15) shows a reduction in the culm length, blade length, and panicle axis (Postma-Haarsma et al. 2002). osh15 also shows defects in internode elongation and the development of epidermal and hypodermal cell types (Sato et al. 1998). Oryza sativa homeobox 59 (HOS59) overexpression lines have lower plant height and smaller grain size (Sheng et al. 2022).
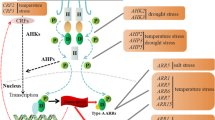
NAC (NAM, ATAF1/2, and CUC2) transcription factors are involved in multiple aspects of plant development and growth. Many genes have been shown to regulate plant height development. OsNAC2 regulates the response of gibberellins, cytokinins, and auxins to affect plant height and root development (Mao et al. 2020). XND1 (ANAC104) regulates lignocellulose synthesis and programmed xylem cell death, thereby affecting plant height (Zhao et al. 2008). OsNAC6 improves drought resistance by increasing the number and diameter of plant roots (Lee et al. 2017). However, the functions of more NAC family genes warrant investigation. Although the previous reports have found that gibberellins, drought, and low-temperature conditions can affect the expression level of OsNAC103, the other functions of OsNAC103 that regulate plant development still remain to be investigated (Jeong et al. 2010; Nuruzzaman et al. 2010, 2012, 2015).
In this study, we constructed transgenic plants and conducted a series of experiments to elucidate the regulation of OsNAC103 on plant growth. We measured the phytohormone contents to determine which phytohormones were affected by OsNAC103. To identify the genes and pathways involved in the plant height regulation of OsNAC103, we treated the seedlings with different phytohormones and detected the expression levels of related genes.
Materials and methods
Generation and cultivation of plant materials
To construct the OsNAC103 overexpression vector, the coding DNA sequence (CDS) of OsNAC103 was obtained from the leaf cDNA library of Taipei 309 (TP309, a japonica rice cultivar) and fused to pCXUN, which is an overexpression vector driven by the maize ubiquitin promoter. For β-glucosidase (GUS) staining, the 3981 bp genomic fragment upstream of ATG was amplified from wild-type (WT, TP309) genomic DNA and inserted into the GUS reporter vector pCXGUS. The target gene interference fragment was recombined with the interference vector pH7GWIWG2 (II) to construct the RNA interference (RNAi) vector. The vectors were transformed into TP309 calluses using the Agrobacterium strain EHA105 to obtain transgenic plants. Vector information for pCXUN and pCXGUS has been published previously (Chen et al. 2009). CRISPR/Csa9 mutant plants were obtained from BIOGLE Gene Tech Co., Ltd. (Jiangsu, China). The mutant rice variety used was Zhonghua 11 (ZH11, a japonica cultivar). The seedlings were grown in a culture room at 28 °C/25 °C (day/night). Seeds were obtained from rice plants grown under natural conditions in a field in Xiamen.
Microscopical observation
Cell length was measured in the middle of the second leaf sheath of the 21-day-old seedlings. Samples were soaked in ethanol for decolorization and photographed using a Leica DM4B microscope.
GUS staining
Samples were obtained from proOsNAC103:GUS transgenic plants. First, the samples were placed in a precooled 90% acetone solution for 30 min. After being washed thrice with pre-chilled ddH2O, the samples were incubated in a staining solution (50 mM sodium phosphate, (pH 7.2), 2 mM K3Fe(CN)6, 2 mM K4Fe(CN)6, 0.2% [v/v] Triton X-100, and 2 mM X-Gluc) for overnight at 37 °C. The samples were soaked in ethanol several times to rinse, decolorize, and remove the chlorophyll. The images were captured using a Leica M165 FC microscope.
Subcellular localization analysis of OsNAC103
To investigate the subcellular distribution of OsNAC103, the CDS of OsNAC103 was cloned and inserted into the pXDG vector to generate the 35S::GFP-OsNAC103 vector. The nuclear localization signal (NLS) sequence was cloned and inserted into the PXDR vector to obtain the vector 35S::RFP-NLS as a nuclear marker. The two vectors were transferred together into rice protoplasts for transient expression. The preparation and transformation of rice protoplasts were performed according to the methods described by Jiang et al. (2018). Fluorescence signals in the protoplasts were visualized using a confocal microscope (Zeiss, LSM780). Vector information for pXDG and pXDR has been published previously (Chen et al. 2009).
qRT-PCR analysis
An Eastep Universal RNA Extraction Kit (Promega) was used to extract total RNA. Promega GoScript was used to perform reverse transcription. qRT-PCR was performed using the SYBR Green Master Mix (Yeasen, Wuhan, China) in a LightCycler 480 system. qRT-PCR is performed as follows: 95 °C for 5 min, followed by 40 cycles at 95 °C for 5 s and 60 °C for 30 s. Three technical replicates and three biological replicates were prepared for every gene. The internal control was rice actin 1.
Treatment of plant materials
To measure the induced expression levels of OsNAC103 under various phytohormone treatments, WT plants were cultivated on 1/2 Murashige and Skoog (MS) medium. The seedlings at 21 days were treated with 100 µM N6-benzyladenine (6-BA) and 100 µM iP. Leaves were harvested at different time points after treatment. To evaluate phytohormone sensitivity, different plants were germinated and transferred to 1/2 MS alone or 1/2 MS with various concentrations of phytohormones (1 µM, 10 µM GA3; 1 µM, 10 µM 6-BA; 1 µM, 10 µM iP) or 10 µM paclobutrazol (PAC, a synthetic inhibitor of gibberellin) as the treatments for 10 days. For dark-induced stress experiments, leaves were subjected to 0, 2, 3, or 4 days without light to assess yellowing.
Yeast two-hybrid assay and yeast one-hybrid assay
The yeast two-hybrid assay was used to confirm whether OsNAC103 has transactivation activity. The OsNAC103 protein sequence was divided into two parts based on the NAM domain, one containing the protein sequence from 1 to 139 amino acids (OsNAC103△C) and the other containing the protein sequence from 140 to 346 amino acids (OsNAC103△N). The full-length OsNAC103 sequence and truncated sequence were cloned and inserted into the vector GAL4-BD (pGBKT7; Clontech). The fused vectors and GAL4-AD (pGADT7; Clontech) were cotransformed into the yeast strain Y2HGold. The yeast transformation protocol was based on the Clontech Yeast Two-Hybrid System. Co-transformant with pGBKT7-53 and pGADT7-T was used as a positive control. Co-transformant with pGBKT7-Lam and pGADT7-T was used as a negative control. The conserved domain range of the NAM domain of OsNAC103 was based on the Rice Genome Annotation Project Database.
To perform the yeast one-hybrid assay, the CDS region of OsNAC103 was fused to the vector pB42AD. Then, the construct was cotransferred into the yeast strain EGY48 with the LacZ reporter vector (pLacZi2µ) driven by the promoter of OSH71 (1774 bp upstream from ATG). The transformants were examined on SD/− Ura/− Trp plates and were chromogenic on plates containing X-gal. Co-transformant with pB42AD-HY5 and proCOP1 was used as a positive control.
Dual-luciferase assay in rice protoplasts
To assess the transcriptional activity of OsNAC103, the CDS of OsNAC103 was linked to the effector vector (pXSN) driven by the 35S promoter. The promoter of OsCYCP2;1 (1500 bp upstream from ATG) was fused to a reporter vector (pGreenII 0800-LUC). Different carrier combinations were co-transfected into the rice protoplasts in a ratio of 1:1. The preparation and transformation of rice protoplasts were performed according to the methods described by Jiang et al. (2018). The protoplasts were collected for dual-luciferase measurements. The detailed experimental methods for the Promega Dual-Luciferase Reporter Assay System are described in the manufacturer's instructions. The transcriptional activity levels were calculated using the relative LUC/REN ratio.
