Abstract
Energetic properties of a protein are a major determinant of its evolutionary fitness. Using a reconstruction algorithm, dating the reconstructed proteins and calculating the interaction network between their amino acids through a coevolutionary approach, we studied how the interactions that stabilise 890 proteins, belonging to five families, evolved for billions of years. In particular, we focused our attention on the network of most strongly attractive contacts and on that of poorly optimised, frustrated contacts. Our results support the idea that the cluster of most attractive interactions extends its size along evolutionary time, but from the data, we cannot conclude that protein stability or that the degree of frustration tends always to decrease.
Similar content being viewed by others
Introduction
The evolutionary fitness of a protein is tightly related to its energetic properties. An important determinant of the evolutionary fitness of structured proteins is their thermodynamic stability. The stability requirement imposes a constraint to viable mutations and acts in a non-trivial, cooperative way at the level of the whole organism (Zeldovich et al. 2007; Rodrigues et al. 2016).
The stability of ancient proteins has been widely studied experimentally by sequence reconstruction, expressing and analysing ancient sequences obtained from extant protein families through maximum-likelihood or Bayesian methods (Wheeler et al. 2016). In general, ancient proteins are more stable than modern ones (Gaucher et al. 2008; Perez-Jimenez et al. 2011; Carstensen et al. 2012; Risso et al. 2013; Akanuma et al. 2013), although counterexamples exist (Hart et al. 2014). This behaviour is usually rationalised in terms of a higher environmental temperature in the Precambrian era (Boussau et al. 2008; Wheeler et al. 2016). Moreover, simple protein models suggest that the evolution of thermodynamic stability is controlled by a cluster of strongly stabilising residues that evolves in a slower way with respect to the others (Tiana et al. 2000, 2004a, b).
Another requirement that affects protein fitness is the kinetic accessibility of the native state, because a slow folding rate would increase the risk of misfolding and aggregation. Since proteins are frustrated systems, namely unfavourable interactions linger in their energy ground state (Ferreiro et al. 2014), they would be expected to display slow, non-exponential kinetics (Bryngelson and Wolynes 1989). It was suggested that evolution minimises the degree of frustration of proteins to avoid kinetic traps (Bryngelson and Wolynes 1987). Even in the absence of consistent frustration, the folding process is regulated by an entropic barrier that determines folding rate. Such a barrier is usually overcome by nucleation of specific parts of the protein chain (Wetlaufer 1973; Abkevich et al. 1994). Both the folding nucleus (Mirny and Shakhnovich 2001) and the folding rate (Tzul et al. 2017) are usually highly conserved along evolutionary time.
The goal of the present work is to study the network of interactions between amino acids in the native state of reconstructed ancient proteins. In particular, we focused on the evolution of the network of strongly attractive two-body contacts, which stabilises their native state, and on frustrated contacts, which affect folding rates; the main intent is to study how contacts evolve as a function of evolutionary time. Understanding the evolution of the elements which stabilise proteins can be relevant both for fundamental reasons, for designing new proteins and for predicting the microbial resistance to drugs and vaccines (Russ et al. 2020).
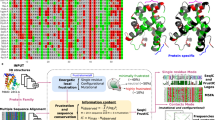
A key problem in pursuing this goal is the quantification of the interaction energies between amino acids. The majority of classical force fields are atom-based and usually require an explicit description of the dynamics of the solvent, thus are not easy to use for our purposes. We then chose to describe the interaction between amino acids with a 2-bodies potential whose parameters are obtained from correlations between mutations in alignments of extant homologous sequences (Morcos et al. 2011). In brief, the sequences of homologous sequences are regarded as equilibrium realisations of a Potts model of unknown parameters. Using techniques of inverse statistics, one can look for the best choice of the interaction energies that are compatible with the empirical correlation functions (Nguyen et al. 2017). Different approximations can be employed to implement the inversion (Morcos et al. 2011, 2013; Ekeberg et al. 2013; Figliuzzi et al. 2015; Cuturello et al. 2020), which anyway perform similarly to each other in obtaining the interaction energies (Franco et al. 2019). For this reason, we employed the original mean-field procedure (Morcos et al. 2011) because of its computational efficiency. This strategy of calculation of the interactions proved efficient in predicting the native conformation protein monomers (Morcos et al. 2011) and dimers (dos Santos et al. 2015), of their conformational fluctuations (Jana et al. 2014; Sutto et al. 2015), to study protein aggregation (Tian et al. 2015; Kassem et al. 2018), the effect of mutations in protein stability (Lui and Tiana 2013; Contini and Tiana 2015), the identification of protein domains (Halabi et al. 2009; Granata et al. 2017) and the identification of interaction hotspots in transmembrane proteins (Baldessari et al. 2020). The use of coevolutionary data to simulate protein evolution (de la Paz et al. 2020) was also useful to investigate the details of neutral evolution theory.
