Abstract
Antigen-specific CD4+ T-lymphocyte responses are restricted by major histocompatibility complex class II molecules, which influence T-cell priming during infection. Human leukocyte antigen (HLA) and bovine leukocyte antigen (BoLA) DRB3 and DQ genes are polymorphic, but unlike HLA, many BoLA haplotypes have duplicated DQ genes, and antibody-blocking studies indicated that BoLA-DQ molecules present various pathogen epitopes. Limited experimentation also suggested that BoLA-DQ molecules formed by interhaplotype pairing of A and B chains are functional. To compare antigen presentation by DR and DQ molecules and to definitively demonstrate functional BoLA-DQ molecules derived from interhaplotype pairing, different combinations of DR or DQ A and B proteins were expressed with CD80 in 293-F cells for use as antigen-presenting cells (APCs). This approach identified 11 unique restriction elements including five DR and six DQ pairs for antigen-specific CD4+ T-cell responses against tick-transmitted bovine hemoparasites Anaplasma marginale or Babesia bovis. Interhaplotype pairing of DQ A and B molecules was demonstrated. Testing of six expressed DQA/B pairs from an animal with duplicated DQ haplotypes (DH16A/DH22H) demonstrated that an interhaplotype pair, DQA*2206/DQB*1301, presented A. marginale peptide B. In DH22H and DH16A homozygous animals, DQA*2206 was tightly linked with DQB*1402, and DQA*22021 was linked with DQB*1301. APCs from these donors could not present peptide B, confirming that DQA*2206/DQB*1301 encoded a functional interhaplotype pair. Functional BoLA-DQ molecules are generated by both intrahaplotype and interhaplotype pairing of A and B chains and play a similar role to BoLA-DR in priming helper T-cell responses to important pathogens.
Similar content being viewed by others
Introduction
The main function of major histocompatibility complex (MHC) class II molecules is to present processed pathogen-derived peptides to CD4+ T lymphocytes. MHC class II alleles are redundant and highly polymorphic, enhancing the repertoire of epitopes that an individual can recognize. Whereas mice express two class II proteins, I-E and I-A, humans express three (HLA-DR, HLA-DQ, and HLA-DP), and heterozygous individuals can therefore express at least four (mice) or six (humans) sets of class II gene products. Cattle express only two class II proteins, DR and DQ. However, in the majority of haplotypes expressed by Holstein cattle, bovine leukocyte antigen (BoLA)-DQ molecules are duplicated, permitting additional diversity through intrahaplotype and, potentially, interhaplotype pairing of DQA and DQB proteins, as reported for human DQA and DQB (Kwok et al. 1993). Interhaplotype pairing refers to pairing of DQA and DQB gene products encoded by different chromosomes. When DQ genes are duplicated, intrahaplotype pairing refers to pairing of DQA and DQB gene products encoded by the same chromosome, which can occur either within a DQA/DQB locus (referred to as “adjacent” pairing) or between the sets of duplicated gene loci (“nonadjacent” pairing) but within the same haplotype. Because the DRA chain in both humans and cattle is monomorphic, polymorphism in DRB genes is the only source of diversity in DR molecules. In cattle, only one DRB (DRB3) gene is known to be functional, whereas humans use DRB1 predominantly, but DRB3, DRB4, and DRB5 genes are functional in some haplotype groups (Schreuder et al. 2005). Unlike DRA, both HLA- and BoLA-DQA and DQB chains exhibit polymorphism, which would intuitively be used by the immune system to increase the antigenic epitope-binding repertoire (Lewin et al. 1999). However, for HLA-DQ, this does not appear to be the case. For reasons that are not understood, the great majority of human peripheral blood derived T-cell clones is restricted by DR molecules (Mølvig et al. 1989; Endl et al. 1997), although HLA-DQ molecules can present peptides from different types of pathogens (Calvo-Calle et al. 1997; Agrewala and Wilkinson 1998; Kwok et al. 2000). The best studied role of HLA-DQ molecules is their association, either positive or negative, with immunologically mediated diseases including insulin-dependent diabetes mellitus (Abbas et al. 2000) and celiac disease (Sollid et al. 1989).
In cattle, BoLA-DRB3 exon 2 has been extensively sequenced and most of the DRB3 alleles in Holstein cattle are known, whereas the sequence information for BoLA-DQ genes and the understanding of the role of BoLA-DQ molecules in bovine immune system are limited. To date, 106 DRB3, 46 DQA, and 52 DQB alleles have been reported (http://www.projects.roslin.ac.uk/bola, http://www.ebi.ac.uk/ipd/mhc/bola). Since BoLA-DRA is monomorphic and DRB3 is the only active DRB gene, it is likely that DQ molecules are as important as DR molecules for priming CD4+ T cells specific for many different pathogens. Using monoclonal antibody (mAb)-blocking assays, Glass et al. (2000) demonstrated presentation by BoLA-DQ molecules of foot-and-mouth disease virus (FMDV) epitopes to CD4+ T cells. We have similarly shown presentation of Babesia bovis and Anaplasma marginale epitopes to CD4+ T cells by BoLA-DR and BoLA-DQ molecules (Brown et al. 2002, 2003; Norimine et al. 2002, 2004). However, in our studies, ambiguous results were obtained with mAb used to block proliferation of a T-cell clone (Norimine et al. 2002; W.C. Brown, unpublished observations), and the specificity (DR vs DQ) of one mAb reagent (IL-A21) was also reportedly uncertain (Fogg et al. 2001). Nevertheless, evidence for functional interhaplotype pairing of DQ molecules was presented in studies using T-cell clones specific for FMDV (Glass et al. 2000) or A. marginale major surface protein (MSP)-1a (Brown et al. 2002). In both studies, the pattern of peptide presentation by antigen-presenting cells (APCs) expressing certain combinations of haplotypes was indicative of interhaplotype pairing of DQA and DQB chains, but definitive proof of this was lacking (Glass et al. 2000; Brown et al. 2002).
The present study used a transfected cell line expressing BoLA-DQ or BoLA-DR A and B proteins to determine whether DQ and DR molecules present antigen to CD4+ T cells with comparable frequency and to definitively demonstrate that interhaplotype pairing of BoLA-DQA and BoLA-DQB molecules results in the formation of functional DQ heterodimers. T-cell lines and clones specific for known epitopes of the bovine pathogens B. bovis and A. marginale were used. In addition, BoLA-DQ and BoLA-DR alleles were sequenced from the T-cell donors, and different gene pairs, together with bovine CD80 or CD86, were expressed in a human embryonic kidney cell line and used as a source of APCs. Our results show that BoLA-DQ molecules function like BoLA-DR molecules to present pathogen-derived antigenic peptides and therefore differ from HLA-DQ molecules.