Measurements of phytohormones
To quantify phytohormones, the leaves of 21-day-old WT and OE-OsNAC103 seedlings were harvested. Each sample consisted of three biological samples. The phytohormone contents were analyzed at the Shanghai Applied Protein Technology Co., Ltd. (Shanghai, China). The samples were ground with liquid nitrogen, and samples weighing 100 ± 5 mg were placed in 2 mL centrifuge tubes. To ensure complete extraction, 30 μL of internal standard solution and 1.17 mL of acetonitrile were added to the 2 mL centrifuge tubes. The solution was vortexed until sufficiently mixed. The resulting mixture was ultrasonicated for 25 min at low temperature and avoiding light and then allowed to stand overnight at − 20 °C. After centrifugation (14,000 g, 4 °C, 20 min), the resulting supernatants were subjected to filtration and subsequently evaporated under N2 until dryness. Before mass spectrometry analysis, the extracts were dissolved again in 200 μL of a mixture of methanol and water (1:1, v/v). An AB SCIEX system was used for the mass spectrometry analysis, with the system set to the positive/negative ionization mode.
Electrophoretic mobility shift assay (EMSA)
The EMSA reaction system was as follows: 0.02% BSA, 8% glycerol, 0.5% Triton X-100, 10 × EMSA Binding Buffer [10 mM MgCl2, 200 mM KCl, 10 mM DTT, and 100 mM Hepes (pH = 7.8)], probe, purified GST-NAC103 protein or GST, and 300 ng of salmon essence, to which ddH2O was added to bring the volume up to 20 μL. The solution was thoroughly mixed and incubated for 30 min in the dark at room temperature. The probes were labeled with Texas Red. The EMSA reaction products were resolved on a 6% native polyacrylamide gel in TBE buffer. After electrophoresis, a Bio-Rad instrument and Texas Red filter were used for exposure.
Results
Comparative analysis of the OsNAC103 protein and its homologs
The OsNAC103 gene encodes a protein of 346 amino acids, that belongs to the NAC transcription factor family. OsNAC103 is a member of the NAP (NAC-Like, Activated by AP3/PI) subfamily, which includes seven rice NAP proteins (Fan et al. 2015). OsNAC58 and OsNAC131 (in the RAP-DB rice database), ANAC029 (ATNAP), and ANAC047 (in the TAIR Arabidopsis database) were highly homologous to OsNAC103. All these genes belong to the NAP family. The protein homologs of OsNAC103 from other species were screened using BLAST from the NCBI database. It was found that OsNAC103 had the highest similarity with QHE23802.1 (Phyllostachys edulis), followed by XP_044969400.7 (Hordeum vulgare) and XP_044319333.1 (Triticum aestivum). The above genes and some reported NAC transcription factors involved in plant development were analyzed by constructing a phylogenetic tree. These findings indicated that OsNAC103 clustered with other NAP family genes (Fig. 1a). Studies have shown that plants overexpressing OsNAC58 exhibit an obvious phenotype of yellowing and senescence (Liang et al. 2014), suggesting that the OsNAC103 gene may function similarly to OsNAC58.
Phylogenetic tree and conserved sequence analysis of OsNAC103 proteins homologs. a Phylogenetic tree analysis of protein homologs of OsNAC103. The neighbor-joining (NJ) phylogenetic tree was constructed using MEGA5. b Conserved motif analysis of OsNAC103 and homologous proteins. The MEME program was used to investigate the conserved motifs. The motif width was set from 6 to 200. The motif number was set to 10. Differently colored rectangles represent different domains. c The green label shows the location of the NAM conserved domains of OsNAC103 and homologous proteins
Researches show that NAC family proteins have a highly conserved NAC domain in their N-terminal regions and a variable C-terminal domain that functions as a transcriptional activation region (Ooka et al. 2003; Puranik et al. 2012). MEME was used for the motif analysis of all proteins in the phylogenetic tree. The conserved motifs are labeled in Fig. 1b, and ten motifs were screened. The NAM domains of the corresponding NAC transcription factor families in the phylogenetic tree are shown in Fig. 1c. Motifs 1–5 represent the conserved subdomains of the N-terminus of NAC transcription factors. OsNAC103 also has a motif 9 at its N-terminus. Compared to other genes, the NAP subfamily has more conserved sequences in the C-terminal region. In the NAP subfamily, in addition to OsNAC16, OsNAC131, ANAC029, and ANAC047, other proteins, including OsNAC103, also contain motif 8, motif 7, and motif 10 in the C-terminal transcriptional activation region. These differences may indicate that this protein has additional functions in plant growth and development.
OsNAC103 subcellular localization and expression pattern analysis
To further study the function of OsNAC103, a 35S::GFP-OsNAC103 vector was constructed and transferred to rice protoplasts for fluorescence observation. Subcellular localization analysis showed that OsNAC103 was mainly located in the nucleus, colocalizing with the signal of RFP fused with the nuclear localization signal (NLS; Fig. 2a). In terms of the expression pattern of OsNAC103, the analysis based on MBKbase revealed that OsNAC103 showed spatiotemporal expression at various stages of rice development (Fig. S1). To directly observe the tissue expression of OsNAC103, the gene promoter was inserted into a vector that included the GUS reporter and then transformed to obtain transgenic plants. GUS activity was detected in leaves, leaf sheaths, and roots (Fig. 2b i–iii). Furthermore, weaker GUS staining was detected in the first internode, stem node, and inner wall of the second internode (Fig. 2b iv–vi). qRT-PCR was performed on tissues during the seedling and mature stages. The highest expression level of OsNAC103 was observed in the leaves of 21-day-old seedlings, followed by the roots and leaf sheaths, and was the lowest in the internodes (Fig. 2c). These findings indicate that OsNAC103 may play a role in leaf and stem development.
OsNAC103 subcellular localization and tissue expression analysis. a The subcellular localization of OsNAC103. Bar = 10 μm. RFP-NLS was used as a nuclear marker. b GUS activity was detected in the young leaf (i), the young leaf sheath (ii), the young root (iii), the first internode (iv), the stem node (v), and the inner wall of the second internode of rice at maturity (vi). Bar = 1 mm. c Relative OsNAC103 expression levels in different tissues. Mean values ± SD, n = 3. Leaf, sheath, and root of 21-day-old seedling; internode I and internode II: the first internode and the second internode of mature plants
Identification of transcriptional activation of the OsNAC103 protein
To ascertain whether OsNAC103 has transcriptional activation activity, the OsNAC103 protein was fused to GAL4-BD. The results showed that yeast transformed with the full-length fusion protein BD-OsNAC103 could grow on the QDO/X/A medium, indicating that OsNAC103 had self-activation activity. Then, the OsNAC103 protein sequence was divided into two segments based on the location of the NAM domain. One contained the protein sequence from 1 to 139 amino acids (OsNAC103△C), while the other contained the protein sequence from 140 to 346 amino acids (OsNAC103△N). However, only yeast cells harboring BD-OsNAC103△N grew, while yeast cells carrying BD-OsNAC103△C did not grow on the QDO/X/A medium. These results indicate the transcriptional activation region at the C-terminus (Fig. 3). To investigate whether a specific portion of OsNAC103△N is the determining factor for transactivation activity, we further performed truncated experiments. Notably, the 186‒208 amino acid region may play a prominent role in regulating the transactivation activity of OsNAC103.
OsNAC103 negatively regulates plant height
To explore the function of OsNAC103 in rice, OsNAC103-overexpressing transgenic lines and RNA interference (RNAi) lines were constructed in the TP309 background, and CRISPR/Csa9 mutants were constructed in the ZH11 background.
OE-OsNAC103 plants exhibited a dwarf phenotype at the 21-day-old seedling stage (Fig. 4a, b). The lengths of the shoots, leaves, and leaf sheaths in OE-OsNAC103 seedlings were shorter than the corresponding values of the WT (Fig. 4c–e). Nevertheless, no notable differences were observed between the RNAi lines and the WT (Fig. 5a–e). At the heading stage, slower vegetative growth and lower plant height than those of the WT were observed in the OE4 and OE7 lines (Fig. 4f, g). The length of the internodes was reduced relative to that of the WT (Fig. 4h, i). Even during rice maturation, the RNAi plants did not exhibit notable variations in plant height (Fig. 5f–j). In addition, the phenotype of the osnac103 mutant also exhibited no notable disparity in plant height compared with that of WT (ZH11) (Fig. S2). Taken together, these results revealed that OsNAC103 negatively regulates plant height and development.
