Abstract
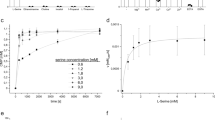
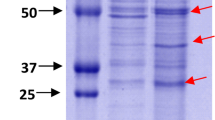
Phosphatidylserine synthase (Pss) is involved in the metabolic pathway in phospholipid synthesis in different organisms. In this study, Pss expression in Vibrio parahaemolyticus was verified through liquid chromatography tandem-mass spectrometry. To analyze the characteristics of Pss, the recombinant Pss was overexpressed and purified from Escherichia coli. The optimum temperature and pH of Pss were 40 °C and 8, respectively. When reacting with divalent metal, Pss activity decreased. In addition, Pss could not only use cytidine diphosphate diacylglycerol (CDP-DAG, 16:0), but also CDP-DAG (18:1) as a substrate to produce cytidine 5′‐monophosphate. Furthermore, the 3D structure of Pss was predicted, and the results revealed that histidine and lysine of the two HKD motifs were present in the catalytic site. Moreover, CDP-DAG (16:0) was docked with the Pss model. To investigate whether the two HKD motifs in Pss are important to its activity, site-directed mutagenesis of histidine was performed. The results revealed that the activities of both H131A and H352A were diminished. Little is known regarding the catalytic site of type I Pss. This is the first report on the biochemical characterization of Pss in V. parahaemolyticus.





Similar content being viewed by others
References
Okada M, Matsuzaki H, Shibuya I, Matsumoto K (1994) Cloning, sequencing, and expression in Escherichia coli of the Bacillus subtilis gene for phosphatidylserine synthase. J Bacteriol 176(24):7456–7461
Cassilly CD, Reynolds TB (2018) PS, It's complicated: the roles of phosphatidylserine and phosphatidylethanolamine in the pathogenesis of Candida albicans and other microbial pathogens. J Fungi (Basel). https://doi.org/10.3390/jof4010028
Li J, Wang X (2019) Phospholipase D and phosphatidic acid in plant immunity. Plant Sci 279:45–50. https://doi.org/10.1016/j.plantsci.2018.05.021
Selvy PE, Lavieri RR, Lindsley CW, Brown HA (2011) Phospholipase D: enzymology, functionality, and chemical modulation. Chem Rev 111(10):6064–6119. https://doi.org/10.1021/cr200296t
Xie Z, Ho WT, Exton JH (2000) Association of the N- and C-terminal domains of phospholipase D. Contribution of the conserved HKD motifs to the interaction and the requirement of the association for Ser/Thr phosphorylation of the enzyme. J Biol Chem 275(32):24962–24969. https://doi.org/10.1074/jbc.M909745199
Bullock WO, Fernandez JM, Short JM (1987) XL1-blue: a high efficiency plasmid transforming recA Escherichia coli strain with beta-galactosidase selection. Biotechniques 5:376–378
Simon R, Priefer U, Puhler A (1983) A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in gram negative bacteria. Nat Biotech 1(9):784–791
Philippe N, Alcaraz JP, Coursange E, Geiselmann J, Schneider D (2004) Improvement of pCVD442, a suicide plasmid for gene allele exchange in bacteria. Plasmid 51(3):246–255. https://doi.org/10.1016/j.plasmid.2004.02.003
Perkins DN, Pappin DJ, Creasy DM, Cottrell JS (1999) Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20(18):3551–3567. https://doi.org/10.1002/(SICI)1522-2683(19991201)20:18<3551::AID-ELPS3551>3.0.CO;2-2
Huang RY, Lee CY (2019) Molecular and functional evidence of phosphatidylserine synthase in Vibrio parahaemolyticus. Microbiol Immunol 63(3–4):119–129. https://doi.org/10.1111/1348-0421.12676
Dowhan W (1992) Phosphatidylserine synthase from Escherichia coli. Methods Enzymol 209:287–298
Kanfer J, Kennedy EP (1964) Metabolism and function of bacterial lipids. II. Biosynthesis of phospholipids in Escherichia coli. J Biol Chem 239:1720–1726
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46(W1):W296–W303. https://doi.org/10.1093/nar/gky427
Tian W, Chen C, Lei X, Zhao J, Liang J (2018) CASTp 3.0: computed atlas of surface topography of proteins. Nucleic Acids Res 46(W1):W363–W367. https://doi.org/10.1093/nar/gky473
Grosdidier A, Zoete V, Michielin O (2011) SwissDock, a protein-small molecule docking web service based on EADock DSS. Nucleic Acids Res 39 (Web Server issue):W270–277. doi: 10.1093/nar/gkr366.
Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, Ferrin TE (2004) UCSF Chimera–a visualization system for exploratory research and analysis. J Comput Chem 25(13):1605–1612. https://doi.org/10.1002/jcc.20084
Xia Y, Chu W, Qi Q, Xun L (2015) New insights into the QuikChange process guide the use of Phusion DNA polymerase for site-directed mutagenesis. Nucleic Acids Res 43(2):e12. https://doi.org/10.1093/nar/gku1189
Zhang YN, Lu FP, Chen GQ, Li Y, Wang JL (2009) Expression, purification, and characterization of phosphatidylserine synthase from Escherichia coli K12 in Bacillus subtilis. J Agric Food Chem 57(1):122–126. https://doi.org/10.1021/jf802664u
Acknowledgement
This work was supported by the Ministry of Science and Technology of Taiwan (Grant No. MOST 106–2313-B-002–024).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Huang, RY., Lee, CY. Characterization of a Phosphatidylserine Synthase of Vibrio parahaemolyticus. Curr Microbiol 77, 710–715 (2020). https://doi.org/10.1007/s00284-019-01854-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01854-x