Materials and methods
Experimental cattle and their BoLA haplotypes
All of the protocols in this study were reviewed and approved by the Washington State University Institutional Animal Care and Use Committee. Brahman–Angus cross cow C97 was infected with the Mexico strain of B. bovis (Brown et al. 1991), Holstein steer 87 was immunized with A. marginale native MSP1, which is a heterodimer of MSP1a and MSP1b (Brown et al. 2001a), and Holstein steer 61 was immunized with A. marginale native MSP2 (Brown et al. 2001b) as previously described. Holstein steer 04B93 and cows 131, 132, and 201 were used as nonimmune controls. The DRB3 alleles of animals C97, 61, 87, and 04B93 were determined by the polymerase chain reaction–restriction fragment length polymorphism (PCR–RFLP) method as developed by van Eijk et al. (1992a). RFLP analysis was particularly important for confirming whether animals had homozygous BoLA haplotypes. Further, each PCR product from exon 2 of each DRB3 gene was cloned into the pCR2.1-TOPO vector (Invitrogen) and sequenced using the M13 forward and reverse primers. Sequencing was performed using the Prism Ready Reaction Dye Deoxy Terminator cycle-sequencing kit and analyzed with the ABI Prism 373 genetic analyzer (Applied Biosystems). For all animals, DQA1 and DQA2 alleles were determined by sequencing their exon 2 regions using recently published primer sets (Park et al. 2004). All DQB1 and DQB2 alleles were identified by sequencing the entire cDNAs using the primer sets listed in Table 1. The class II haplotypes of all cattle are shown in Table 2.
Antigens and synthetic peptides
Babesia bovis Mexico strain merozoite antigen enriched in parasite cell membranes (CM) was prepared from cultured, infected erythrocytes following homogenization with a French pressure cell and centrifugation as described (Brown et al. 1991). A. marginale Florida strain organisms were isolated from thawed, infected bovine erythrocytes by sonication and differential centrifugation as described (Palmer and McGuire 1984). Antigens were diluted in phosphate-buffered saline (PBS; pH 7.4) containing 25 μg/ml of the protease inhibitors antipain and E-64 (Boehringer Mannheim) and 300 μg/ml phenylmethylsulfonyl fluoride (Sigma) and stored at −20°C. All peptides were synthesized by Gerhardt Munske, Laboratory for Biotechnology and Bioanalysis I, Washington State University, Pullman. The peptides were diluted to 1 mg/ml in PBS and stored at −20°C. The amino acid sequences of the peptides and their origin are listed in Table 3.
Transfection of 293-F cells with MHC class II, CD80, and CD86 pCR3.1 constructs
Total RNA was obtained from peripheral blood mononuclear cells (PBMCs) of each animal using Trizol Reagent (Invitrogen) and reverse-transcribed to cDNA in a 50-μl volume by using oligo (dT)16 (Perkin Elmer) according to the manufacturer's instructions. Amplification of full-length cDNA for DRA, DRB3, DQA, DQB, CD80, and CD86 was performed by PCR using the primer sets listed in Table 1. The PCR parameters were 94°C 10 min, 35 cycles at 94°C for 15 s, 60°C for 15 s, and 72°C for 2 min, with an extension of 72°C for 10 min, and finally 4°C. Each PCR product was cloned into the eukaryotic expression vector pCR3.1 (Invitrogen) using T4 ligase (Invitrogen). The ligation was performed overnight at 15°C. The sequence and direction of each insert was confirmed by sequencing in both direction using T7 forward and BGHR primers (Invitrogen). For use in transfection, 90% confluent human embryonic kidney 293-F cells (Invitrogen) were cultured in a six-well plate in Dulbecco's Minimal Essential Medium (DMEM, Invitrogen) supplemented with 10% heat-inactivated fetal calf serum (FCS; Atlanta Biologicals) and penicillin–streptomycin. Prior to transfection, the cells were washed once with DMEM (without FCS and antibiotics), and 800 μl of DMEM (without FCS and antibiotics) was added to each well. A plasmid DNA mixture consisting of 2 μg of each plasmid DNA was added to 100 μl of DMEM (without FCS and antibiotics), mixed with 6 μl of Plus Lipofectamine (Invitrogen), and incubated at room temperature for 15 min. The DNA mixture was then combined with a Lipofectamine mixture consisting of 100 μl DMEM (without FCS and antibiotics) and 4 μl of Lipofectamine and incubated at room temperature for 15 min. The DNA–Lipofectamine complex was added to the 293-F cells. After incubation for 3 h at 37°C, 1 ml of DMEM with 4% FCS (without antibiotics) was added, and the cells were cultured for 2 days. Medium was removed, complete RPMI-1640 medium (Brown et al. 1991) containing 50 μg/ml Mitomycin C (Sigma) was added, and the cells were incubated for 2 h at 37°C in a humidified atmosphere of 5% CO2 in air. The transfected cells were harvested by repetitive pipetting, washed three times with complete RPMI-1640 medium, and used as APCs.
Flow cytometric analysis of 293-F cells transfected with BoLA-class II, CD80, and CD86
Expression of BoLA-class II and costimulatory molecules on transfected 293-F cells was verified by flow cytometry using bovine DR-specific mAb IL-A21, DQ-specific mAbs TH22A and CC158, CD80-specific mAb IL-A159, and CD86-specific mAb IL-A190. mAb TH22A was purchased from the Washington State University Monoclonal Antibody Center. mAbs CC158, IL-A159, and IL-A190 were kindly provided by Chris Howard, Institute of Animal Health, Compton, UK, and Niall MacHugh, University of Edinburgh, UK. mAbs IL-A21, IL-A159, and IL-A190 were originally obtained from the former International Laboratory for Research on Animal Diseases (current International Livestock Research Institute), Nairobi, Kenya. For the secondary antibody, fluorescein isothiocyanate (FITC)-conjugated goat antimouse immunoglobulins (IgA, IgG, IgM) (CAPPEL) was used. To confirm coexpression of CD80 and DR or DQ, the mixture of antibovine CD80 mAb IL-A159 (IgG1) and either anti-BoLA-DR mAb IL-A21 (IgG2a) or anti-BoLA-DQ mAb CC158 (IgG2a) was used for the primary antibody (15 μg/ml each mAb). For the secondary antibody, the mixture of FITC-conjugated goat antimouse IgG2a and R-phycoerythrin (R-PE)-conjugated goat antimouse IgG1 (CALTAG Laboratories) was used.