Phenotypes of OE-OsNAC103 plants. a The relative expression level of OsNAC103 of 21-day-old WT and OE-OsNAC103 plants. Mean values ± SD, n = 3. b The phenotype of 21-day-old WT and OE-OsNAC103 plants. Bar = 2 cm. c The shoot length of 21-day-old WT and OE-OsNAC103 plants. Mean ± SD, n = 20. d The leaf length of 21-day-old WT and OE-OsNAC103 plants. Mean ± SD, n = 25. e The leaf sheath length of of 21-day-old WT and OE-OsNAC103 plants. Mean ± SD, n = 28. f The plant height of mature WT and OE-OsNAC103 plants. Mean ± SD, n = 10. g The phenotype of mature-stage WT and OE-OsNAC103 plants. Bar = 5 cm. h The different internodes of mature WT and OE-OsNAC103 plants. Bar = 2 cm. i The internode lengths of mature WT and OE-OsNAC103 plants (from the top of the stem to the bottom). Mean ± SD, n = 13. The WT was used as a control for significance difference analysis. *P < 0.05; **P < 0.01; ***P < 0.001; t test
Phenotypes of RNAi-OsNAC103 plants. a The relative expression level of OsNAC103 of 21-day-old WT and RNAi lines. Mean values ± SD, n = 3. b The phenotype of 21-day-old WT and RNAi plants. Bar = 5 cm. c The shoot length of 21-day-old WT and RNAi plants. Mean ± SD, n = 15. d The leaf length of 21-day-old WT and RNAi plants. Mean ± SD, n = 15. e The leaf sheath length of 21-day-old WT and RNAi plants. Mean ± SD, n = 15. f The plant height of mature WT and RNAi plants. Mean ± SD, n = 8. g The phenotype of mature-stage WT. Bar = 10 cm. h The phenotype of mature-stage RNAi plants. Bar = 10 cm. i The different internodes of mature WT and RNAi lines. Bar = 3 cm. j The internode lengths of the WT and RNAi plants (from the top of the stem to the bottom). Mean ± SD, n = 8. The WT was used as a control for significance difference analysis. *P < 0.05; ns, no significant difference, t test
OsNAC103 regulates gibberellin metabolism but does not affect cell length
A previous study showed that gibberellins are the main factors determining plant height (Salas Fernandez et al. 2009). Therefore, we investigated the relationship between the dwarf phenotype and gibberellins. The results showed that the expression of the gibberellin biosynthesis-related genes OsKS, OsKO2, and OsKAO were suppressed in OE-OsNAC103 plants, but increased in RNAi lines (Fig. 6a). These results suggested that gibberellin synthesis was negatively regulated. We examined the growth of the WT, RNAi lines, and OE-OsNAC103 plants treated with PAC and exogenous GA3. After 10 days of 10 μM PAC treatment, both the WT and transgenic lines were significantly inhibited, and the difference in plant height was dramatically reduced. The plant height increased (compared to normal growth) after the application of 1 μM GA3 and 10 μM GA3. In contrast, the plant height of the RNAi lines was similar to that of the WT except in plants treated with 10 μM GA3. However, plant height remained lower in the overexpressing plants than in WT plants and RNAi lines (Fig. 6b, c). These results indicated that the increased expression of OsNAC103 decreases sensitivity to gibberellins.
OsNAC103 regulates gibberellin metabolism and cell numbers. a The relative expression level of genes related to gibberellin synthesis in WT, RNAi, and OE-OsNAC103 plants. Mean values ± SD, n = 3. b The phenotype of WT, RNAi, and OE-OsNAC103 plants incubated in 1/2 MS medium or medium-containing PAC, GA3 for 10 days. Bar = 5 cm. c The shoot length of WT, RNAi, and OE-OsNAC103 plants incubated in 1/2 MS medium or medium-containing PAC, GA3 for 10 days. Mean ± SD, n = 10. d The epidermal cells in the second leaf sheath of 21-day-old seedlings. Bar = 50 μm. e The cell length of WT, RNAi, and OE-OsNAC103 plants. Mean ± SD. Every line has at least 300 cells. f Estimation of cell numbers in the second leaf sheath of 21-day-old seedlings
Gibberellins exert effects on growth by altering cell length (Thingnaes et al. 2003; Chen et al. 2015). Accordingly, we compared the cell lengths of the second leaf sheaths, and the statistical analysis showed no significant differences (Fig. 6d, e). This suggests that dwarfing is not caused by changes in cell length. Cell numbers were estimated using the ratio of leaf sheath length to cell length. The results showed that the overexpression of OsNAC103 significantly reduces the number of cells (Fig. 6f). The decrease in cell numbers is the main cause of dwarfing.
OsNAC103 overexpression results in a reduction in cytokinin content
To confirm whether phytohormone levels were altered in OsNAC103-related dwarf plants, endogenous phytohormone levels were measured. This finding indicated a significant decrease in iP levels in OE-OsNAC103 plants, which was only 40% of that in WT plants. In addition, there were no notable differences in the levels of other cytokinin components and other phytohormones (Fig. 7a, Fig. S3). Dark-induced stress experiments showed that cytokinins can delay chlorophyll degradation in mature green leaves (Zhang et al. 2021). Therefore, we conducted dark induction experiments on WT and transgenic plants. After 2 days of dark treatment, the leaves of OE-OsNAC103 plants showed an early yellowing trend. On the third day of treatment, the WT and RNAi strains showed delayed leaf yellowing. The above results indicate that the decrease in cytokinin content was more likely to exhibit a yellowing phenotype (Fig. 7b). OsNAC103 can affect plant growth by reducing the cytokinin content.
Overexpression of OsNAC103 decreased the cytokinin content and sensitivity to iP. a The comparison of WT and OE-OsNAC103 plants’ cytokinin content. Mean values ± SD, n = 3. b Observation of dark-induced leaf yellowing phenotypes in WT, RNAi, and OE-OsNAC103 plants. c The relative expression level of OsNAC103 under iP (100 µM) treatment in WT plants. Mean ± SD, n = 3. d Phenotypes of WT, RNAi, and OE-OsNAC103 plants treated with different concentrations of iP for 10 days. Bar = 5 cm. e The relative shoot length of WT, RNAi, and OE-OsNAC103 plants treated with different concentrations of iP for 10 days. Mean ± SD, n = 8. The WT was used as a control for significance difference analysis. **P < 0.01; ***P < 0.001; ns, no significant difference, t test
Overexpression of OsNAC103 decreased plant susceptibility to iP
In plants, iP is considered as one of the primary active forms of cytokinins. To further explore the relationship between OsNAC103 and iP, the expression level of OsNAC103 in plants treated with exogenous cytokinin iP was tested. As shown in Fig. 7c, upon exogenous iP treatment, the OsNAC103 transcript level was significantly increased in the WT plants, indicating that OsNAC103 participates in the regulation of iP response.
Then, the WT, RNAi, and OE-OsNAC103 strains were treated with 1 µM iP and 10 µM iP. Compared with seedlings grown on 1/2 MS medium, the growth of plants was suppressed under 1 µM iP treatment, and the relative height of the RNAi plants was significantly lower than that of the WT. The relative plant height of the OE-OsNAC103 plants was considerably higher than that of the WT. When treated with 10 μM iP, plants of all lines showed restricted growth. The RNAi strain exhibited a higher degree of inhibition than the OE-OsNAC103 strain. The relative plant height of OE-OsNAC103 was significantly higher than that of the WT and RNAi plants (Fig. 7d, e). Similarly, when the osnac103 mutant was treated with iP, the phenotype of the osnac103 plants was consistent with that of the RNAi lines (Fig. S4a). These results indicated that OE-OsNAC103 plants were not sensitive to iP. Overexpression of OsNAC103 not only reduced the cytokinin content but also reduced the sensitivity to cytokinins.