In this work, we analysed five protein families, reconstructing their evolution and calculating the interaction network in every extant and reconstructed molecule, for a total of 890 proteins. We then analysed how the interaction network depends on evolutionary time.
Calculation of interaction energies along evolution
We studied the evolution of the energetic properties of β-lactamase (BLM), thioredoxin (TRD), nucleoside-diphosphate kinase (NDK), cytochrome c (CYC) and ribonuclease H (RDH). These are well-characterised protein families, they are evolutionarily quite old and contain a large number of sequences.
From the alignment of the extant proteins in each family, we calculated the interaction tensor εij(σ, ρ) with the mean-field approach of ref. (Morcos et al. 2011), which is remarkably fast. The key idea is that pairs of residues which are close in space and that attract strongly each other undergo highly correlated mutations (while each of the two residues are not necessarily more conserved, see below). Inverting the problem, most correlated residues are expected to be close in space and strongly interacting, and this interaction can be quantified within an inverse Potts model (Nguyen et al. 2017).
Operatively, the “full” alignments are obtained from the Pfam database (Punta et al. 2012), those with less than 30% gaps are retained and those with sequence identity larger than 70% were down-weighted as in ref. (Morcos et al. 2011). For all families, there are more than 104 sequences (cf. Table 1). We calculated one- and two-point frequencies using pseudocounts on the overall fraction of residues types (x = 0.5), on the fraction of residues types in the alignment (y = 0.1) and on the fraction of residues type in the specific position (z = 1.0) as in ref. (Lui and Tiana 2013).
Energies are expressed in units of the evolutive temperature (Shakhnovich and Gutin 1993a), which cannot be determined within the model and which does not relate straightforwardly to the environmental temperature, being expected to be smaller than that (Morcos et al. 2014). They are gauged setting to zero the interaction of the gap, regarded as the 21st type of residue, with all the others (Lui and Tiana 2013). The (N × N × 21 × 21)-tensor contains the contact energies between all pair of sites for any kind of amino acid that they can host.
The sequence of ancient proteins of each family is reconstructed through a maximum-likelihood scheme with PAML (Yang 2007) from the Pfam alignment and from the phylogenetic tree that defines the links between proteins over time (see, e.g. Fig. S1 in the Supp. Mat.). We selected a subset of extant sequences from the Pfam alignment that have mutually less than 20% of gaps (cf. Table 1), selecting a single protein per organism. When multiple proteins are associated with the same organism, that with minimum number of gaps is preferred. We then built a tree that defines the relationship between the selected organisms which host the proteins from TimeTree (Hedges et al. 2015), that also gives an estimate of the age of each reconstructed ancient sequence. The sequences of the proteins corresponding to the nodes of the tree, that is the ancestors of extant proteins belonging to the Pfam dataset, are then reconstructed with PAML. The resulting alignment is free of gaps.
For any sequence of a family, putative native conformations are predicted by homology modelling using Modeller (Webb and Sali 2016). Homologs of known structure are selected with E value < 0.01, giving a variable number of templates, usually between 1 and 5. Subsequently, the obtained structure is optimised through a short minimization run with Gromacs (Van Der Spoel et al. 2005) using the Amber99SB force field.
The energy tensor is then filtered, setting to zero the elements that are not in contact in the (crystallographic or putative) native structure. Two residues are assumed to be in contact if their Cβ (Cα in case of glycine) are closer than 6.5 Å.
As a consequence of this procedure, the energy tensor εij(σ, ρ) is the same for all proteins of each family (but in general different between different families); on the other hand, the projection of the four-dimensional tensor on the specific sequence to obtain the two-dimensional interaction matrix εij between its amino acids depends on the specific protein.