T-cell proliferation assays
To establish short-term T-cell lines from animals 61 and 87, PBMCs were initially depleted of CD8+ and γδ T lymphocytes using antibody and complement lysis. Briefly, PBMCs were incubated for 30 min at 4°C with anti-CD8 mAb 7C2B and anti-TCR γ chain mAb CACT61A purchased from the Washington State University Monoclonal Antibody Center and diluted to 15 μg/ml in complete RPMI-1640 medium. The cells were washed once and incubated for 30 min at 37°C with rabbit complement (Sigma) diluted 1:16 in complete medium and washed twice. Viable cells were purified using a Histopaque 1083 (Sigma) gradient. Subsequently, 4×106 cells were cultured in 24-well plates (Costar) in 1.5 ml of complete medium with 10 μg/ml homogenate prepared from the Florida strain of A. marginale (Brown et al. 1998, 2001b). After 7 days, lymphocytes were washed in complete medium, and 7×105 cells/well were cultured for 7 days with 2×106 irradiated autologous PBMCs without antigen. Short-term T-cell lines from animal C97 were similarly established without CD8+ and γδ T-lymphocyte depletion and using 10 μg/ml B. bovis CM antigen to stimulate the T cells. CD4+ T-cell clones specific for A. marginale MSP1a, MSP2, or B. bovis small heat shock protein Hsp20 were described elsewhere (Brown et al. 2002, 2004; Norimine et al. 2004). Proliferation assays were performed for 3–4 days in duplicate or triplicate wells of round-bottomed 96-well plates (Costar) at 37°C in a humidified atmosphere of 5% CO2 in air. Briefly, 3×104 T cells and 2×105 autologous APCs were cultured with 0.1–10 μg/ml of antigen or peptide. The cells were radiolabeled during the last 18 h of culture with 3H-thymidine, and the results are reported as mean cpm of replicate cultures. In one experiment, APCs were prepared from nonautologous donors as described in the text. When transfected 293-F cells were used as APCs, 293-F cells were treated with 50 μg/ml Mitomycin C, plated in 96-well U-bottomed plates (5×104 cells/well), loaded with 0.1–10 μg/ml of peptide in triplicate wells in a total volume of 100 μl complete RPMI-1640 medium, and incubated for 1 h at 37°C in a humidified atmosphere of 5% CO2 in air. The plates were washed 2–3 times with complete RPMI-1640 medium by centrifugation at 900×g, and CD4+ T cells derived from either short-term cell lines or CD4+ T-cell clones (3×104 cells/well) were then added in a total volume of 100 μl complete RPMI-1640 medium. Cells were cultured for 3–4 days, radiolabeled, harvested, and counted.
Statistical significance of antigen-specific T-cell proliferation to peptide presented by different irradiated PBMCs as a source of APCs, untransfected 293-F cells, or 293-F cells transfected with different MHC class II A and B chains was evaluated by the Student's two-tailed t test. P values less than 0.05 are considered significant.
Results
BoLA-DR and BoLA-DQ alleles and their linkages in the experimental animals
The BoLA class II alleles identified are based on BoLA nomenclature web site (http://www.projects.roslin.ac.uk/bola, http://www.ebi.ac.uk/ipd/mhc/bola) and are shown in Table 2. Because the names of one DQA and one DQB allele from Brahman–Angus cross cow C97 were not available on the web site, the DQA allele (accession number AY730727) and DQB allele (accession number AY730728) have been designated DQA*C97 and DQB*C97 alleles, respectively. Animal 87 had heterozygous BoLA haplotypes (DH16A/DH22H), each of which contained duplicated DQ pairs. The DR-DQ linkages were confirmed by sequencing DR and DQ genes from animals 04B93 and 131 (DH16A homozygous) and from animals 132 and 201 (DH22H homozygous). As shown in Table 2, the DQA*2206 and DQB*1402 alleles are linked with the DRB3*1101 allele (RFLP 22) in haplotype DH22H, while the DQA* 22021 and DQB* 1301 alleles are linked with the DRB*1501 allele (RFLP 16) in haplotype DH16A. The pair of DQA*10011 and DQB*10021 alleles was consistently observed in both DH22H and DH16A haplotypes. Extensive sequencing and DRB3 RFLP analyses confirmed these homozygosities and linkages, and the linkages were consistent with results from sequencing DRB3 exon 2 from other Holstein cattle (data not shown). Animal C97 has unique heterozygous BoLA haplotypes (both not defined) with nonduplicated DQ alleles. The linkage between DR and DQ genes in this animal is unknown since no animals sharing these DR or DQ genes were available for analysis.
Flow cytometric analysis of DR, DQ, CD80, and CD86 expression on 293-F cells
Bovine CD80 and CD86 cDNAs were also cloned into the expression vector pCR3.1 and examined for their expression on transfected 293-F cells by flow cytometric analysis using mAb IL-A159 (anti-CD80) and IL-A190 (anti-CD86). Both proteins were expressed at similar levels, and no cross-reactivities between the mAbs were observed. Coexpression of CD80 and BoLA-DR or BoLA-DQ molecules on 293-F cells was confirmed by double staining. The level of MHC class II expression varied among combinations of A and B chains, ranging from approximately 40 to 80% (data not shown). Class II expression was never detected when 293-F cells were transfected with a single A or B chain pCR3.1 construct (data not shown). Table 4 summarizes the expression profiles of all possible DRA/B and DQA/B pairs in transfected 293-F cells. All DRA/B pairs were expressed, while expression of four DQA/B combinations, DQA*12011/DQB*1201 and DQA*2201/DQB*10051 (animal 61) and DQA*10011/DQB*1301 and DQA*10011/DQB*1402 (animal 87) was not detected. The DQA*22021 and DQA*2206 alleles were expressed in combination with all three DQB alleles tested, indicating that these alleles could form intra- and interhaplotype DQ molecules. The DQA*10011 allele was expressed only with the DQB*10021 allele, which was tightly linked. Although there are several conflicting reports regarding the specificity of IL-A21 (anti-DR) and TH22A (anti-DQ) mAbs (Fogg et al. 2001; Davis et al. 1987; Davies et al. 1994; Bissumbhar et al. 1994; Fraser et al. 1996), we did not observe any cross-reactivity with these reagents. The specificity of anti-DQ mAb CC158 was consistent with previous reports (Howard et al. 1997; Russell et al. 2000).
