Abstract
Purpose
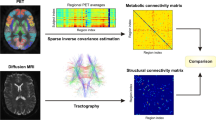
Sparse inverse covariance estimation (SICE) is increasingly utilized to estimate inter-subject covariance of FDG uptake (FDGcov) as proxy of metabolic brain connectivity. However, this statistical method suffers from the lack of robustness in the connectivity estimation. Patterns of FDGcov were observed to be spatially similar with patterns of structural connectivity as obtained from DTI imaging. Based on this similarity, we propose to regularize the sparse estimation of FDGcov using the structural connectivity.
Methods
We retrospectively analyzed the FDG-PET and DTI data of 26 healthy controls, 41 patients with Alzheimer’s disease (AD), and 30 patients with frontotemporal lobar degeneration (FTLD). Structural connectivity matrix derived from DTI data was introduced as a regularization parameter to assign individual penalties to each potential metabolic connectivity. Leave-one-out cross validation experiments were performed to assess the differential diagnosis ability of structure weighted SICE approach. A few approaches of structure weighted were compared with the standard SICE.
Results
Compared to the standard SICE, structural weighting has shown more stable performance in the supervised classification, especially in the differentiation AD vs. FTLD (accuracy of 89–90%, while unweighted SICE only 85%). There was a significant positive relationship between the minimum number of metabolic connection and the robustness of the classification accuracy (r = 0.57, P < 0.001). Shuffling experiments showed significant differences between classification score derived with true structural weighting and those obtained by randomized structure (P < 0.05).
Conclusion
The structure-weighted sparse estimation can enhance the robustness of metabolic connectivity, which may consequently improve the differentiation of pathological phenotypes.





Similar content being viewed by others
References
Yakushev I, Chételat G, Fischer FU, Landeau B, Bastin C, Scheurich A, et al. Metabolic and structural connectivity within the default mode network relates to working memory performance in young healthy adults. Neuroimage. 2013;79:184–90. https://doi.org/10.1016/j.neuroimage.2013.04.069.
Wang M, Jiang J, Yan Z, Alberts I, Ge J, Zhang H, et al. Individual brain metabolic connectome indicator based on Kullback-Leibler divergence similarity estimation predicts progression from mild cognitive impairment to Alzheimer's dementia. Eur J Nucl Med Mol Imaging. 2020;47(12):2753–64. https://doi.org/10.1007/s00259-020-04814-x.
Huang S, Li J, Sun L, Ye J, Fleisher A, Wu T, et al. Learning brain connectivity of Alzheimer's disease by sparse inverse covariance estimation. Neuroimage. 2010;50(3):935–49. https://doi.org/10.1016/j.neuroimage.2009.12.120.
Yakushev I, Drzezga A, Habeck C. Metabolic connectivity: methods and applications. Current opinion in neurology. 2017;30(6):677–85. https://doi.org/10.1097/wco.0000000000000494.
Morbelli S, Perneczky R, Drzezga A, Frisoni GB, Caroli A, van Berckel BN, et al. Metabolic networks underlying cognitive reserve in prodromal Alzheimer disease: a European Alzheimer disease consortium project. J Nucl Med. 2013;54(6):894–902. https://doi.org/10.2967/jnumed.112.113928.
Perani D, Farsad M, Ballarini T, Lubian F, Malpetti M, Fracchetti A, et al. The impact of bilingualism on brain reserve and metabolic connectivity in Alzheimer's dementia. Proc Natl Acad Sci U S A. 2017;114(7):1690–5. https://doi.org/10.1073/pnas.1610909114.
Titov D, Diehl-Schmid J, Shi K, Perneczky R, Zou N, Grimmer T, et al. Metabolic connectivity for differential diagnosis of dementing disorders. J Cereb Blood Flow Metab. 2017;37(1):252–62. https://doi.org/10.1177/0271678X15622465.
Jeong Y, Cho SS, Park JM, Kang SJ, Lee JS, Kang E, et al. 18F-FDG PET findings in frontotemporal dementia: an SPM analysis of 29 patients. J Nucl Med. 2005;46(2):233–9.
Toussaint PJ, Perlbarg V, Bellec P, Desarnaud S, Lacomblez L, Doyon J, et al. Resting state FDG-PET functional connectivity as an early biomarker of Alzheimer's disease using conjoint univariate and independent component analyses. Neuroimage. 2012;63(2):936–46. https://doi.org/10.1016/j.neuroimage.2012.03.091.
Friedman J, Hastie T, Tibshirani R. Sparse inverse covariance estimation with the graphical lasso. Biostatistics. 2008;9(3):432–41. https://doi.org/10.1093/biostatistics/kxm045.