The distribution of the native contact energies between residues in all proteins belonging to each family is displayed in the upper-left panel of Fig. 1. It displays a sharp peak centred in zero and a long tail towards negative values. The distribution of energies over all sequences of a family is similar to that of single sequences (cf. Fig. S2 in the Supp. Mat.), so it can be regarded as representative of any sequence.
a With solid lines, the distribution of the interaction energies εij for the native contact of all family members of β-lactamase (BLM), thioredoxin (TRD), nucleoside diosolphate kinase (NDK), Cytochrome c (CYC) and Ribonuclease H (RDH). Dashed lines indicate the energies obtained from a random bootstrap of the sequences. b–f the native energy EN of the extant and reconstructed proteins of each family as a function of evolutionary time; the continuous line is the linear fit, while the dashed line is a horizontal reference
In Fig. 1a, it is also displayed the distribution of energies associated with a null model, obtained from a bootstrap procedure in which the residues at each position are randomly reshuffled among the sequences (thus keeping one-site frequencies unchanged).
While the distribution of energies in the null model is rather symmetric, Gaussian-like and centred around negative values, that of protein energies displays a long tail towards negative elements, stemming from a sharp peak centred close to zero. Energies in proteins seem, thus, much more polarised than in the null model. On the side of positive energies, the distributions associated with the five protein families do not display any tail but a decay similar to that of the null model.
Considering that the distribution displayed in Fig. 1a is limited to interactions that are in contact in the native conformation, its shape supports the idea that proteins are stabilised by a core of strong interactions (those belonging to the negative tail of the distribution), that constrain the rest of weakly interacting residues, corresponding to the peak around zero (Tiana et al. 1998; Mirny and Shakhnovich 1999). This shape is also consistent with the asymmetrical distribution of mutational energies obtained for several proteins (Tokuriki et al. 2007).
Thermodynamic stability of ancestral proteins
The stability of a protein is essentially determined by the energy of its native state, because the competing, denatured states are self-averaging, i.e. their thermodynamic properties do not depend on the detailed sequence (Shakhnovich and Gutin 1993a, b). In Fig. 1, we plotted the native energies EN of extant and reconstructed proteins as a function of evolutionary time. This quantity is calculated simply summing all the coevolutionary energies of pairs of residues that are in contact; the lower the value of EN, the more stable is the protein.
It should be noted that the self-averaging character of the denatured state can be guaranteed only if the composition of the protein in terms of type of amino acids, especially in terms of hydrophobic residues, remains constant. The reconstructed proteins display a rather constant composition (cf. Fig. S3 in the Supp. Mat.) and, thus, comply with this requirement.
The trend of EN appears as system dependent. In the case of BLM, CYC and RNH, stability decreases towards recent proteins. In this case, the slope obtained from a linear fit of the energies as a function of time is significantly larger than that of a null model obtained from a random bootstrap of the calculated energies; also Kendall’s tau test indicates significant monotonicity (cf. the p values in Table 2). Also a different null model in which we calculate the energies of a random set of sequences gives in all cases a constant temporal trend, with standard deviation on the slope < 10–5 My−1 (cf., e.g. Fig. S4 in the Supp. Mat.).
In the case of NDK, the decreasing values of EN indicate a significant increase of stability with time; while for TRD, we cannot spot any monotonic behaviour. Anyway, for all the considered proteins, the variability of EN is quite large (cf. the standard deviation in Table 2), even at similar times. The variability within different kingdoms is the same as that in the overall set of proteins (cf. Fig. S5 in the Supp. Mat.).
The overall tendency of proteins to destabilise towards the present age (but with several exceptions) has been already recognised (Wheeler et al. 2016). This tendency was explained either as a selective advantage of marginally stable proteins in terms of adapting to new functions (Bloom et al. 2004) or as an entropic effect in sequence space (Taverna and Goldstein 2002). However, at variance with our findings, the reconstructed proteins belonging to the TRD and NDK families display decreasing stability as measured by differential scanning calorimetry (Perez-Jimenez et al. 2011) and circular dichroism (Akanuma et al. 2013), respectively. One should consider that these conclusions are drawn by the analysis of 7 and 12 proteins, respectively, which is a small subset of the 187 and 305, respectively, considered in our analysis. For example, the analysis of the of the TRD proteins reconstructed in ref. (Perez-Jimenez et al. 2011) shows that the experimental denaturation temperature is negatively correlated with the predicted native-state energy (cf. Fig. 2), as expected for the equilibrium of a two-state system, and thus, the model predicts for these seven proteins a decreasing stability along evolutionary time. This means not only that the model is able to predict the thermodynamic properties of proteins characterised by calorimetry, but also that selecting few proteins one can observe a trend that is different from that of the whole set. Note that filtering only the energies associated with native contacts is useful, because not doing it leads to an unphysical positive correlation between denaturation temperature and native energy (cf. Fig. S6).