Presentation of antigen by BoLA-DR and BoLA-DQ transfected cells requires coexpression of CD80 or CD86
To determine if the MHC class II molecules expressed on 293-F cells were functional, T-cell proliferation assays were performed. Previous studies had identified numerous epitopes within B. bovis rhoptry-associated protein 1 (RAP-1) (Norimine et al. 2002, 2003), including CD4+ T-cell epitopes P3 (aa 174–203), P9 (aa 294–316), and CT-P2 (aa 386–408) using lymphocytes from animal C97 infected with B. bovis (Norimine et al. 2002). Since CD4+ T-lymphocyte responses against peptides P3, P9, and CT-P2 were apparently restricted by DR molecules as determined by mAb-blocking assays (Norimine et al. 2002), we examined products of DRA/DRB3*3001 and DRA/DRB3*4501 derived from animal C97 for their ability to present antigen. As shown in Fig. 1, CD4+ T-lymphocyte responses against peptides P3, P9, and CT-P2 were restricted by DRA/DRB3*3001-encoded (for P3 and P9) and DRA/DRB3*4501-endoded (for CT-P2) class II molecules. Importantly, T-cell proliferation was not observed in the absence of CD80 or CD86 expression, indicating that expression of a B7 molecule was required in this system. Because the efficiency of costimulation was comparable between CD80 and CD86, CD80 was used in subsequent MHC class II transfection experiments.
Peptide-specific T-cell proliferation using DR-transfectants as APCs requires coexpression of either CD80 or CD86. 293-F cells were transfected with a single set of DR or DQ A and B chain alleles from animal C97 with or without the CD80 or CD86 pCR3.1 construct. T cells from short-term cell lines (C97 CL) were cultured with the indicated transfected cells loaded with the following peptides: RAP-1 peptide P9 (a), RAP-1 peptide P3 (b), and RAP-1 peptide CT-P2 (c). Results are presented as the mean cpm of triplicate or duplicate cultures. Responses significantly higher than responses to peptide in the presence of nontransfected 293-F cells are indicated by an asterisk (P<0.05) (a). Responses significantly higher than responses to peptide in the presence of 293-F cells expressing the other DRB3 allele are indicated by an asterisk (P<0.05) (b and c)
Presentation of A. marginale MSP2 epitopes by DR and DQ molecules
Anaplasma marginale MSP2-immunized animal 61 (Brown et al. 2001b) has a homozygous MHC haplotype (DH8A/DH8A) as shown in Table 2. The DH8A haplotype contains duplicated DQ alleles, so there are four potential DQ A and B pairs. However, only DQA*12011/DQB*10051 and DQA*2201/DQB*1201 allelic pairs were expressed on transfected 293-F cells (Table 4). This indicated that animal 61 expressed three MHC class II molecules, DRA/DRB3*1201, DQA12011/DQB*10051, and DQA*2201/DQB*1201. We determined if these MHC class II molecules were functional using three different MSP2 peptides previously shown to stimulate Th cells (Brown et al. 2001b, 2004; and W.C. Brown, unpublished data). Peptides P16-7, P25, and P12-AM5 from the conserved carboxy region of MSP2 were presented to CD4+ T cells by DRA/DRB3*1201, DQA*12011/DQB*10051, and DQA*2201/DQB*1201 molecules, respectively (Fig. 2), confirming that all three MHC class II molecules were functional.
Presentation of A. marginale MSP2 peptides by DR and DQ molecules. 293-F cells were transfected with CD80 and DRA/DRB3*1201, DQA*12011/DQB*10051, or DQA*2201/DQB*1201 and tested for presentation of loaded peptide. Short‐term T–cell lines were stimulated with 293‐F cells loaded with either 0.1–10 µg/ml peptide P25 (a) or peptide P12‐AM5 (c), and CD4+ T–cell clones 61.10.2D7 (b) and 61.3B5 (d) were stimulated with 293‐F cells loaded with 0.01–10 µg/ml peptide P25 or 16-7, respectively. Results are presented as the mean CPM of triplicate cultures. Responses significantly higher than responses to peptide in the presence of non transfected 293‐F cells or 293‐F cells expressing different class II alleles are indicated with an asterisk (P<0.05)
Evidence that heterozygous BoLA haplotypes generate functional interhaplotype DQ molecules
Previous mAb-blocking studies suggested that the B. bovis Hsp20 peptide P1 (aa 11–40), recognized by cow C97, was DQ-restricted (Norimine et al. 2004). To verify this and to determine the DQ restriction element, transfected 293-F cells were pulsed with peptide P1 and tested for stimulation of Hsp20-specific CD4+ T-cell clone 3B11. Interestingly, this clone, which was shown to be derived from a single cell by sequencing the TCR α and β chains (Norimine et al. 2004), responded to peptide P1 presented by the products of two DQ allelic pairs, DQA*0301/DQB*0402 and DQA*C97/DQB*0402 (Fig. 3). This result was also obtained using CD4+ T-cell clone 3G5 (Norimine et al. 2004) specific for the same epitope (data not shown). Because cow C97 does not have a duplicated haplotype (Table 2), this finding indicates that the DQB*0402 allelic product paired with two different DQA allelic products, one provided from each haplotype (Tables 2 and 4), providing evidence for both intra- and interhaplotype pairing in the generation of functional DQA/B heterodimers.
Two DQA/B pairs present B. bovis Hsp20 peptide P1 to CD4+ T-cell clone 3B11. B. bovis Hsp20-specific CD4+ T-cell clone 3B11 was assayed with 293-F cells transfected with the indicated DRB3 and DQ alleles and CD80 and loaded with 0.1–10 μg/ml peptide P1. Responses significantly higher than responses to peptide in the presence of 293-F cells transfected the other MHC class II allelic pairs are indicated by an asterisk (P<0.05)
Numerous epitopes on A. marginale MSP1a are recognized by CD4+ T cells from MSP1-immunized cattle expressing DH16A and/or DH22H haplotypes (Brown et al. 2002). One of these, the N-terminal-repeated, 29-amino-acid-sequence peptide B, appeared to be restricted by interhaplotype-paired DQA and DQB molecules. Presentation of this epitope to T cells from heterozygous DH16A/DH22H animal 87 was blocked by DQ-specific mAb. Furthermore, this epitope was presented to CD4+ T-cell clones only by autologous APCs or APCs from a donor animal expressing the identical heterozygous BoLA haplotype DH16A/DH22H, but not by APCs from cattle with one of the two sets of parental haplotypes (Brown et al. 2002). To confirm the presentation of peptide B by an interhaplotype DQ molecule, we first examined whether MSP1a peptide B-specific T cells from animal 87 could recognize the peptide presented by APCs from animals with homozygous DH16A (animals 04B93 and 132) or DH22H (animals 132 and 201) haplotypes (Table 2). Whereas autologous APCs presented peptide B, APCs from BoLA-homozygous donors expressing either DH16A/DH16A or DH22H/DH22H haplotypes were unable to present peptide B to a CD4+ T-cell line derived from animal 87 expressing the DH16A/DH22A haplotype (Fig. 4a). These results support the previous observation that the presentation of MSP1a peptide B requires the heterozygous BoLA haplotype DH16A/DH22H, which would be necessary for generation of interhaplotype MHC class II molecules. In contrast, MSP1a peptide F3-5 (aa 330–359), also previously identified as a T-cell epitope recognized by animal 87 (Brown et al. 2002), was presented by APCs derived from DH22H-homozygous but not DH16A- homozygous animals (Fig. 4b).