Tucholka A, Grau-Rivera O, Falcon C, Rami L, Sánchez-Valle R, Lladó A, et al. Structural connectivity alterations along the Alzheimer's disease continuum: reproducibility across two independent samples and correlation with cerebrospinal fluid amyloid-β and Tau. J Alzheimer's Dis : JAD. 2018;61(4):1575–87. https://doi.org/10.3233/jad-170553.
Alm KH, Bakker A. Relationships between diffusion tensor imaging and cerebrospinal fluid metrics in early stages of the Alzheimer's disease continuum. J Alzheimer's Dis : JAD. 2019;70(4):965–81. https://doi.org/10.3233/jad-181210.
Yakushev I, Ripp I, Wang M, Savio A, Schutte M, Lizarraga A, et al. Mapping covariance in brain FDG uptake to structural connectivity. Eur J Nucl Med Mol Imaging. 2021. https://doi.org/10.1007/s00259-021-05590-y.
Broser PJ, Groeschel S, Hauser TK, Lidzba K, Wilke M. Functional MRI-guided probabilistic tractography of cortico-cortical and cortico-subcortical language networks in children. Neuroimage. 2012;63(3):1561–70. https://doi.org/10.1016/j.neuroimage.2012.07.060.
Sreedharan RM, Menon AC, James JS, Kesavadas C, Thomas SV. Arcuate fasciculus laterality by diffusion tensor imaging correlates with language laterality by functional MRI in preadolescent children. Neuroradiology. 2015;57(3):291–7. https://doi.org/10.1007/s00234-014-1469-1.
Zhu D, Li K, Guo L, Jiang X, Zhang T, Zhang D, et al. DICCCOL: dense individualized and common connectivity-based cortical landmarks. Cereb Cortex. 2013;23(4):786–800. https://doi.org/10.1093/cercor/bhs072.
Bowman FD, Zhang L, Derado G, Chen S. Determining functional connectivity using fMRI data with diffusion-based anatomical weighting. Neuroimage. 2012;62(3):1769–79. https://doi.org/10.1016/j.neuroimage.2012.05.032.
Ng B, Varoquaux G, Poline JB, Thirion B. A novel sparse graphical approach for multimodal brain connectivity inference. Med Image Comput Comput Assist Interv. 2012;15(Pt 1):707–14. https://doi.org/10.1007/978-3-642-33415-3_87.
McKhann G, Drachman D, Folstein M, Katzman R, Price D, Stadlan EM. Clinical diagnosis of Alzheimer's disease: report of the NINCDS-ADRDA Work Group under the auspices of Department of Health and Human Services Task Force on Alzheimer's Disease. Neurology. 1984;34(7):939–44. https://doi.org/10.1212/wnl.34.7.939.
Neary D, Snowden JS, Gustafson L, Passant U, Stuss D, Black S, et al. Frontotemporal lobar degeneration: a consensus on clinical diagnostic criteria. Neurology. 1998;51(6):1546–54. https://doi.org/10.1212/wnl.51.6.1546.
Gonzalez-Escamilla G, Lange C, Teipel S, Buchert R, Grothe MJ. PETPVE12: an SPM toolbox for partial volume effects correction in brain PET - application to amyloid imaging with AV45-PET. Neuroimage. 2017;147:669–77. https://doi.org/10.1016/j.neuroimage.2016.12.077.
Hammers A, Allom R, Koepp MJ, Free SL, Myers R, Lemieux L, et al. Three-dimensional maximum probability atlas of the human brain, with particular reference to the temporal lobe. Hum Brain Mapp. 2003;19(4):224–47. https://doi.org/10.1002/hbm.10123.
Wilkins B, Lee N, Gajawelli N, Law M, Leporé N. Fiber estimation and tractography in diffusion MRI: development of simulated brain images and comparison of multi-fiber analysis methods at clinical b-values. Neuroimage. 2015;109:341–56. https://doi.org/10.1016/j.neuroimage.2014.12.060.
Thirion B, Varoquaux G, Dohmatob E, Poline JB. Which fMRI clustering gives good brain parcellations? Front Neurosci. 2014;8:167. https://doi.org/10.3389/fnins.2014.00167.
Funding
This work was supported by the National Natural Science Foundation of China (82020108013) and the research project of Shanghai Health Commission (2020YJZX0111).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study was performed in line with the principles of the Declaration of Helsinki and with ethical standards of the institutional and/or national research committee. Informed consent was obtained from all individual participants included in the study.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Advanced Image Analyses (Radiomics and Artificial Intelligence).
Supplementary Information
ESM 1
(DOCX 435 kb)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, M., Schutte, M., Grimmer, T. et al. Reducing instability of inter-subject covariance of FDG uptake networks using structure-weighted sparse estimation approach. Eur J Nucl Med Mol Imaging 50, 80–89 (2022). https://doi.org/10.1007/s00259-022-05949-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00259-022-05949-9