Abstract
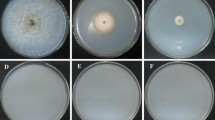
Agrobacterium tumefaciens-mediated transformation (ATMT) has been proven to be a powerful strategy for gene disruption in plants and fungi. Patterns associated with transferred DNA (T-DNA) integration in plants and yeast have been studied comprehensively, whereas no detailed analysis of T-DNA integration has been reported yet in filamentous fungi. Here, we reported the T-DNA insertion patterns in the genome of filamentous fungus Magnaporthe oryzae. Using ATMT, a T-DNA tagged population consisting of 6,179 transformants of M. oryzae was constructed. With thermal asymmetric interlaced-PCR (TAIL-PCR), 623 right border (RB) flanking sequences and 124 left border (LB) flanking sequences were generated. Analysis of these flanking sequences indicated a significant integration bias toward non-coding sequences, suggesting distribution of T-DNAs was not random. Comparing to T-DNA RB, LB was nicked inaccurately and truncated frequently during integration. Chromosomal rearrangements, such as deletion, inversion, and translocation, were associated with T-DNA integration in some transformants. Our data suggest that, comparing with plant cells, T-DNA integrates into this filamentous fungus with more precise and simpler patterns. Some phenotypic mutants were observed in our T-DNA tagged population, and these transformants will be very useful for functional genomics research of M. oryzae.





Similar content being viewed by others
References
Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, Gadrinab C, Heller C, Jeske A, Koesema E, Meyers CC, Parker H, Prednis L, Ansari Y, Choy N, Deen H, Geralt M, Hazari N, Hom E, Karnes M, Mulholland C, Ndubaku R, Schmidt I, Guzman P, Aguilar-Henonin L, Schmid M, Weigel D, Carter DE, Marchand T, Risseeuw E, Brogden D, Zeko A, Crosby WL, Berry CC, Ecker JR (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301:653–657
Brunaud V, Balzergue S, Dubreucq B, Aubourg S, Samson F, Chauvin S, Bechtold N, Cruaud C, DeRose R, Pelletier G, Lepiniec L, Caboche M, Lecharny A (2002) T-DNA integration into the Arabidopsis genome depends on sequences of pre-insertion sites. Embo Rep 3:1152–1157
Castle LA, Errampalli D, Atherton TL, Franzmann LH, Yoon ES, Meinke DW (1993) Genetic and molecular characterization of embryonic mutants identified following seed transformation in Arabidopsis. Mol Gen Genet 241:504–514
Chen S, Jin W, Wang M, Zhang F, Zhou J, Jia Q, Wu Y, Liu F, Wu P (2003) Distribution and characterization of over 1000 T-DNA tags in rice genome. Plant J 36:105–113
Chilton MD, Que Q (2003) Targeted integration of T-DNA into the tobacco genome at double-stranded breaks: new insights on the mechanism of T-DNA integration. Plant Physiol 133:956–965
Daley JM, Palmbos PL, Wu D, Wilson TE (2005) Non-homologous end joining in yeast. Annu Rev Genet 39:431–451
De Buck S, Jacobs A, Van Montagu M, Depicker A (1999) The DNA sequences of T-DNA junctions suggest that complex T-DNA loci are formed by a recombination process resembling T-DNA integration. Plant J 20:295–304
De Neve M, De Buck S, Jacobs A, Van Montagu M, Depicker A (1997) T-DNA integration patterns in co-transformed plant cells suggest that T-DNA repeats originate from co-integration of separate T-DNAs. Plant J 11:15–29
Dean RA, Talbot NJ, Ebbole DJ, Farman ML, Mitchell TK, Orbach MJ, Thon M, Kulkarni R, Xu JR, Pan H, Read ND, Lee YH, Carbone I, Brown D, Oh YY, Donofrio N, Jeong JS, Soanes DM, Djonovic S, Kolomiets E, Rehmeyer C, Li W, Harding M, Kim S, Lebrun MH, Bohnert H, Coughlan S, Butler J, Calvo S, Ma LJ, Nicol R, Purcell S, Nusbaum C, Galagan JE, Birren BW (2005) The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434:980–986
Forsbach A, Schubert D, Lechtenberg B, Gils M, Schmidt R (2003) A comprehensive characterization of single-copy T-DNA insertions in the Arabidopsis thaliana genome. Plant Mol Biol 52:161–176
Gheysen G, Villarroel R, Van Montagu M (1991) Illegitimate recombination in plants: a model for T-DNA integration. Genes Dev 5:287–297
Hill MA (1999) Radiation damage to DNA: the importance of track structure. Radiat Meas 31:15–23
Kim SR, Lee J, Jun SH, Park S, Kang HG, Kwon S, An G (2003) Transgene structures in T-DNA-inserted rice plants. Plant Mol Biol 52:761–773
Köhler F, Cardon G, Pöhlman M, Gill R, Schieder O (1989) Enhancement of transformation rates in higher plants by low-dose irradiation: are DNA repair systems involved in the incorporation of exogenous DNA into the plant genome? Plant Mol Biol 12:189–199
Kononov ME, Bassuner B, Gelvin SB (1997) Integration of T-DNA binary vector ‘backbone’ sequences into the tobacco genome: evidence for multiple complex patterns of integration. Plant J 11:945–957
Lacroix B, Tzfira T, Vainstein A, Citovsky V (2006) A case of promiscuity: Agrobacterium’s endless hunt for new partners. Trends Genet 22:29–37
