Abstract
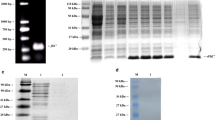
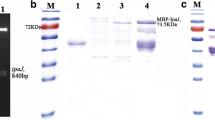
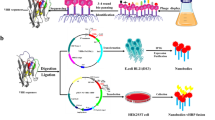
Salmonella Enteritidis (SE) causes both horizontal and vertical transmission of diseases in poultry industry and is also one of the main causes of human food poisoning. Sequence analysis of the sef operon of poultry-derived Salmonella serotypes showed the presence of an entire sef operon in SE, whereas only sef pseudogenes were found in Salmonella Gallinarum and Salmonella Pullorum. Subsequently, the sef operon of SE was cloned into the pBR322 plasmid and expressed in a modified Escherichia coli strain SE5000. sef operon expression was demonstrated using sodium dodecyl sulfate-polyacrylamide gel electrophoresis, western blot, agglutination assay, and transmission electron microscopy. The results showed that SE5000+Sef, but not SE5000+pBR322, could specifically react with SE-positive chicken serum in an agglutination assay, which could be clearly visualized by the naked eye within less than 2 min. In contrast, SE5000+Sef could not be recognized in Salmonella Gallinarum– and Salmonella Pullorum–positive chicken sera. Next, taking advantage of the exclusive presence of an entire sef operon in SE, we set up an agglutination-based detection system to monitor the dynamics of Sef-targeted antibody from SE-infected chicks for 47 days. Using the proposed detection method, SE was readily detectable starting from 2 weeks post-infection. Finally, we compared the proposed SE5000+Sef-based detection system with commercially available agglutination antigen using the classical bacterial isolation and identification procedure as reference. The results showed that the SE5000+Sef system was more consistent with the results of bacterial isolation and identification with almost 100% accuracy. We established a simple, sensitive, and cheap agglutination method for rapid and specific detection of SE-infected chickens, which can facilitate epidemiological investigation and eradication of SE infections.
Key points
• Only the Salmonella Enteritidis serotype expressed Sef fimbriae in chicken infected with SE.
• A rapid, large-scale method of detection by the naked eye of detection of SE-infected chicken is presented.






Similar content being viewed by others
Data availability
The authors declare that the data supporting the findings of this study are available within the article.
References
Ahmed A, Wang M, Khan R, Shah AA, Guo H, Malik S, Xia K, Hu Z (2021) A splice-site variant (C.3289-1G>T) in OTOF underlies profound hearing loss in a Pakistani kindred. BMC Med Genet 14:2. https://doi.org/10.1186/s12920-020-00859-x
Eriksson E, Aspan A (2007) Comparison of culture, ELISA and PCR techniques for Salmonella detection in faecal samples for cattle, pig and poultry. BMC Vet Res 3:21. https://doi.org/10.1186/1746-6148-3-21
Fasano RM, Sullivan HC, Bray RA, Gebel HM, Meyer EK, Winkler AM, Josephson CD, Stowell SR, Duncan A, Roback JD (2017) Genotyping applications for transplantation and transfusion management: the Emory experience. Arch Pathol Lab Med 141(3):329–340. https://doi.org/10.5858/arpa.2016-0277-SA
Gal-Mor O, Boyle EC, Grassl GA (2014) Same species, different diseases: how and why typhoidal and non-typhoidal Salmonella enterica serovars differ. Front Microbiol 391(5):1–10. https://doi.org/10.3389/fmicb.2014.00391
Gantois I, Ducatelle R, Pasmans F, Haesebrouck F, Gast R, Humphrey TJ, Van Immerseel F (2009) Mechanisms of egg contamination by Salmonella Enteritidis. FEMS Microbiol Rev 33(4):718–738. https://doi.org/10.1111/j.1574-6976.2008.00161.x
Gast RK (1997) Detecting infections of chickens with recent Salmonella Pullorum isolates using standard serological methods. Poult Sci 76(1):17–23. https://doi.org/10.1093/ps/76.1.17
Kang M-S, Kwon Y-K, Jung B-Y, Kim A, Lee K-M, An B-K, Song E-A, Kwon J-H, Chung G-S (2011) Differential identification of Salmonella enterica subsp enterica serovar Gallinarum biovars Gallinarum and Pullorum based on polymorphic regions of glgC and speC genes. Vet Microbiol 147(1-2):181–185. https://doi.org/10.1016/j.vetmic.2010.05.039
Kasturi K, Drgon T (2017) Real-time PCR method for detection of Salmonella spp. in environmental samples. Appl Environ Microbiol 83(14). https://doi.org/10.1128/aem.00644-17