OsNAC103 regulates cytokinin synthesis, degradation, and signal transduction
Cytokinin levels are determined by the balance between their synthesis and metabolism. Because the iP content was decreased in OE-OsNAC103, we measured the expression levels of genes involved in cytokinin metabolism. As shown in Fig. 8a, the cytokinin biosynthesis-related genes OsIPT3 and OsIPT8 were activated in the RNAi plants but were inhibited in the overexpression plants. In contrast, OsCKX4 and OsCKX5 showed a more significant upregulation trend in OE-OsNAC103 plants. Previous studies have shown that the CKX enzymes can irreversibly degrade iP (Zurcher and Muller 2016), and the enhanced CKX function promotes cytokinin degradation, leading to an increase in inactive cytokinins, thereby inhibiting plant growth and development. These results indicated that OsNAC103 reduces bioactive cytokinin accumulation by promoting cytokinin degradation and inhibiting cytokinin synthesis.
OsNAC103 regulates cytokinin synthesis, degradation, and signal transduction. a The relative expression level of genes related to cytokinin synthesis, degradation, and signaling response in WT, RNAi, and OE-OsNAC103 plants. Mean values ± SD, n = 3. b The relative expression level of OsNAC103 in the WT under 6-BA (100 µM) treatment. Mean ± SD, n = 3. c Phenotypes of WT, RNAi, and OE-OsNAC103 plants on 1/2 MS alone or treated with different concentrations of 6-BA for 10 days. Bar = 5 cm. d The relative shoot length of WT, RNAi, and OE-OsNAC103 plants on 1/2 MS alone or treated with different concentrations of 6-BA for 10 days. Mean ± SD, n = 8. The WT was used as a control for significance difference analysis. *P < 0.05; **P < 0.01; ***P < 0.001; t test
To validate the connection between dwarfing and cytokinin, we investigated the response of OsNAC103 transcripts to 100 μM 6-BA in WT leaves and found that the expression of OsNAC103 was downregulated within 1 h (Fig. 8b). The sensitivity of the WT and transgenic lines to 6-BA was evaluated with 1 μM and 10 μM 6-BA. After treatment for 10 days, the plant height of both the WT and transgenic lines was restricted in comparison to that of plants cultivated under standard growth conditions. The inhibition rate was higher in the RNAi lines and osnac103 plants than in the overexpression lines (Fig. 8c, d, Fig. S4b). These results indicate that the OE-OsNAC103 plants were hyposensitive to 6-BA. To explore whether cytokinin signaling is affected, cytokinin response genes type-A and type-B RRs were analyzed. Type-A RRs negatively regulate cytokinin signaling, whereas type-B RRs are activators (To and Kieber 2008). The results showed that OsARR6 was suppressed in the RNAi lines but increased in the overexpression lines. OsBRR1 was downregulated (Fig. 8a). Thus, OsNAC103 negatively regulates cytokinin biosynthesis and signaling pathways in rice.
OsNAC103 represses cell cycle progression and OsCYCP2;1 may act as a target gene
Cytokinins can accelerate cell division, and there is a positive correlation between their levels and cell division (Yang et al. 2002). For example, rice G-protein γ subunit (RGG1) inhibits cell division by significantly reducing cytokinin biosynthesis, ultimately reducing plant height and panicle elongation (Tao et al. 2020). In OsCKX3 overexpressing plants, the decrease in the expression of the cyclin-related gene CYCP4;1 reduced cell proliferation (Huang et al. 2022). Because cell numbers were reduced in dwarf plants, we speculated that cell division might be inhibited in OE-OsNAC103 plants. qRT-PCR analysis showed that OsCYCP1;1, OsCYCP2;1, and OsCYCB2;2 were downregulated in the overexpression lines (Fig. 9a). Physical interactions between AtCYCP2;1 and CDKs affect the G2/M transition. Furthermore, atcycp2;1 mutant plants exhibit seedling growth arrest (Torres Acosta et al. 2004; Peng et al. 2014). The findings indicated that OsNAC103 hinders the regular progression of the cell cycle and restrains cell division, leading to a dwarf phenotype.
OsNAC103 affects cell cycle progression genes. a The relative expression levels of cell cycle genes in WT, RNAi, and OE-OsNAC103 plants. Mean values ± SD, n = 3. b Transactivation activity of OsNAC103 on the promoter of OsCYCP2;1 was tested by dual-luciferase assay. Mean ± SD, n = 5. c A schematic diagram of the promoter of OsCYCP2;1 and DNA-binding activities of OsNAC103 proteins on the CACG motifs of OsCYCP2;1 was tested by EMSA. The WT was used as a control for significance difference analysis. *P < 0.05; **P < 0.01; ***P < 0.001; t test
OsNAC103 is a transcription factor; therefore, it is important to identify its downstream regulatory target genes. We performed dual-luciferase experiments to determine whether OsNAC103 regulates the promoter activity of cytokinin-related genes and cell cycle-related genes. It has been found that OsNAC103 can regulate the promoter activity of OsCYCP2;1. Compared to the control, the overexpression of OsNAC103 led to a significant increase in the activity of the LUC reporter (Fig. 9b).
The CACG motif is a core-binding site for NAC transcription factors (He et al. 2015; Tang et al. 2019). There was more than one CACG motif in the OsCYCP2;1 promoter (Fig. 9c). Therefore, we asked whether OsNAC103 could bind to the promoter of OsCYCP2;1. Next, the interaction between OsNAC103 protein and proOsCYCP2;1 was tested by performing EMSA for further verification with purified OsNAC103 protein. These experiments confirmed the above results, showing that the OsNAC103 protein binds to the CACG motif in the promoter of OsCYCP2;1 (Fig. 9c). Together, these data confirmed that OsCYCP2;1 as a target of OsNAC103.
OsNAC103 affects plant development by regulating KNOX family genes
The above findings indicated that both genes associated with cytokinins and gibberellins were impacted. We speculated that OsNAC103 had disrupted the crosstalk and balance of phytohormones. Therefore, KNOX genes involved in gibberellin and cytokinin crosstalk were examined. The results showed that OSH71 exhibited a significant increase in overexpression plants but was downregulated in RNAi lines, indicating that the balance was regulated by the expression of OsNAC103 (Fig. 10a). As OSH71 also plays a significant role in plant morphogenesis, OsNAC103 may also affect plant growth by regulating OSH71. However, OsNAC103 did not directly regulate the expression of OSH71 (Fig. S5).
OsNAC103 is involved in the homeostasis regulation of plant hormones. a The relative expression level of OSH71 in WT, RNAi, and OE-OsNAC103 plants. Mean values ± SD, n = 3. b Working model for OsNAC103 regulating plant height. The circles and rectangles marked with question marks represent unknown proteins and promoter elements, respectively. The WT was used as a control for difference significance analysis. *P < 0.05; **P < 0.01; ***P < 0.001; t test
Discussion
The NAC transcription factor family is engaged in diverse processes of plant growth and development. OsNAC103 overexpression reduced plant height, but there was no significant variation observed in terms of plant height among the WT, RNAi, and osnac103 mutants. These indicated that OsNAC103 negatively regulates rice plant height.
NAC family genes can affect the regulation of various phytohormones (Fujita et al. 2004; Bu et al. 2008; Puranik et al. 2012; Mao et al. 2020). Cytokinins play an important role in regulating cell proliferation by positively influencing cell division (Schaller et al. 2014). The reduction in cytokinin levels also inhibits growth (Duan et al. 2019). CKX is the only enzyme that irreversibly degrades cytokinins and nucleosides. In Arabidopsis thaliana, the enzymatic activity of CKX4 is enhanced in 35S::AtCKX4 transgenic plants, leading to a reduction in cytokinin levels (Werner et al. 2003). OsCKX4-overexpressing plants showed poor agronomic traits, such as dwarfing and late flowering (Gao et al. 2014; Chen et al. 2019), similar to the phenotype of OE-OsNAC103 plants. The activation of CKXs led to a reduction in iP content, while the levels of other active cytokinins remained relatively unchanged compared to those in the WT. The upregulation of OsARR6 and the repression of OsBRR1 in plants overexpressing OsNAC103 inhibited cytokinin signaling. Although the expression of OsIPTs increased and that of OsCKXs decreased in the RNAi lines, OsBRR1 was also downregulated, impairing the cytokinin response. This may be one of the reasons why there was no significant difference in plant height between the RNAi lines and WT.