Presentation of A. marginale MSP1a peptides B and F3-5 by APCs from BoLA-homozygous animals. T cells from a short-term cell line (87 CL) were cultured with 0.1–10 μg/ml peptide B (a) or peptide F3-5 (b) with autologous APCs or APCs derived from the indicated donors expressing homozygous DH16A or DH22H haplotypes. Results are presented as the mean cpm of triplicate cultures +1 SD. Responses significantly higher than responses without peptide are indicated by an asterisk (P<0.05)
To verify the use of interhaplotype-paired DQ molecules in the presentation of MSP1a peptide B, 293-F cells transfected with different BoLA DQA/B pairs were used to present the peptide. The intrahaplotype pair, DQA*22021/DQB*1301, was not tested since expression of this pair of alleles was not detected when the initial experiments were performed. As hypothesized, MSP1a peptide B was clearly presented by the interhaplotype pair encoded by DQA*2206/DQB*1301 (Fig. 5a). We also confirmed that the intrahaplotype pair encoded by DQA*2206/DQB*1402 within haplotype DH22H was functional for presenting a different peptide, F3-5 (Fig. 5b). These results indicate that, as demonstrated in vitro (Table 4), in vivo pairing of the DQA*2206 allelic product with either DQB*1301 or DQB*1402 allelic product resulted in the formation of different functional DQA/B heterodimers that primed CD4+ T cells against two different MSP1a epitopes following immunization with MSP1.
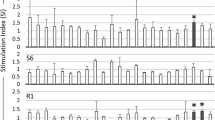
Identification of BoLA-class II restriction elements for MSP1a epitopes using DR- and DQ-transfectants. T cells from a short-term T-cell line (87 CL) were cultured for 3 days with either irradiated, autologous APCs plus 10 μg/ml of the indicated peptide or with the indicated DR- or DQ- and CD80-transfected 293-F cells loaded with 10 μg/ml of A. marginale MSP1a peptide B (a), peptide F3-5 (b), peptide F3-3 (c), or peptide F2-5B (d). Results are presented as the mean cpm of triplicate cultures +1 SD. Responses significantly higher than responses without peptide are indicated by an asterisk (P<0.005)
Additional MSP1a-derived peptides, F2-5B (aa 243–258) and F3-3 (aa 290–319), also previously shown to contain CD4+ T-cell epitopes (Brown et al. 2002), were included in the T-cell proliferation assays with MHC class II transfectants. DQA*10011/DQB*10021 was identified as encoding the restriction element for CD4+ T lymphocytes specific for MSP1a peptide F3-3 (Fig. 5c). Because this encoded DQA/B pair is present in both DH16A and DH22H haplotypes (Tables 2 and 4), animal 87 (DH16A/DH22H) might have an increased frequency of memory CD4+ T lymphocytes restricted by DQ molecule. The relatively high level of T-cell proliferation to peptide F3-3 supports this possibility. Finally, DRB3*1101 was identified as encoding the restriction element for MSP1a peptide F2-5B (Fig. 5d), which again is consistent with previous studies showing that a DRB3 product linked to RFLP 22 presented this peptide to CD4+ T-lymphocyte clones (Brown et al. 2002).
Comparison of DR vs DQ presentation of epitopes for the proteins studied
The results of antigen presentation by BoLA-DQ or BoLA-DR are summarized in Table 5. A total of 11 unique restriction elements were identified in these studies; four encoded by DR-AB allelic pairs and seven encoded by DQ-AB allelic pairs. Furthermore, five different peptides were presented by DR molecules, and six different peptides were presented by DQ molecules. These results indicate that, at least among the proteins studied, comparable numbers of epitopes were presented by BoLA-DR and BoLA-DQ.
Discussion
This study has definitively shown functional BoLA-DQA/B heterodimers formed by interhaplotype pairing of DQA and DQB molecules. Interhaplotype pairing of DQA and DQB chains, together with duplication of BoLA-DQ genes, intrahaplotype DQA and DQB chain pairing, and polymorphism in BoLA-DRB3 and BoLA-DQ genes, serves to increase the complexity of restriction element usage in cattle, which may influence the immunological outcome during infection (Lewin et al. 1999; Ellis and Ballingall 1999; Glass et al. 2000; Park et al. 2004;). In mice, mixed-haplotype H-2 molecules play an important role in causing autoimmune symptoms or a dominant immune response to hen egg lysozyme (HEL) (Moreno et al. 1990; Gotoh et al. 1993; Nygard et al. 1993). Strikingly, in (H-2k×H-2b)F1 mice, 86% of T-cell hybridomas specific for HEL were restricted by mixed haplotypes, either I-Aαkβb or I-Aαbβk (Moreno et al. 1990). This observation is probably relevant to antigen presentation by HLA- and BoLA-DQ molecules because most humans and cattle have heterozygous haplotypes and both DQA and DQB chains are highly polymorphic, similar to mouse I-A molecules.