Khan AS, Schifferli DM (1994) A minor 987P protein different from the structural fimbrial subunit is the adhesin. Infect Immun 62(10):4233–4243. https://doi.org/10.1128/iai.62.10.4233-4243
Kuehn BM (2010) Salmonella cases traced to egg producers findings trigger recall of more than 500 million eggs. JAMA-J AM Med Assoc 304(12):1316. https://doi.org/10.1001/jama.2010.1330
Kurien BT, Scofield RH (2015) Western blotting: an introduction: methods and protocols. Methods Mol Biol 1312:17–30
Li J, Ma B, Fang J, Zhi A, Chen E, Xu Y, Yu X, Sun C, Zhang M (2020) Recombinase polymerase amplification (RPA) combined with lateral flow immunoassay for rapid detection of Salmonella in food. Foods 9(27):1–12. https://doi.org/10.3390/foods9010027
Liquan Huang YW (2006) Study on the antibacterial activity of Leontopodium leontopodioides (Willd.) Beauv. in vitro. J Traditional Chinese Vet Med 1:5–7. https://doi.org/10.13823/j.cnki.jtcvm.2006.01.001
Lynch MJB, Leon-Velarde CG, McEwen S, Odumeru JA (2004) Evaluation of an automated immunomagnetic separation method for the rapid detection of Salmonella species in poultry environmental samples. J Microbiol Methods 58(2):285–288. https://doi.org/10.1016/j.mimet.2004.04.005
Machado I, Garrido V, Hernandez LI, Botero J, Bastida N, San-Roman B, Grillo M-J, Hernandez FJ (2019) Rapid and specific detection of Salmonella infections using chemically modified nucleic acid probes. Anal Chim Acta 1054:157–166. https://doi.org/10.1016/j.aca.2018.12.027
Majowicz SE, Musto J, Scallan E, Angulo FJ, Kirk M, O’Brien SJ, Jones TF, Fazil A, Hoekstra RM, Int Collaboration Enteric Dis B (2010) The global burden of nontyphoidal Salmonella gastroenteritis. Clin Infect Dis 50(6):882–889. https://doi.org/10.1086/650733
Margot H, Saegesser G, Zweifel C, Stephan R (2013) Performance of a commercially available latex agglutination test kit for the identification of Salmonella. J Food Saf Food Qual 64(3):75–77. https://doi.org/10.2376/0003-925x-64-75
Miao J, Han N, Qiang Y, Zhang T, Li X, Zhang W (2017) 16SPIP: a comprehensive analysis pipeline for rapid pathogen detection in clinical samples based on 16S metagenomic sequencing. BMC Bioinformatics 18:255–259. https://doi.org/10.1186/s12859-017-1975-3
Oliveira GH, Berchieri A Jr, Montassier HJ (2006) Chicken serologic response to Salmonella enterica serotype Typhimurium assessed by ELISA. Braz J Poult Sci 8(1):51–54. https://doi.org/10.1590/s1516-635x2006000100008
Rajashekara G, Munir S, Lamichhane CM, Back A, Kapur V, Halvorson DA, Nagaraja KV (1998) Application of recombinant fimbrial protein for the specific detection of Salmonella Enteritidis infection in poultry. Diagn Microbiol Infect Dis 32(3):147–157. https://doi.org/10.1016/s0732-8893(98)00091-1
Ricke SC, Khatiwara A, Kwon YM (2013) Application of microarray analysis of foodborne Salmonella in poultry production: a review. Poult Sci 92(9):2243–2250. https://doi.org/10.3382/ps.2012-02740
Roychoudhury P, Murugkar HV, Rahman H, Subudhi PK (2009) Dot-ELISA for detection of enterotoxin production by Salmonella isolates from Japanese quail. Indian J Anim Sci 79(2):161–163
Schifferli DM, Beachey EH, Taylor RK (1991) 987P fimbrial gene identification and protein characterization by T7 RNA polymerase induced transcription and TnphoA mutagenesis. Mol Microbiol 5:61–70
Schreier S, Doungchawee G, Chadsuthi S, Triampo D, Triampo W (2013) Leptospirosis: current situation and trends of specific laboratory tests. Expert Rev Clin Immunol 9(3):263–280. https://doi.org/10.1586/eci.12.110
Song Q, Shen X, Yang Y, Zhang D, Gao H (2016) Genetically similar isolates of Salmonella enterica serotype Enteritidis persistent in China for a long-term period. J Food Sci 81(7):M1778–M1781. https://doi.org/10.1111/1750-3841.13339
Stoica C, Ancuta PN, Lucaciu IE, Banciu AR, Sorea S, Atanasescu A, Lazar MN (2018) Computerized high-tech detection technology of immunofluorescence labelled waterborne pathogenic bacteria. Rev Chim-Bucharest 69(11):4166–4170
Thorns CJ, Sojka MG, Chasey D (1990) Detection of a novel fimbrial structure on the surface of Salmonella Enteritidis by using a monoclonal antibody. J Clin Microbiol 28(11):2409–2414. https://doi.org/10.1128/jcm.28.11.2409-2414.1990