Cytokinins regulate the G1/S and G2/M transitions (Schaller et al. 2014). The decreased expression level of OsCYCB2.2 also indicated that G2/M was indeed affected. These data indicated that OsNAC103 negatively regulates cytokinin synthesis, weakens the cytokinin response, and suppresses the expression of genes related to the cell cycle, ultimately leading to a dwarfing phenotype. However, the expression level of OsCYCP2;1 in OE-OsNAC103 plants was inconsistent with the transactivation activities of OsNAC103 in rice protoplasts. Similar expression patterns have been reported for OsNAC2 (Chen et al. 2015; Mao et al. 2018) and OsNAC24 (Jin et al. 2023) negatively regulating downstream genes. OsNAC2 may not be the only factor that regulates OsKO2 and OsCOX11 expression. Studies have shown that interactions between transcription factors and cofactors may alter DNA-binding affinity and convert activators into repressors (Ren et al. 2021; Li et al. 2023). Based on the previous studies, we deduced that OsNAC103 may cooperate with other proteins to regulate OsCYCP2;1 expression in plant cells via a more complex mechanism. Furthermore, OsNAC103 may regulate gene expression in a non-linear way with enhancers or silencers, and cofactors on the promoter sequence of OsCYCP2;1. The relationships between OsNAC103, OsCKXs or other proteins, and OsCYCP2;1 warrant further exploration. In addition, AtCYCP1;1 can interact with AtCDKA1 (Torres Acosta et al. 2004), and OsCYCP4 competes with other typical OsCYCs to bind OsCDKs under phosphate starvation (Xu et al. 2020). The interaction between OsCYCP2;1 and OsCDKs, and the effect of OsNAC103 on protein interaction need further experimental investigations.
Genes associated with gibberellin synthesis were affected in OsNAC103 overexpression and RNAi plants (Fig. 6a). Changes in intermediate levels of the gibberellin-synthesis pathway are unknown. However, the content of GA3, one of the final active gibberellin components, did not vary significantly compared to that in the WT (Fig. S3). Studies have shown that changes in intermediates can also affect the gene expression level regulated by them (Su et al. 2021). In addition, gibberellins also affect the cell cycle (Nagai et al. 2020). In the NAC transcription factor family, ANAC019 and ANAC055 participate in the crosstalk between abscisic acid and methyl jasmonate in plant defense response (Bu et al. 2008; Jiang et al. 2009). OsNAC016 plays a role in maintaining the balance between abscisic acid and brassinosteroids (Wu et al. 2022). Therefore, we believe that OsNAC103 also affects communication between cytokinins and gibberellins.
It can be confirmed that the levels of cytokinins and gibberellin-related genes and OSH71, involved in plant hormone regulation, were altered in the OsNAC103 transgenic lines. The phenotypes of plants overexpressing KNOX class I genes were similar to those of plants with excessive cytokinin content (Ori et al. 1999). OSH71 expression was upregulated in OE-OsNAC103 plants (Fig. 10a), while the cytokinin content was decreased (Fig. 7a). This suggests that gibberellin–cytokinin crosstalk was affected. In rice, OSH71 is expressed at a low level in mature and young leaves, but at a relatively high level during panicle and seed development. Ectopic overexpression of OSH71 causes abnormal plant development and affects the communication between cytokinins and gibberellins. OsNAC103 may also regulate plant growth by modulating OSH71 expression. However, the specific regulation mode needs further exploration.
Members of the NAC gene family have functional redundancy. osnac20–osnac26 double mutants showed decreased storage proteins in the grain, while the single-gene mutant showed no phenotype (Wang et al. 2020). OsNAC20 and OsNAC26 co-regulate the synthesis of starch in grains. The RNAi lines and mutant of OsNAC103 were comparable to WT plants in terms of plant height. The lower transcripts’ accumulation of OsNAC103 is sufficient to meet normal requirements. On the contrary, the overexpression of OsNAC103 may have a strong effect on plant development. OsNAC58 is the homologous gene of OsNAC103 in rice. OsNAC58 overexpression plants showed the phenotype of leaf senescence after entering the tillering stage (Liang et al. 2014). Overexpression of OsNAC103 also promoted leaf yellowing under dark conditions. These suggest a functional similarity between OsNAC103 and OsNAC58. The transgenic plants that OsNAC103 fused with a chimeric dominant repressor (OsNAC103-SRDX) are necessary to generate to explore the functional redundancy problem.
Based on these findings, we propose a working model of OsNAC103 that regulating plant height in rice (Fig. 10b). When OsNAC103 was overexpressed, the expression of cytokinin synthetase IPT genes was downregulated, and the upregulation of CKX genes enhanced the degradation of cytokinins, eventually leading to a reduction in cytokinin content. Upregulated expression of the response factor OsARR6 inhibits signal transduction. These changes also suppressed the expression of cyclins. In addition, OsNAC103 regulates gene expression by influencing the promoter activity of OsCYCP2;1. On the other hand, overexpression of OSH71 affected the homeostasis of cytokinins and gibberellins and the normal growth of plants. These eventually lead to a dwarfing phenotype in plants. Further research is required to enhance the model.
Data availability
Data are contained within the article or supplementary material.
Abbreviations
- 6-BA:
-
N6-benzyladenine
- CDK:
-
Cyclin-dependent kinase
- CKX:
-
Cytokinin oxidase/dehydrogenase
- iP:
-
N6-Δ2-isopentenyladenine
- IPT:
-
Isopentenyl transferase
- PAC:
-
Paclobutrazol
References
Bu Q, Jiang H, Li CB, Zhai Q, Zhang J, Wu X, Sun J, Xie Q, Li C (2008) Role of the Arabidopsis thaliana NAC transcription factors ANAC019 and ANAC055 in regulating jasmonic acid-signaled defense responses. Cell Res 18(7):756–767. https://doi.org/10.1038/cr.2008.53
Camut L, Gallova B, Jilli L, Sirlin-Josserand M, Carrera E, Sakvarelidze-Achard L, Ruffel S, Krouk G, Thomas SG, Hedden P, Phillips AL, Daviere JM, Achard P (2021) Nitrate signaling promotes plant growth by upregulating gibberellin biosynthesis and destabilization of DELLA proteins. Curr Biol 31(22):4971–4982, e4974. https://doi.org/10.1016/j.cub.2021.09.024
Chen S, Songkumarn P, Liu J, Wang GL (2009) A versatile zero background T-vector system for gene cloning and functional genomics. Plant Physiol 150(3):1111–1121. https://doi.org/10.1104/pp.109.137125
Chen X, Lu S, Wang Y, Zhang X, Lv B, Luo L, Xi D, Shen J, Ma H, Ming F (2015) OsNAC2 encoding a NAC transcription factor that affects plant height through mediating the gibberellic acid pathway in rice. Plant J 82(2):302–314. https://doi.org/10.1111/tpj.12819
Chen L, Zhao J, Song J, Jameson PE (2019) Cytokinin dehydrogenase: a genetic target for yield improvement in wheat. Plant Biotechnol J 18(3):614–630. https://doi.org/10.1111/pbi.13305