The denaturation temperatures Tm of the seven reconstructed TRD proteins of ref. (Perez-Jimenez et al. 2011) as a function of the native energy EN calculated with the present model. The labelling corresponds to that of the referenced article. The dashed line indicates a linear fit, expected for a two-state model. The correlation coefficient is − 0.63. The increase of energy with respect to time has a slope of 0.6 My−1
Evolution of strongly attractive contacts
We defined operatively “strongly attractive” contacts as those with energy εij below a threshold εth such that globally 5% of the contacts of the null models of the five proteins lie below εth. Using the energies displayed in Fig. 1, we found that εth = − 0.57.
The fraction a of strongly attractive contacts of extant and reconstructed proteins is displayed in Fig. 3. The value of a is significantly increasing for TRD and CYC (cf. Table 3). A linear fit of a as a function of time gives for these two proteins an increase rate of the order of 10–5 year−1. The p values obtained calculating the slopes of randomly bootstrapped data are 0.002 and 0.001, respectively. We also computed the p values associated with the null hypothesis that the increase in a is not monotonic, using Kendall’s tau. The monotonicity of a is significant as well (cf. Table 3).
For the other three protein families (BLM, NDK and RNH), no statistically significant trend could be identified.
Interaction network analysis
We then analysed the network whose nodes are all the amino acids of each protein and whose links are the strongly attractive contacts (see, e.g. upper-left panel of Fig. 4). All networks display one, or few, large clusters and several orphans (i.e., nodes without links). The largest-cluster size (LCS) is in all cases significantly smaller than that of randomly generated proteins (p value < 10–6 and green points in Fig. 4). LCS increases from more ancient to more recent proteins (see blue points in Fig. 4) for all families in a statistically significant way, as calculated from a random bootstrap of the energies of reconstructed proteins (see Table 3). Also, the comparison with randomly generated sequences give p values < 10–6 (see Fig. S7 in the Supp. Mat.). The number of orphans and the clustering coefficient are significantly larger than those expected from random networks, but they do not display a regular temporal trend (cf. Fig. S8 in the Supplementary Material). These data agree with the literature that all studied proteins display a core of contacts (Mirny and Shakhnovich 1999; Tokuriki et al. 2007) that strongly stabilise the native state (often, but not always hydrophobic) and suggest that, although the total number of strongly attractive contacts does not always increase along evolution, the set of strongly stabilising residues does.
The upper-left panel is an example of network of strongly attractive interactions of BLM. The other panels display (in blue) the largest-cluster size (LCS), normalised to the total number of nodes, of extant and reconstructed proteins. Green points indicate the mean LCS of proteins with randomly reshuffled strongly attractive contacts
The above results are quite robust with respect to the threshold εth used to define the strongly attractive contacts (cf., e.g. Fig. S9 in the Supp. Mat.).
The evolution of the position of the amino acids involved in the strongly attractive contacts can be found in Fig. 5, that displays the sum Ei of attractive interactions for each site i. One can notice that strongly interacting sites (whatever are the residues hosted there) are rather conserved. Those present in ancient proteins tend to remain in extant proteins and sometimes new ones are added during evolution.
Interestingly, the correlation between strongly interacting and highly conserved residues in the alignment is poor (cf. Fig. S10 in the Supp. Mat.). In particular, there are many more highly conserved sites than strongly interacting sites, suggesting that there could be several reasons why residues are evolutionary conserved (Mirny and Shakhnovich 1999).
Evolution of frustrated contacts
Frustration is the property of some complex systems not to be able to get rid of unfavourable interactions even in the ground state. As discussed by Phil Anderson in ref. (Anderson 1978), it is not straightforward to quantify if a system is frustrated if not for spin systems. His suggestion was to identify the ground state of the system, to partition it into subsystems and to quantify the scaling of the interaction energy between them as a function of the area of the separation surface. For finite-range interactions, as those acting between amino acids, the scaling is expected to be linear if the system is ferromagnetic-like. If it is frustrated, one expects a sub-linear scaling because of the compensation between attractive and repulsive interactions.
In the upper-left panel of Fig. 6 it is displayed, as an example, the square of the interfacial energy between the segments of various length L of a BLM and the rest of the protein. The fact that the mean square energy \(\overline{{E^{2} }} (L)\) is a decreasing function supports the accepted idea that proteins are frustrated systems.