In cattle, duplication of DQ molecules provides an increased opportunity for inter- and intrahaplotype pairing of A and B chains to form functional DQ heterodimers. We identified the product of the interhaplotype DQ molecule, DQA*2206/DQB*1301, as the MHC class II restriction element for MSP1a peptide B. We also found that the product of the intrahaplotype-matched allelic pair, DQA*2206/DQB*1402, was functional in presenting MSP1a peptide F3-5. This indicates that the DQA*2206 allelic product plays a role in presenting at least two different CD4+ T-cell epitopes derived from MSP1a by pairing with DQB allelic products derived from the same and different haplotypes. In vitro expression of DQA*2206/DQB*10021 in 293-F cells (Table 4) further suggests that the DQA*2206 allele is flexible to pair with multiple DQB alleles and may play a dominant role for helper T-cell responses in animals carrying the DH22H haplotype. Further, we have identified functional DQ molecules, DQA*10011/DQB*10021, DQA*12011/DQB*10051, and DQA*2201/DQB*1201, which present peptides MSP1a F3-3, MSP2 P25, and MSP2 P12-AM5, respectively. These functional DQ pairs are most likely intrahaplotype DQ molecules consisting of products of adjacent loci according to our linkage data. In this study, functional intrahaplotype DQ molecules consisting of products of nonadjacent loci were not identified, although this type of intrahaplotype DQ molecule was expressed on 293-F cells in some combinations. This is likely explained by the limited numbers of antigens tested in our study, and more extensive analysis may reveal that this type of intrahaplotype DQ molecule is also functional.
It is of interest that some DQA and DQB combinations were not detected on transfected 293-F cells. Preferential pairing of DQ chains has also been reported for humans and mice. With human cells, interchange between DQA and DQB chains occurs within the DQw1 family of haplotypes, but not between this family and the other non-DQw1-associated haplotypes, whereas DQw2-, 3-, and 4-associated haplotypes are freely interchangeable (Kwok et al. 1993). As in sheep, bovine class II DQA*10011 and DQA*12011 alleles are grouped in the DQA1 cluster, while DQA*22021, DQA*2206, and DQA*2201 alleles are grouped in the DQA2 cluster (Hickford et al. 2004; Chris Davies, personal communication). Our observation that products of DQA*10011 and DQA*12011 alleles paired with only one DQB allelic product is consistent with the possibility that DQA alleles within the DQA1 cluster may have more limited structural constraints for pairing. However, additional studies are needed to confirm this possibility.
Additional evidence for functional interhaplotype pairing of DQA and DQB proteins is the observation that two CD4+ T-cell clones specific for B. bovis Hsp20 peptide P1 were restricted by products of both DQA*0301/DQB*0402 and DQA*C97/DQB*0402 pairs. Because duplicated DQ alleles were not identified in this donor animal (C97), one of these allelic pairs appears to result from interhaplotype pairing. Unfortunately, the Brahman–Angus cross cow C97 expresses a rare BoLA haplotype, so it was not possible to prove this using haplotype homozygous APCs. However, this finding is similar to the report that some T-cell hybridomas specific for HEL in (H-2k×H-2b) F1 mice were also restricted by two different I-A molecules, I-Aαbβk and I-Aαkβk (Moreno et al. 1990). The ability of Hsp20-specific T-cell clones to respond to peptide presented by two different DQA/B heterodimers may be attributed to the high degree of homology between the two DQα chains, which are 94% identical in the amino acid sequence encoded by exon 2.
In humans, HLA-heterozygosity has drawn attention largely from association of HLA haplotype with immunologically mediated disease (McDevitt and Bodmer 1972; Todd et al. 1987; Nepom and Erlich 1991). For example, people with HLA-DQ2 (DQA1*05/DQB1*02) are strongly predisposed to celiac disease, and DQ2 can be generated by interhaplotype or intrahaplotype pairing (Sollid et al. 1989; Sollid 2002). Interestingly, a dose effect of HLA-DQ2 expression appears to confer susceptibility to developing celiac disease depending on the homozygous or heterozygous status of specific haplotypes containing DQA1*05 and DQB1*02 alleles (Vader et al. 2003). These reports indicate that MHC class II DQ molecules generated by interhaplotype A/B chain pairing may occur rather commonly in outbred species like humans and cattle, where most individuals express heterozygous haplotypes. Although association of diseases with BoLA haplotypes is largely unknown, several lines of evidence indicate that certain BoLA haplotypes are associated with resistance to persistent lymphocytosis caused by bovine leukemia virus infection (Lewin 1989; van Eijk et al. 1992b; Xu et al. 1993; Nagaoka et al. 1999). Furthermore, an association between mastitis and BoLA-class II haplotypes was observed (Park et al. 2004; Lundén et al. 1990; Starkenburg et al. 1997; Sharif et al. 1998), and two BoLA-DRB3 alleles were significantly associated with protection against Theileria parva challenge (Ballingall et al. 2004). However, in spite of the clear indication that BoLA-DR and BoLA-DQ molecules may play a role in protective immunity or susceptibility to infectious disease, functional analysis of BoLA-class II has rarely been reported.
In summary, using a transient class II and CD80/CD86 expression system, this study provides evidence that BoLA-DQ molecules contribute to priming helper T cells against epitopes of important pathogens of cattle such as B. bovis and A. marginale by generating DQA/B heterodimers formed by interhaplotype and intrahaplotype pairing. These results and those of previous studies (Brown et al. 2001b, 2002, 2003, 2004; Norimine et al. 2002, 2004) indicate that CD4+ T-cell responses against the proteins examined are not skewed toward DQ-restriction as observed in FMDV-immunized animals (Glass et al. 2000), but consist of both DR- and DQ-restricted responses. Nevertheless, it appears that some alleles play more dominant roles in antigen presentation than others. If protective immune responses against bovine pathogens are strongly influenced by certain MHC class II alleles, characterization of the role of BoLA-DQ is extremely important for effective vaccine development or breeding strategies.