Chen Y, Sun S, Wang X (2020) The epidermis-specific cyclin CYCP3;1 is involved in the excess brassinosteroid signaling-inhibited root meristem cell division. J Integr Plant Biol 62(11):1674–1687. https://doi.org/10.1111/jipb.12975
Chevalier C (2008) Cell cycle control and fruit development. In: Inzé D (ed) Annual plant reviews, vol. 32: cell cycle control and fruit development. https://doi.org/10.1002/9780470988923.ch12
de Saint GA, Ligerot Y, Dun EA, Pillot JP, Ross JJ, Beveridge CA, Rameau C (2013) Strigolactones stimulate internode elongation independently of gibberellins. Plant Physiol 163(2):1012–1025. https://doi.org/10.1104/pp.113.220541
Deng M, Hu B, Xu L, Liu Y, Wang F, Zhao H, Wei X, Wang J, Yi K (2014) OsCYCP1;1, a PHO80 homologous protein, negatively regulates phosphate starvation signaling in the roots of rice (Oryza sativa L.). Plant Mol Biol 86(6):655–669. https://doi.org/10.1007/s11103-014-0254-1
Duan J, Yu H, Yuan K, Liao Z, Meng X, Jing Y, Liu G, Chu J, Li J (2019) Strigolactone promotes cytokinin degradation through transcriptional activation of CYTOKININ OXIDASE/DEHYDROGENASE 9 in rice. Proc Natl Acad Sci USA 116(28):14319–14324. https://doi.org/10.1073/pnas.1810980116
Fan K, Shen H, Bibi N, Li F, Yuan S, Wang M, Wang X (2015) Molecular evolution and species-specific expansion of the NAP members in plants. J Integr Plant Biol 57(8):673–687. https://doi.org/10.1111/jipb.12344
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran LSP, Yamaguchi-Shinozaki K, Shinozaki K (2004) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39(6):863–876
Gao S, Fang J, Xu F, Wang W, Sun X, Chu J, Cai B, Feng Y, Chu C (2014) CYTOKININ OXIDASE/DEHYDROGENASE4 integrates cytokinin and auxin signaling to control rice crown root formation. Plant Physiol 165(3):1035–1046. https://doi.org/10.1104/pp.114.238584
Ha S, Vankova R, Yamaguchi-Shinozaki K, Shinozaki K, Tran L-SP (2012) Cytokinins: metabolism and function in plant adaptation to environmental stresses. Trends Plant Sci 17(3):172–179
Haga N, Kobayashi K, Suzuki T, Maeo K, Kubo M, Ohtani M, Mitsuda N, Demura T, Nakamura K, Jurgens G, Ito M (2011) Mutations in MYB3R1 and MYB3R4 cause pleiotropic developmental defects and preferential down-regulation of multiple G2/M-specific genes in Arabidopsis. Plant Physiol 157(2):706–717. https://doi.org/10.1104/pp.111.180836
He X, Qu B, Li W, Zhao X, Teng W, Ma W, Ren Y, Li B, Li Z, Tong Y (2015) The nitrate-inducible NAC transcription factor TaNAC2-5A controls nitrate response and increases wheat yield. Plant Physiol 169(3):1991–2005. https://doi.org/10.1104/pp.15.00568
Hirose N, Makita N, Kojima M, Kamada-Nobusada T, Sakakibara H (2007) Overexpression of a type-A response regulator alters rice morphology and cytokinin metabolism. Plant Cell Physiol 48(3):523–539. https://doi.org/10.1093/pcp/pcm022
Huang P, Zhao J, Hong J, Zhu B, Xia S, Engao Z, Han P, Zhang K (2022) Cytokinins regulate rice lamina joint development and leaf angle. Plant Physiol 191:56–69. https://doi.org/10.1093/plphys/kiac401
Hwang I, Sheen J, Muller B (2012) Cytokinin signaling networks. Annu Rev Plant Biol 63:353–380. https://doi.org/10.1146/annurev-arplant-042811-105503
Itoh H, Ueguchi-Tanaka M, Sakamoto T, Kayano T, Tanaka H, Ashikari M, Matsuoka M (2002) Modification of rice plant height by suppressing the height-controlling gene, D18, in rice. Breed Sci 52(3):215–218
Jasinski S, Piazza P, Craft J, Hay A, Woolley L, Rieu I, Phillips A, Hedden P, Tsiantis M (2005) KNOX action in Arabidopsis is mediated by coordinate regulation of cytokinin and gibberellin activities. Curr Biol 15(17):1560–1565
Jeong J, Kim Y, Baek K, Jung H, Ha S, Choi Y, Kim M, Reuzeau C, Kim J (2010) Root-specific expression of OsNAC10 improves drought tolerance and grain yield in rice under field drought conditions. Plant Physiol 153(1):185–197
Jiang H, Li H, Bu Q, Li C (2009) The RHA2a-interacting proteins ANAC019 and ANAC055 may play a dual role in regulating ABA response and jasmonate response. Plant Signal Behav 4(5):464–466
Jiang Y, Cheng K, Zhao X, Ouyang Y (2018) Isolation and transformation of rice protoplasts. Bio-Protoc 101:e1010125. https://doi.org/10.21769/BioProtoc.1010125
Jin S, Xu L, Leng Y, Zhang M, Yang Q, Wang S, Jia S, Song T, Wang R, Tao T, Liu Q, Cai X, Gao J (2023) The OsNAC24-OsNAP protein complex activates OsGBSSI and OsSBEI expression to fine-tune starch biosynthesis in rice endosperm. Plant Biotechnol J 21(11):2224–2240. https://doi.org/10.1111/pbi.14124
Kudo T, Kiba T, Sakakibara H (2010) Metabolism and long-distance translocation of cytokinins. J Integr Plant Biol 52(1):53–60. https://doi.org/10.1111/j.1744-7909.2010.00898.x
La H, Li J, Ji Z, Cheng Y, Li X, Jiang S, Venkatesh PN, Ramachandran S (2006) Genome-wide analysis of cyclin family in rice (Oryza sativa L.). Mol Genet Genomics 275(4):374–386. https://doi.org/10.1007/s00438-005-0093-5
Lee D, Chung P, Jeong J, Jang G, Bang S, Jung H, Kim Y, Ha S, Choi Y, Kim J (2017) The rice OsNAC6 transcription factor orchestrates multiple molecular mechanisms involving root structural adaptions and nicotianamine biosynthesis for drought tolerance. Plant Biotechnol J 15(6):754–764
Li J, Qiu J, Zeng Q, Yi Z, Zhang N, Xu S, Jin J, Dong Z, Chen L, Huang W (2023) OsTOC1 plays dual roles in the regulation of plant circadian clock by functioning as a direct transcription activator or repressor. Cell Rep 42(7):112765. https://doi.org/10.1016/j.celrep.2023.112765
Liang C, Wang Y, Zhu Y, Tang J, Hu B, Liu L, Ou S, Wu H, Sun X, Chu J, Chu C (2014) OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. Proc Natl Acad Sci USA 111(27):10013–10018. https://doi.org/10.1073/pnas.1321568111
Liu Y, Liu Y, He Y, Yan Y, Yu X, Ali M, Pan C, Lu G (2023) Cytokinin-inducible response regulator SlRR6 regulates plant height through gibberellin and auxin pathways in tomato. J Exp Bot 74:4471–4488. https://doi.org/10.1093/jxb/erad159
Lo S, Yang S, Chen K, Hsing Y, Zeevaart JA, Chen L, Yu S (2008) A novel class of gibberellin 2-oxidases control semidwarfism, tillering, and root development in rice. Plant Cell 20(10):2603–2618. https://doi.org/10.1105/tpc.108.060913
Lomin SN, Krivosheev DM, Steklov MY, Arkhipov DV, Osolodkin DI, Schmulling T, Romanov GA (2015) Plant membrane assays with cytokinin receptors underpin the unique role of free cytokinin bases as biologically active ligands. J Exp Bot 66(7):1851–1863. https://doi.org/10.1093/jxb/eru522
Mao C, Ding J, Zhang B, Xi D, Ming F (2018) OsNAC2 positively affects salt-induced cell death and binds to the OsAP37 and OsCOX11 promoters. Plant J 94(3):454–468. https://doi.org/10.1111/tpj.13867
Mao C, He J, Liu L, Deng Q, Yao X, Liu C, Qiao Y, Li P, Ming F (2020) OsNAC2 integrates auxin and cytokinin pathways to modulate rice root development. Plant Biotechnol J 18(2):429–442