Liu YG, Whittier RF (1995) Thermal asymmetric interlaced PCR: automatable amplification and sequencing of insert end fragments from P1 and YAC clones for chromosome walking. Genomics 25:674–681
Mayerhofer R, Koncz-Kalman Z, Nawrath C, Bakkeren G, Crameri A, Angelis K, Redei GP, Schell J, Hohn B, Koncz C (1991) T-DNA integration: a mode of illegitimate recombination in plants. Embo J 10:697–704
Michielse CB, Hooykaas PJ, van den Hondel CA, Ram AF (2005) Agrobacterium-mediated transformation as a tool for functional genomics in fungi. Curr Genet 48:1–17
Mullins ED, Chen X, Romaine P, Raina R, Geiser DM, Kang S (2001) Agrobacterium-mediated transformation of Fusarium oxysporum: an efficient tool for insertional mutagenesis and gene transfer. Phytopathology 91:173–180
Nacry P, Camilleri C, Courtial B, Caboche M, Bouchez D (1998) Major chromosomal rearrangements induced by T-DNA transformation in Arabidopsis. Genetics 149:641–650
Ohba T, Yoshioka Y, Machida C, Machida Y (1995) DNA rearrangement associated with the integration of T-DNA in tobacco: an example for multiple duplications of DNA around the integration target. Plant J 7:157–164
Oleinick NL, Balasubramaniam U, Xue L, Chiu S (1994) Nuclear structure and the microdistribution of radiation damage in DNA. Int J Radiat Biol 66:523–529
Rho HS, Kang S, Lee YH (2001) Agrobacterium tumefaciens-mediated transformation of the plant pathogenic fungus, Magnaporthe grisea. Mol Cells 12:407–411
Sallaud C, Meynard D, van Boxtel J, Gay C, Bes M, Brizard JP, Larmande P, Ortega D, Raynal M, Portefaix M, Ouwerkerk PB, Rueb S, Delseny M, Guiderdoni E (2003) Highly efficient production and characterization of T-DNA plants for rice (Oryza sativa L.) functional genomics. Theor Appl Genet 106:1396–1408
Sambrook J, Fritsch E, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York
Sha Y, Li S, Pei Z, Luo L, Tian Y, He C (2004) Generation and flanking sequence analysis of a rice T-DNA tagged population. Theor Appl Genet 108:306–314
Soanes DM, Skinner W, Keon J, Hargreaves J, Talbot NJ (2002) Genomics of phytopathogenic fungi and the development of bioinformatic resources. Mol Plant Microbe Interact 15:421–427
Stachel SE, Timmerman B, Zambryski P (1986) Generation of single-stranded T-DNA molecules during the initial stages of T-DNA transfer from Agrobacterium tumefaciens to plant cells. Nature 322:706–712
Talbot NJ, Salch YP, Ma M, Hamer JE (1993) Karyotypic variation within clonal lineages of the rice blast fungus, Magnaporthe grisea. Appl Environ Microbiol 59:585–593
Tinland B, Schoumacher F, Gloeckler V, Bravo-Angel AM, Hohn B (1995) The Agrobacterium tumefaciens virulence D2 protein is responsible for precise integration of T-DNA into the plant genome. Embo J 14:3585–3595
Tzfira T, Frankman LR, Vaidya M, Citovsky V (2003) Site-specific integration of Agrobacterium tumefaciens T-DNA via double-stranded intermediates. Plant Physiol 133:1011–1023
Tzfira T, Li J, Lacroix B, Citovsky V (2004) Agrobacterium T-DNA integration: molecules and models. Trends Genet 20:375–383
Valent B (1990) Rice blast as a model system for plant pathology. Phytopathology 80:33–36
Virts EL, Gelvin SB (1985) Analysis of transfer of tumor-inducing plasmids from Agrobacterium tumefaciens to petunia protoplasts. J Bacteriol 162:1030–1038
Wang K, Stachel SE, Timmerman B, Montagu MV, Zambryski PC (1987) Site-specific nick in the T-DNA border sequence as a result of Agrobacterium vir gene expression. Science 235:587–591
Wenck A, Czako M, Kanevski I, Marton L (1997) Frequent collinear long transfer of DNA inclusive of the whole binary vector during Agrobacterium-mediated transformation. Plant Mol Biol 34:913–922
Windels P, De Buck S, Van Bockstaele E, De Loose M, Depicker A (2003) T-DNA integration in Arabidopsis chromosomes. Presence and origin of filler DNA sequences. Plant Physiol 133:2061–2068
Xu JR, Peng YL, Dickman MB, Sharon A (2006) The dawn of fungal pathogen genomics. Annu Rev Phytopathol 44: 337–366
Yin Z, Wang G-L (2000) Evidence of multiple complex patterns of T-DNA integration into the rice genome. Theor Appl Genet 100:461–470
Zeigler RS, Leong SA, Teng PS (1994) Rice Blast Disease. CAB International, Wallingford
Acknowledgments
This project was supported by the Ministry of Science and Technology of China (Grant 2002BA711A15).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by K. Borkovich.
Guihua Li and Zhuangzhi Zhou contributed equally to the work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Li, G., Zhou, Z., Liu, G. et al. Characterization of T-DNA insertion patterns in the genome of rice blast fungus Magnaporthe oryzae . Curr Genet 51, 233–243 (2007). https://doi.org/10.1007/s00294-007-0122-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-007-0122-5