References
Abbas AK, Lichtman AH, Pober JS (2000) The major histocompatibility complex. In: Cellular and molecular immunology, 4th edn. WB Saunders, Philadelphia, pp 63–78
Agrewala JN, Wilkinson RJ (1998) Differential regulation of Th1 and Th2 cells by p. 91–110 and p. 21–40 peptides of the 16kD α-crystallin antigen of Mycobacterium tuberculosis. Clin Exp Immunol 114:392–397
Allred DR, McGuire TC, Palmer GH, Leib SR, Harkins TM, McElwain T, Barbet AF (1990) Molecular basis for surface antigen size polymorphisms and conservation of a neutralization-sensitive epitope in Anaplasma marginale. Proc Natl Acad Sci U S A 87:3220–3224
Ballingall KT, Luyai A, Rowlands GJ, Sales J, Musoke AJ, Morzaria SP, McKeever DJ (2004) Bovine leukocyte antigen major histocompatibility complex class II DRB3*2703 and DRB3*1501 alleles are associated with variation in levels of protection against Theileria parva challenge following immunization with the sporozoite p67 antigen. Infect Immun 72:2738–2741
Bissumbhar B, Nilsson PR, Hensen EJ, Davis WC, Joosten I (1994) Biochemical characterization of bovine MHC DQ allelic variants by one-dimensional isoelectric focusing. Tissue Antigens 44:100–109
Brown WC, Logan KS, Wagner GG, Tetzlaff CL (1991) Cell-mediated immune responses to Babesia bovis antigens in cattle following infection with tick-derived or cultured parasites. Infect Immun 59:2418–2426
Brown WC, Shkap V, Zhu D, McGuire TC, Tuo W, McElwain TF, Palmer GH (1998) CD4+ T-lymphocyte and immunoglobulin G2 responses in calves immunized with Anaplasma marginale outer membranes and protected against homologous challenge. Infect Immun 66:5406–5413
Brown WC, Palmer GH, Lewin HA, McGuire TC (2001a) CD4+ T lymphocytes from calves immunized with Anaplasma marginale major surface protein 1 (MSP1), a heteromeric complex of MSP1a and MSP1b, preferentially recognize the MSP1a carboxyl terminus that is conserved among strains. Infect Immun 69:6853–6862
Brown WC, McGuire TC, Zhu D, Lewin HA, Sosnow J, Palmer GH (2001b) Highly conserved regions of the immunodominant major surface protein 2 (MSP2) of the ehrlichial pathogen Anaplasma marginale are rich in naturally derived CD4+ T lymphocyte epitopes that elicit strong recall responses. J Immunol 166:1114–1124
Brown WC, McGuire TC, Mwangi W, Kegerreis KA, Macmillan H, Lewin HA, Palmer GH (2002) Major histocompatibility complex class II DR-restricted memory CD4+ T lymphocytes recognize conserved immunodominant epitopes of Anaplasma marginale major surface protein 1a. Infect Immun 70:5521–5532
Brown WC, Brayton KA, Styer CM, Palmer GH (2003) The hypervariable region of Anaplasma marginale major surface protein 2 (MSP2) contains multiple immunodominant CD4+ T lymphocyte epitopes that elicit variant-specific proliferative and IFN-γ responses in MSP2 vaccinates. J Immunol 170:3790–3798
Brown WC, Palmer GH, Brayton KA, Meuss PF, Barbet AF, Kegerreis KA, McGuire TC (2004) CD4+ T lymphocytes from Anaplasma marginale major surface protein 2 (MSP2) vaccinates recognize naturally processed epitopes conserved in MSP3. Infect Immun 72:3688–3692
Calvo-Calle JM, Hammer J, Singaglia F, Clavijo P, Moya-Castro ZR, Nardin EH (1997) Binding of malaria T cell epitopes to DR and DQ molecules in vitro correlates with immunogenicity in vivo. J Immunol 159:1362–1373
Davies CJ, Joosten I, Andersson L, Arriens MA, Bernoco D, Bissumbhar B, Byrns G, van Eijk MJT, Kristensen B, Lewin HA, Mikko S, Morgan ALG, Muggli-Cockett NE, Nilsson PhR, Oliver RA, Park CA, van der Poel JJ, Polli M, Spooner RL, Stewart JA (1994) Polymorphism of bovine MHC class II genes. Joint report of the Fifth International Bovine Lymphocyte (BoLA) Workshop, Interlaken, Switzerland, 1 August 1992. Eur J Immunogenet 21:259–289
Davis WC, Marusic S, Lewin HA, Splitter GA, Perryman LE, McGuire TC, Gorham JR (1987) The development and analysis of species specific and cross reactive monoclonal antibodies to leukocyte differentiation antigens and antigens of the major histocompatibility complex for use in the study of the immune system in cattle and other species. Vet Immunol Immunopathol 15:337–376
Ellis SA, Ballingall KT (1999) Cattle MHC: evolution? Immunol Rev 167:159–168
Endl J, Otto H, Jung G, Driesbusch B, Donie F, Stahl P, Elbracht R, Schmitz G, Meinl E, Hummel M, Ziegler AG, Wank R, Schendel DJ (1997) Identification of naturally processed T cell epitopes from glutamic acid decarboxylase presented in the context of HLA-DR alleles by T lymphocytes of recent onset IDDM patients. J Clin Invest 99:1415–2405
Fogg MH, Parsons KR, Thomas LH, Taylor G (2001) Identification of CD4+ T cell epitopes on the fusion (F) and attachment (G) proteins of bovine respiratory syncytial virus (BRSV). Vaccine 19:3226–3240
Fraser DC, Craigmile S, Campbell JDM, Oliver RA, Brown DJ, Russell GC, Spooner RL, Glass EJ (1996) Functional expression of a cattle MHC class II DR-like antigen on mouse L cells. Immunogenetics 43:296–303
Glass EJ, Oliver RA, Russell GC (2000) Duplicated DQ haplotypes increase the complexity of restriction element usage in cattle. J Immunol 165:134–138
Gotoh Y, Takashima H, Noguchi K, Nishimura H, Tolushima M, Shirai T, Kimoto M (1993) Mixed haplotype Aβz/Aαd class II molecule in (NZB x NZW)F1 mice detected by T cell clones. J Immunol 150:4777–4787
Hickford JGH, Zhou H, Slow S, Fang Q (2004) Diversity of the ovine DQA2 gene. J Anim Sci 82:1553–1563
Howard CJ, Sopp P, Brownlie J, Kwong L-S, Parsons KR, Taylor G (1997) Identification of two distinct populations of dendritic cells in afferent lymph that vary in their ability to stimulate T cells. J Immunol 159:5372–5382
Kwok WW, Kovats S, Thurtle P, Nepom GT (1993) HLA-DQ allelic polymorphisms constrain patterns of class II heterodimer formation. J Immunol 150:2263–2272
Kwok WW, Liu AW, Novak EJ, Gebe JA, Ettinger RA, Nepom GT, Reymond SN, Koelle DM (2000) HLA-DQ tetramers identify epitope-specific T cells in peripheral blood of herpes simplex virus type 2-infected individuals: direct detection of immunodominant antigen-responsive cells. J Immunol 164:4244–4249