Margis-Pinheiro M, Zhou X, Zhu Q, Dennis ES, Upadhyaya NM (2005) Isolation and characterization of a Ds-tagged rice (Oryza sativa L.) GA-responsive dwarf mutant defective in an early step of the gibberellin biosynthesis pathway. Plant Cell Rep 23(12):819–833. https://doi.org/10.1007/s00299-004-0896-6
Miyawaki K, Tarkowski P, Matsumoto-Kitano M, Kato T, Sato S, Tarkowska D, Tabata S, Sandberg G, Kakimoto T (2006) Roles of Arabidopsis ATP/ADP isopentenyltransferases and tRNA isopentenyltransferases in cytokinin biosynthesis. Proc Natl Acad Sci USA 103(44):16598–16603
Nagai K, Mori Y, Ishikawa S, Furuta T, Gamuyao R, Niimi Y, Hobo T, Fukuda M, Kojima M, Takebayashi Y, Fukushima A, Himuro Y, Kobayashi M, Ackley W, Hisano H, Sato K, Yoshida A, Wu J, Sakakibara H, Sato Y, Tsuji H, Akagi T, Ashikari M (2020) Antagonistic regulation of the gibberellic acid response during stem growth in rice. Nature 584(7819):109–114. https://doi.org/10.1038/s41586-020-2501-8
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465(1–2):30–44. https://doi.org/10.1016/j.gene.2010.06.008
Nuruzzaman M, Sharoni AM, Satoh K, Moumeni A, Venuprasad R, Serraj R, Kumar A, Leung H, Attia K, Kikuchi S (2012) Comprehensive gene expression analysis of the NAC gene family under normal growth conditions, hormone treatment, and drought stress conditions in rice using near-isogenic lines (NILs) generated from crossing Aday Selection (drought tolerant) and IR64. Mol Genet Genomics 287(5):389–410. https://doi.org/10.1007/s00438-012-0686-8
Nuruzzaman M, Sharoni AM, Satoh K, Karim MR, Harikrishna JA, Shimizu T, Sasaya T, Omura T, Haque MA, Hasan SM, Ahmad A, Kikuchi S (2015) NAC transcription factor family genes are differentially expressed in rice during infections with rice dwarf virus, rice black-streaked dwarf virus, rice grassy stunt virus, rice ragged stunt virus, and rice transitory yellowing virus. Front Plant Sci 6:676. https://doi.org/10.3389/fpls.2015.00676
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10(6):239–247
Ori N, Juarez MT, Jackson D, Yamaguchi J, Banowetz GM, Hake S (1999) Leaf senescence is delayed in tobacco plants expressing the maize homeobox gene knotted1 under the control of a senescence-activated promoter. Plant Cell 11(6):1073–1080
Peng L, Skylar A, Chang P, Bisova K, Wu X (2014) CYCP2;1 integrates genetic and nutritional information to promote meristem cell division in Arabidopsis. Dev Biol 393(1):160–170. https://doi.org/10.1016/j.ydbio.2014.06.008
Polyn S, Willems A, De Veylder L (2015) Cell cycle entry, maintenance, and exit during plant development. Curr Opin Plant Biol 23:1–7. https://doi.org/10.1016/j.pbi.2014.09.012
Postma-Haarsma AD, Rueb S, Scarpella E, Den Besten W, Hoge JHC, Meijer AH (2002) Developmental regulation and downstream effects of the knox class homeobox genes Oskn2 and Oskn3 from rice. Plant Mol Biol 48(4):423–441
Puranik S, Sahu PP, Srivastava PS, Prasad M (2012) NAC proteins: regulation and role in stress tolerance. Trends Plant Sci 17(6):369–381
Qi F, Zhang F (2019) Cell cycle regulation in the plant response to stress. Front Plant Sci 10:1765. https://doi.org/10.3389/fpls.2019.01765
Ren Y, Huang Z, Jiang H, Wang Z, Wu F, Xiong Y, Yao J, Gibbs D (2021) A heat stress responsive NAC transcription factor heterodimer plays key roles in rice grain filling. J Exp Bot 72(8):2947–2964. https://doi.org/10.1093/jxb/erab027
Riou-Khamlichi C, Huntley R, Jacqmard A, Murray JA (1999) Cytokinin activation of Arabidopsis cell division through a D-type cyclin. Science 283(5407):1541–1544
Rosin FM, Hart JK, Horner HT, Davies PJ, Hannapel DJ (2003) Overexpression of a knotted-like homeobox gene of potato alters vegetative development by decreasing gibberellin accumulation. Plant Physiol 132(1):106–117. https://doi.org/10.1104/pp.102.015560
Sakamoto T, Miura K, Itoh H, Tatsumi T, Ueguchi-Tanaka M, Ishiyama K, Kobayashi M, Agrawal GK, Takeda S, Abe K, Miyao A, Hirochika H, Kitano H, Ashikari M, Matsuoka M (2004) An overview of gibberellin metabolism enzyme genes and their related mutants in rice. Plant Physiol 134(4):1642–1653. https://doi.org/10.1104/pp.103.033696
Salas Fernandez MG, Becraft PW, Yin Y, Lubberstedt T (2009) From dwarves to giants? Plant height manipulation for biomass yield. Trends Plant Sci 14(8):454–461. https://doi.org/10.1016/j.tplants.2009.06.005
Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Swapan D, Ishiyama K, Saito T, Kobayashi M, Khush GS (2002) A mutant gibberellin-synthesis gene in rice. Nature 416(6882):701–702
Sato Y, Sentoku N, Nagato Y, Matsuoka M (1998) Isolation and characterization of a rice homebox gene, OSH15. Plant Mol Biol 38(6):983–997
Schaller GE, Street IH, Kieber JJ (2014) Cytokinin and the cell cycle. Curr Opin Plant Biol 21:7–15. https://doi.org/10.1016/j.pbi.2014.05.015
Sheng M, Ma X, Wang J, Xue T, Li Z, Cao Y, Yu X, Zhang X, Wang Y, Xu W (2022) KNOX II transcription factor HOS59 functions in regulating rice grain size. Plant J 110:863–880
Su S, Hong J, Chen X, Zhang C, Chen M, Luo Z, Chang S, Bai S, Liang W, Liu Q, Zhang D (2021) Gibberellins orchestrate panicle architecture mediated by DELLA-KNOX signalling in rice. Plant Biotechnol J 19(11):2304–2318. https://doi.org/10.1111/pbi.13661
Sun S, Wang T, Wang L, Li X, Jia Y, Liu C, Huang X, Xie W, Wang X (2018) Natural selection of a GSK3 determines rice mesocotyl domestication by coordinating strigolactone and brassinosteroid signaling. Nat Commun 9(1):2523. https://doi.org/10.1038/s41467-018-04952-9
Tanabe S, Ashikari M, Fujioka S, Takatsuto S, Yoshida S, Yano M, Yoshimura A, Kitano H, Matsuoka M, Fujisawa Y, Kato H, Iwasaki Y (2005) A novel cytochrome P450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant, dwarf11, with reduced seed length. Plant Cell 17(3):776–790. https://doi.org/10.1105/tpc.104.024950
Tang W, Ye J, Yao X, Zhao P, Xuan W, Tian Y, Zhang Y, Xu S, An H, Chen G, Yu J, Wu W, Ge Y, Liu X, Li J, Zhang H, Zhao Y, Yang B, Jiang X, Peng C, Zhou C, Terzaghi W, Wang C, Wan J (2019) Genome-wide associated study identifies NAC42-activated nitrate transporter conferring high nitrogen use efficiency in rice. Nat Commun 10(1):5279. https://doi.org/10.1038/s41467-019-13187-1
Tao Y, Miao J, Wang J, Li W, Xu Y, Wang F, Jiang Y, Chen Z, Fan F, Xu M, Zhou Y, Liang G, Yang J (2020) RGG1, involved in the cytokinin regulatory pathway, controls grain size in rice. Rice (n y) 13(1):76. https://doi.org/10.1186/s12284-020-00436-x
Thingnaes E, Torre S, Ernstsen A, Moe R (2003) Day and night temperature responses in Arabidopsis: effects on gibberellin and auxin content, cell size, morphology and flowering time. Ann Bot 92(4):601–612. https://doi.org/10.1093/aob/mcg176
To JP, Kieber JJ (2008) Cytokinin signaling: two-components and more. Trends Plant Sci 13(2):85–92. https://doi.org/10.1016/j.tplants.2007.11.005
Torres Acosta JA, de Almeida EJ, Raes J, Magyar Z, De Groodt R, Inze D, De Veylder L (2004) Molecular characterization of Arabidopsis PHO80-like proteins, a novel class of CDKA;1-interacting cyclins. Cell Mol Life Sci 61(12):1485–1497. https://doi.org/10.1007/s00018-004-4057-4
