Abstract
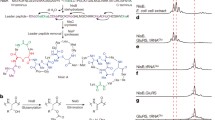
The lantibiotic nukacin ISK-1 is an antimicrobial peptide containing unusual amino acids such as lanthionine and dehydrobutyrine. The nukacin ISK-1 prepeptide (NukA) undergoes posttranslational modifications, such as the dehydration and cyclization reactions required to form the unusual amino acids by the modification enzyme NukM. We have previously constructed a system for the introduction of unusual amino acids into NukA by coexpression of NukM in Escherichia coli. Using this system, we describe the substrate specificity of NukM by the coexpression of a series of NukA mutants. Our results revealed the following characteristics of NukM: (1) its dehydration activity is not coupled to its cyclization activity; (2) its dehydration activity is site-specific; (3) the length of the substrate is important for its dehydration activity. Furthermore, we succeeded in introducing a novel thioether bridge in NukA by replacing an unmodified Ser at position 27 with a Cys residue.


Similar content being viewed by others
References
Bierbaum G, Szekat C, Josten M, Heidrich C, Kempter C, Jung G, Sahl HG (1996) Engineering of a novel thioether bridge and role of modified residues in the lantibiotic Pep5. Appl Environ Microbiol 71:562–565
Chatterjee C, Paul M, Xie L, van der Donk WA (2005a) Biosynthesis and mode of action of lantibiotics. Chem Rev 105:633–684
Chatterjee C, Miller LM, Leung YL, Xie L, Yi M, Kelleher NL, van der Donk WA (2005b) Lacticin 481 synthetase phosphorylates its substrate during lantibiotic production. J Am Chem Soc 127:15332–15333
Chatterjee C, Patton GC, Cooper L, Paul M, van der Donk WA (2006) Engineering dehydro amino acids and thioethers into peptides using lacticin 481 synthetase. Chem Biol 13:1109–1117
Helfrich M, Entian KD, Stein T (2007) Structure-function relationships of the lanthionine cyclase SpaC involved in biosynthesis of the Bacillus subtilis peptide antibiotic subtilin. Biochemistry 46:3224–3233
Ishizaki A, Takese E, Ikai T, Kumai S, Nagano R, Sonomoto K, Doi K, Ogata S, Kawamura Y, Ezaki T (2001) Taxonomic position of new bacteriocin (nukacin ISK-1) producer isolated from long-aged Nukadoko. J Gen Appl Microbiol 47:143–147
Kimura H, Matsusaki H, Sashihara K, Sonomoto K, Ishizaki A (1998) Purification and partial identification of bacteriocin ISK-1, a new lantibiotic produced by Pediococcus sp. ISK-1. Biosci Biotechnol Biochem 62:2341–2345
Kluskens LD, Kuipers A, Rink R, de Boef E, Fekken S, Driessen AJM, Kuipers OP, Moll GN (2005) Post-translational modification of therapeutic peptides by NisB, the dehydratase of the lantibiotic nisin. Biochemistry 44:12827–12834
Koponen O, Tolonen M, Qiao M, Wahlstrom G, Helin J, Saris PEJ (2002) NisB is required for the dehydration and NisC for the lanthionine formation in the post-translational modification of nisin. Microbiology 148:3561–3568
Levengood MR, van der Donk WA (2008) Use of lantibiotic synthetases for the preparation of bioactive constrained peptides. Bioorg Med Chem Lett 18:3025–3028
Li B, Yu JPJ, Brunzelle JS, Moll GN, van der Donk WA, Nair SK (2006) Structure and mechanism of the lantibiotic cyclase involved in nisin biosynthesis. Science 311:1464–1467
McClerren AL, Cooper LE, Quan C, Thomas PM, Kelleher NL, van der Donk WA (2006) Discovery and in vitro biosynthesis of haloduracin, a two component lantibiotic. Proc Natl Acad Sci USA 14:17243–17248
Nagao J, Harada Y, Shioya K, Aso Y, Zendo T, Nakayama J, Sonomoto K (2005a) Lanthionine introduction into nukacin ISK-1 prepeptide by co-expression with modification enzyme NukM in Escherichia coli. Biochem Biophys Res Commun 336:507–513
Nagao J, Aso Y, Sashihara T, Shioya K, Adachi A, Nakayama J, Sonomoto K (2005b) Localization and interaction of the biosynthetic proteins for the lantibiotic, nukacin ISK-1. Biosci Biotechnol Biochem 69:1341–1347
Nagao J, Asaduzzaman SM, Aso Y, Okuda K, Nakayama J, Sonomoto K (2006) Lantibiotics: insight and foresight for new paradigm. J Biosci Bioeng 102:139–149
Nagao J, Aso Y, Shioya K, Nakayama J, Sonomoto K (2007) Lantibiotic engineering: molecular characterization and exploitation of lantibiotic-synthesizing enzyme for peptide engineering. J Mol Microbiol Biotechnol 13:235–242
Patton GC, Paul M, Cooper LE, Chatterjee C, van der Donk WA (2008) The importance of the leader sequence for directing lanthionine formation in lacticin 481. Biochemistry 47:7342–7351
Paul M, Patton GC, van der Donk WA (2007) Mutants of the zinc ligands of lacticin 481 synthetase retain dehydration activity but have impaired cyclization activity. Biochemistry 46:6268–6276
Rink R, Kuipers A, de Boef E, Leenhouts KJ, Driessen AJM, Moll GN, Kuipers OP (2005) Lantibiotic structures as guidelines for the design of peptides that can be modified by lantibiotic enzymes. Biochemistry 44:8873–8882
Sambrook J, Frisch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY
Wu J, Yang Y, Watson JT (1998) Trapping of intermediates during the refolding of recombinant human epidermal growth factor (hEGF) by cyanylation, and subsequent structural elucidation by mass spectrometry. Protein Sci 7:1017–1028
Xie L, Miller LM, Chatterjee C, Averin O, Kelleher NL, van der Donk WA (2004) Lacticin 481: in vitro reconstitution of lantibiotic synthetase. Science 303:679–681
You YO, van der Donk WA (2007) Mechanistic investigation of the dehydration reaction of lacticin 481 synthetase using site-direct mutagenesis. Biochemistry 46:5991–6000
Zendo T, Nakayama J, Fujita K, Sonomoto K (2008) Bacteriocin detection by liquid chromatography/mass spectrometry for rapid identification. J Appl Microbiol 104:499–507
Acknowledgments
We thank N. Futami (Toray Research Center, Tokyo, Japan) for the amino acid sequence analysis. Our work on lantibiotics nukacin ISK-1 has been partially supported by grants from the following organizations: the Japan Society for the Promotion of Science, the Japan Science Society, the Novartis Foundation (Japan) for the Promotion of Science, the Novozymes Japan Research Fund, and the Nagase Science and Technology Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shioya, K., Harada, Y., Nagao, Ji. et al. Characterization of modification enzyme NukM and engineering of a novel thioether bridge in lantibiotic nukacin ISK-1. Appl Microbiol Biotechnol 86, 891–899 (2010). https://doi.org/10.1007/s00253-009-2334-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-009-2334-8