Lewin HA (1989) Disease resistance and immune response genes in cattle: strategies for their detection and evidence of their existence. J Dairy Sci 72:1334–1348
Lewin HA, Russell GC, Glass EJ (1999) Comparative organization and function of the major histocompatibility complex of domesticated cattle. Immunol Rev 167:145–158
Lundén A, Sigurdardóttir S, Edfors-Lilja I, Danell B, Rendel J, Andersson L (1990) The relationship between bovine major histocompatibility complex class II polymorphism and disease studied by use of bull breeding values. Anim Genet 21:221–232
McDevitt HO, Bodmer WF (1972) Histocompatibility antigens, immune responsiveness and susceptibility to disease. Am J Med 52:1–8
Mølvig J, Sonderstrup McDevitt G, Thomsen M, Bossmann B, Blangero C, de Preval C, Baek L, Nerup J (1989) Characterization of PPD specific T cell lines generated in type 1 (insulin dependent) diabetic and healthy individuals. Scand J Immunol 30:615–629
Moreno J, Adorini L, Hämmerling GJ (1990) Co-dominant restriction by a mixed-haplotype I-A molecule (αkβb) for the lysozyme peptide 52-61 in H-2k×H-2b F1 mice. J Immunol 144:3296–3304
Nagaoka Y, Kebaya H, Onuma M, Kasai N, Okada K, Aida Y (1999) Ovine MHC Class II DRB1 alleles associated with resistance or susceptibility to development of bovine leukemia virus-induced ovine lymphoma. Cancer Res 59:975–981
Nepom GT, Erlich H (1991) MHC class-II molecules and autoimmunity. Annu Rev Immunol 9:493–525
Norimine J, Suarez CE, McElwain TF, Florin-Christensen M, Brown WC (2002) Immunodominant epitopes in Babesia bovis rhoptry-associated protein 1 that elicit memory CD4+ T-lymphocyte responses in B. bovis-immune individuals are located in the amino-terminal domain. Infect Immun 70:2039–2048
Norimine J, Mosqueda J, Suarez CE, Palmer GH, McElwain TF, Mbassa G, Brown WC (2003) Stimulation of T-helper cell gamma interferon and immunoglobulin G responses specific for Babesia bovis rhoptry-associated protein 1 (RAP-1) or a RAP-1 protein lacking the carboxy-terminal repeat region is insufficient to provide protective immunity against virulent B. bovis challenge. Infect Immun 71:5021–5032
Norimine J, Mosqueda J, Palmer GH, Lewin HA, Brown WC (2004) Conservation of Babesia bovis small heat shock protein (Hsp20) among strains and definition of T helper cell epitopes recognized by cattle with diverse major histocompatibility complex class II haplotypes. Infect Immun 72:1096–1106
Nygard NR, McCarthy DM, Schiffenbauer J, Schwartz BD (1993) Mixed haplotypes and autoimmunity. Immunol Today 14:53–56
Palmer GH, McGuire TC (1984) Immune serum against Anaplasma marginale initial bodies neutralizes infectivity for cattle. J Immunol 133:1010–1015
Park YH, Joo YS, Park JY, Moon JS, Kim SH, Kwon NH, Ahn JS, Davis WC, Davies CJ (2004) Characterization of lymphocyte subpopulations and major histocompatibility complex haplotypes of mastitis-resistant and susceptible cows. J Vet Sci 5:29–39
Russell GC, Fraser DC, Craigmile S, Oliver RA, Dutia BM, Glass EJ (2000) Sequence and transfection of BoLA-DRB3 cDNAs. Anim Genet 31:219–222
Schreuder GMT, Hurley CK, Marsh SGE, Lau M, Fernandez-Vina MA, Noreen HJ, Setterholm M, Maiers M (2005) HLA dictionary 2004: summary of HLA-A, -B, -C, -DRB1/3/4/5, -DQB1 alleles and their association with serologically defined HLA-A, -B, -C, -DR, and -DQ antigens. Hum Immunol 66:170–210
Sharif S, Mallard BA, Wilkie BN, Sargeant JM, Scott HM, Dekkers JCM, Leslie KE (1998) Associations of the bovine major histocompatibility complex DRB3 (BoLA-DRB3) alleles with occurrence of disease and milk somatic cell score in Canadian dairy cattle. Anim Genet 29:185–193
Sollid LM (2002) Coeliac disease: dissecting a complex inflammatory disorder. Nat Rev Immunol 2:647–655
Sollid LM, Markussen G, Ek J, Gjerde H, Vartdal F, Thorsby E (1989) Evidence for a primary association of celiac disease to a particular HLA-DQ α/β heterodimer. J Exp Med 169:345–350
Starkenburg RJ, Hansen LB, Kehrli ME Jr, Chester-Jones H (1997) Frequencies and effects of alternative DRB3.2 alleles of bovine lymphocyte antigen for Holsteins in milk selection and control lines. J Dairy Sci 80:3411–3419
Todd JA, Bell JI, McDevitt HO (1987) HLA-DQβ gene contributes to susceptibility and resistance to insulin-dependent diabetes mellitus. Nature 329:599–604
Vader W, Stepniak D, Kooy Y, Mearin L, Thompson A, van Rood JJ, Koning L (2003) The HLA-DQ2 gene dose effect in celiac disease is directly related to the magnitude and breadth of gluten-specific T cell responses. Proc Natl Acad Sci U S A 100:12390–12395
van Eijk MJT, Stewart-Haynes JA, Lewin HA (1992a) Extensive polymorphism of the BoLA-DRB3 gene distinguished by PCR–RFLP. Anim Genet 23:483–496
van Eijk MJT, Stewart-Haynes JA, Beever JE, Fernando RL, Lewin HA (1992b) Development of persistent lymphocytosis in cattle is closely associated with DRB2. Immunogenetics 37:64–68
Xu A, van Eijk MJT, Park C, Lewin HA (1993) Polymorphism in BoLA-DRB3 exon 2 correlates with resistance to persistent lymphocytosis caused by bovine leukemia virus. J Immunol 151:6977–6985
Acknowledgements
We thank Kimberly Kegerreis, Shelley Whidbee, and Emma Karel for excellent technical assistance, Dr. Christopher J. Davies for information of BoLA-DQ expression in cattle with homozygous haplotypes and for helpful comments on the manuscript, and Dr. John Swain for providing blood samples from calves at the WSU Dairy. This work was supported by grants R01-AI30136, R01-AI44005, and R01-AI053692 from the National Institute of Allergy and Infectious Diseases, National Institutes of Health (NIH) and by the US Department of Agriculture (USDA) Cooperative Agreement 58-5348-044. These experiments are in compliance with USDA and NIH guidelines for vertebrate animal use.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Norimine, J., Brown, W.C. Intrahaplotype and interhaplotype pairing of bovine leukocyte antigen DQA and DQB molecules generate functional DQ molecules important for priming CD4+ T-lymphocyte responses. Immunogenetics 57, 750–762 (2005). https://doi.org/10.1007/s00251-005-0045-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-005-0045-6