Wang B, Smith SM, Li J (2018) Genetic regulation of shoot architecture. Annu Rev Plant Biol 69:437–468. https://doi.org/10.1146/annurev-arplant-042817-040422
Wang J, Chen Z, Zhang Q, Meng S, Wei C (2020) The NAC transcription factors OsNAC20 and OsNAC26 regulate starch and storage protein synthesis. Plant Physiol 184(4):1775–1791. https://doi.org/10.1104/pp.20.00984
Werner T, Motyka V, Strnad M, Schmülling T (2001) Regulation of plant growth by cytokinin. Proc Natl Acad Sci USA 98(18):10487–10492
Werner T, Motyka V, Laucou V, Smets R, Van Onckelen H, Schmulling T (2003) Cytokinin-deficient transgenic Arabidopsis plants show multiple developmental alterations indicating opposite functions of cytokinins in the regulation of shoot and root meristem activity. Plant Cell 15(11):2532–2550. https://doi.org/10.1105/tpc.014928
Wu Y, Wang Y, Mi X, Shan J, Li X, Xu J, Lin H (2016) The QTL GNP1 encodes GA20ox1, which increases grain number and yield by increasing cytokinin activity in rice panicle meristems. PLoS Genet 12(10):e1006386
Wu Q, Liu Y, Xie Z, Yu B, Sun Y, Huang J (2022) OsNAC016 regulates plant architecture and drought tolerance by interacting with the kinases GSK2 and SAPK8. Plant Physiol 189(3):1296–1313. https://doi.org/10.1093/plphys/kiac146
Xu L, Wang F, Li R, Deng M, Fu M, Teng H, Yi K (2020) OsCYCP4s coordinate phosphate starvation signaling with cell cycle progression in rice. J Integr Plant Biol 62(7):1017–1033. https://doi.org/10.1111/jipb.12885
Yamaguchi S (2008) Gibberellin metabolism and its regulation. Annu Rev Plant Biol 59:225–251. https://doi.org/10.1146/annurev.arplant.59.032607.092804
Yang J, Zhang J, Huang Z, Wang Z, Zhu Q, Liu L (2002) Correlation of cytokinin levels in the endosperms and roots with cell number and cell division activity during endosperm development in rice. Ann Bot 90(3):369–377
Zhang W, Peng K, Cui F, Wang D, Zhao J, Zhang Y, Yu N, Wang Y, Zeng D, Wang Y (2021) Cytokinin oxidase/dehydrogenase OsCKX11 coordinates source and sink relationship in rice by simultaneous regulation of leaf senescence and grain number. Plant Biotechnol J 19(2):335–350
Zhao C, Avci U, Grant EH, Haigler CH, Beers EP (2008) XND1, a member of the NAC domain family in Arabidopsis thaliana, negatively regulates lignocellulose synthesis and programmed cell death in xylem. Plant J 53(3):425–436. https://doi.org/10.1111/j.1365-313X.2007.03350.x
Zheng H, Zhang F, Wang S, Su Y, Ji X, Jiang P, Chen R, Hou S, Ding Y (2018) MLK1 and MLK2 coordinate RGA and CCA1 activity to regulate hypocotyl elongation in Arabidopsis thaliana. Plant Cell 30(1):67–82. https://doi.org/10.1105/tpc.17.00830
Zurcher E, Muller B (2016) Cytokinin synthesis, signaling, and function–advances and new insights. Int Rev Cell Mol Biol 324:1–38. https://doi.org/10.1016/bs.ircmb.2016.01.001
Acknowledgements
The authors express our gratitude to Xiaoling Guo (College of the Environment & Ecology, XMU) for her assistance in ensuring the smooth progress of the experiment.
Funding
This work was supported by grants from the Joint Fund of the National Natural Science Foundation of China, the Karst Science Research Center of Guizhou Province (U1812401), the National Key Research and Development Program of China (2016YFD0100604), and the Major Science and Technology Special Project of Fujian Province (2020NZ08016).
Author information
Authors and Affiliations
Contributions
LC, YC, and YL designed the experiment. YL and LZ undertook the main experimental work and YL wrote the manuscript. YL and YC analyzed the data. CG, MT, WL, XX, SC, YP, YY, and CL assisted with experiments and rice material planting. LC, YC, and CG helped revise the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
No competing or financial interests were declared.
Accession numbers
The RGAP database contains the information on the sequence data. The accession numbers: OsNAC103 (LOC_Os07g48450), OsKS (LOC_Os04g52230), OsKO2 (LOC_Os06g37364), OsKAO (LOC_Os06g02019), OsCYCB2.2 (LOC_Os06g51110), OsCYCP1;1 (LOC_Os05g33040), OsCYCP2;1 (LOC_Os04g53680), OsIPT3 (LOC_Os05g24660), OsIPT8 (LOC_Os01g49390), OsCKX4 (LOC_Os01g71310), OsCKX5 (LOC_Os01g56810), OsARR6 (LOC_Os04g57720), OsBRR1 (LOC_Os03g12350), OSH71 (LOC_Os05g03884), ACTIN1 (LOC_Os03g50885).
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file 1. Fig. S1
: RNA-seq results of the MBKBASE database to analyze the tissue expression pattern of OsNAC103. Fig. S2: Phenotypes of osnac103 mutant. a Schematic diagram indicating the gRNA location in the genomic region of OsNAC103 and the comparison of osnac103-c2 and osnac103-c6. OsNAC103: normal protein sequence. OsNAC103 insertion T: partial protein sequences of mutants inserted with T. OsNAC103 deletion CG: partial protein sequence of a mutant lacking CG. b The phenotype of 21-day-old WT (ZH11), osnac103-c2, and osnac103-c6 plants. Bar = 5 cm. c The plant height of 21-day-old WT (ZH11), osnac103-c2, and osnac103-c6 plants. Mean values ± SD, n = 9. d The phenotype of WT (ZH11) at the mature stage. Bar = 5 cm. e The phenotype of osnac103-c6 at the mature stage. Bar = 5 cm. f The plant height of mature WT (ZH11) and osnac103-c6 plants. Mean ± SD, n = 8. g The different internodes of mature WT (ZH11) and osnac103-c6 plants (from the top of the stem to the bottom). Bar = 5 cm. h The internode lengths of WT (ZH11) and osnac103-c6 plants (from the top of the stem to the bottom). Mean ± SD, n = 5. ns, no significant difference, t test. Fig. S3: Analysis of endogenous plant hormone contents in WT and OE plants. Mean values ± SD, n = 3. IAA (indole-3-acetic acid), ABA (cis-abscisic acid), CS (castasterone), JA (jasmonic acid), JA-ILE (jasmonic acid-isoleucine), cis-OPDA (cis–12-oxophytodienoic acid), SA (salicylic acid), ACC (1-aminocyclopropane-1-carboxylic acid). Fig. S4: Phenotypes of osnac103 treated with iP and 6-BA. a The phenotype and relative shoot length of WT, osnac103-c2, and osnac103-c6 plants incubated in 1/2 MS medium or treated with different concentrations of iP for 10 days. Bar = 5 cm. Mean values ± SD, n = 8. b The phenotype and relative shoot length of WT, osnac103-c2, and osnac103-c6 plants incubated in 1/2 MS medium or treated with different concentrations of 6-BA for 10 days. Bar = 5 cm. Mean ± SD, n = 8. The WT was used as a control for significance difference analysis.*P < 0.05; **P < 0.01; ***P < 0.001; ns, no significant difference, t test. Fig. S5: The regulation between OsNAC103 and OSH71 promoter was tested by yeast one-hybrid assay. Co-transformant with pB42AD-HY5 and proCOP1 was used as a positive control. Table S1 Primer sequences were used in the experiment. (PDF 733 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Y., Zhao, L., Guo, C. et al. OsNAC103, an NAC transcription factor negatively regulates plant height in rice. Planta 259, 35 (2024). https://doi.org/10.1007/s00425-023-04309-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-023-04309-7