Thorns CJ, McLaren IM, Sojka MG (1994) The use of latex particle agglutination to specifically detect Salmonella Enteritidis. Int J Food Microbiol 21(1-2):47–53. https://doi.org/10.1016/0168-1605(94)90199-6
Turcotte C, Woodward MJ (1993) Cloning, DNA nucleotide sequence and distribution of the gene encoding the Sef fimbrial antigen of Salmonella Enteritidis. J Gen Microbiol 139(7):1477–1485. https://doi.org/10.1099/00221287-139-7-1477
Wang X, Wang H, Li T, Liu F, Cheng Y, Guo X, Wen G, Luo Q, Shao H, Pan Z, Zhang T (2020) Characterization of Salmonella spp. isolated from chickens in central China. BMC Vet Res 16(1):299–308. https://doi.org/10.1186/s12917-020-02513-1
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ (2009) Jalview version 2-a multiple sequence alignment editor and analysis workbench. Bioinformatics 25(9):1189–1191. https://doi.org/10.1093/bioinformatics/btp033
Xia P, Song Y, Zou Y, Yang Y, Zhu G (2015) F4+ enterotoxigenic Escherichia coli (ETEC) adhesion mediated by the major fimbrial subunit FaeG. J Basic Microbiol 55(9):1118–1124. https://doi.org/10.1002/jobm.201400901
Yan M, Tam FCH, Kan B, Lim PL (2011) Combined rapid (TUBEX) test for typhoid-paratyphoid a fever based on strong anti-O12 response: design and critical assessment of sensitivity. PLoS One 6(9):e24743. https://doi.org/10.1371/journal.pone.0024743
Yang F-H, Zhang Q, Liang Q-Y, Wang S-Q, Zhao B-X, Wang Y-T, Cai Y, Li G-F (2015) Bioavailability enhancement of paclitaxel via a novel oral drug delivery system: paclitaxel-loaded glycyrrhizic acid micelles. Molecules 20(3):4337–4356. https://doi.org/10.3390/molecules20034337
Yang B, Niu Q, Yang Y, Dai P, Yuan T, Xu S, Pan X, Zhu G (2019) Self-made Salmonella Pullorum agglutination antigen development and its potential practical application. Poult Sci 98(12):6326–6332. https://doi.org/10.3382/ps/pez453
Yang Y, Wang P, Xia P, Yang B, Dai P, Hong T, Li J, Meng X, El Qaidi S, Zhu G (2020) Rapid detection of flagellated and non-flagellated Salmonella by targeting the common flagellar hook gene flgE. Appl Microbiol Biotechnol 104(22):9719–9732. https://doi.org/10.1007/s00253-020-10925-0
Yue M, Rankin SC, Blanchet RT, Nulton JD, Edwards RA, Schifferli DM (2012) Diversification of the Salmonella fimbriae: a model of macro- and microevolution. PLoS One 7(6):e38596. https://doi.org/10.1371/journal.pone.0038596
Zhu CH, Musa HH, Wu SL, Zhu GQ (2009) The role of Sef fimbriae in pathogenesis and enhancing the immunity of Salmonella Enteritidis. Afr J Microbiol Res 3(5):191–194
Zhu CH, Wu J, Chen WW, Hassan HM, Zhu GQ (2010) Difference and variation of the sef operon gene clusters in Salmonella Pullorum. J Basic Microbiol 50:S120–S123. https://doi.org/10.1002/jobm.200900262
Acknowledgements
The authors gratefully acknowledge Professor Song Gao in Yangzhou University, College of Veterinary Medicine, for providing the serum for this study.
Funding
This work was supported by the College Students Research & Practice Innovation Program, Graduates Research & Practice Innovation Program of Jiangsu Province (No. KYCX20_3004), International Collaboration Program from Science and Technology Agency of Jiangsu Province (2019), and Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Contributions
XG and QH performed the experiments, analyzed the data, and wrote the manuscript. JL and PX participated in the data analysis and wrote the paper. QD and GZ conceived and designed the study, participated in experimental work, and wrote the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This article does not contain any studies with human participants performed by any of the authors. The current study was approved by the Institutional Animal Care and Use Committee of the Yangzhou University College of Veterinary Medicine of China.
Consent for publication
All authors listed on this manuscript have read and agreed to the publication of this research.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Gu, X., Hou, Q., Liu, J. et al. Sef fimbria operon construction, expression, and function for direct rapid detection of Salmonella Enteritidis. Appl Microbiol Biotechnol 105, 5631–5641 (2021). https://doi.org/10.1007/s00253-021-11400-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11400-0