The scaling of the square interfacial energies E2 between segments of a BLM (GI number: gi116251120) of length L and the rest of the protein as a function of L, normalised by L2 and plotted in logarithmic scale. The dashed line is the mean square energy. The other plots display the scaling coefficient α of the mean square energy, calculated in the linear region, for the proteins of the five families
The shape of \(\overline{{E^{2} }} (L)\) displays a power law \({1/L}^{\alpha }\) followed by a drop. The largest is \(\alpha\), the more frustrated is the system; \(\alpha =0\) for a ferromagnetic-like system. The values of \(\alpha\) for the extant and reconstructed proteins are displayed in the various panels of Fig. 6. In the case of BLM and RNH, the degree of frustration increases with time in a significant way; while for the other three analysed proteins, a specific trend cannot be established (cf. the p values in Table 4).
Another way of quantifying the degree of frustration is counting the number of contacts with energy εij > 0, thus relying on the gauge we chose that sets the zero to the interaction of any residue with gaps. The fraction f of frustrated contacts over the total number of contacts is displayed in Fig. 7. TRD and CYC have a significant monotonic behaviour, decreasing for the former and increasing for the latter; while the other proteins do not show significant monotonicity (cf. the p values in Table 5; the most significant are in bold).
Notice that \(\alpha\) and f do not provide exactly the same information, because the scaling of the interfacial energy depends not only on the number of frustrated contacts but also on their spatial arrangement.
Overall, these data do not support a regular evolutionary trend for the number of frustrated contacts.
We then studied how frustration is localised within proteins. In Fig. 8, it is shown for each family the number fi of frustrated contacts in each site, averaged over all proteins of the same age. All proteins appear to display few sites concentrating most frustration and these sites are highly conserved along evolution. In few cases (three in TRD, three in CYC, one in RNH) sites that were not frustrated in more ancient proteins become frustrated. The opposite is never observed.
Frustrated contacts tend to aggregate into clusters, see the network of frustrated interactions in Fig. 9. As a result, most amino acids are not connected by frustrated contacts, i.e. they are “orphans”. The number of orphans is much larger than that one would observe in a random graph with the same number of nodes and links (p value < 10–6, see also Fig. 9). In fact, random networks display a wide distribution of cluster sizes with a negligible number of orphans (see, e.g. Fig. S11 in the Supp. Mat.), while proteins display few main clusters and a large number of orphans. The amount of orphans either decreases with time (as in CYC and NDK) or fluctuates non-monotonically (as BLM, TRD and RNH, see Fig. 9 and the p values in Table 5).
The size of the largest cluster is comparable to the one of a random network and does not display a regular trend with respect to evolutionary time (cf. Fig. S12 in the Supp. Mat.). On the other hand, its clustering coefficient is significantly larger than that of a random cluster and either decreases significantly with time (for TRD and NDK) or does not display a specific trend (for the other proteins, cf. Fig. S13).
Summing up, frustrated contacts concentrate into few clusters that are not particularly large but are highly connected. There is a signal, although weak, that this tendency increases towards more recent proteins.
A popular tool to study frustration contacts within proteins is the frustratometer developed by Ferreiro and coworkers (Parra et al. 2016). We have performed a rough comparison of the pattern of frustrated contacts of the coevolutionary model with that obtained from the online version of the frustratometer. The results of the comparison are highly family dependent; for example, they are quite good in the case of TRD and much worse for NDK (cf. Fig. S14 in the Supp. Mat.).
Discussion and conclusions
We studied the energetic properties of five protein families and of their reconstructed ancestors; that is in total 890 proteins. The two key tools that permitted this analysis are coevolutionary potentials and the reconstruction algorithms of ancient proteins. Coevolutionary potentials are an efficient and realistic way of describing the interactions that stabilise proteins at the scale of amino acids. They are predictive of many protein features (Halabi et al. 2009; Morcos et al. 2011, 2013; Lui and Tiana 2013; Jana et al. 2014; Tian et al. 2015; Contini and Tiana 2015; dos Santos et al. 2015; Sutto et al. 2015; Granata et al. 2017; Kassem et al. 2018; Baldessari et al. 2020), but need large sequence alignments as input, and, thus, cannot be applied to all protein families. Moreover, they suffer of systematic errors in estimating the interaction energy of residues involved in active sites, that could coevolve for reasons which are not related to the stabilisation of the native state and, thus, cannot be described in terms of the inverse Potts model. In fact, while one would expect that the interactions of the active site can be frustrated, the model predicts erroneously strongly attractive interactions (see Fig. S15 in the Supp. Mat.).
Also, the maximum-likelihood reconstruction of ancient proteins is powerful but not error free. In fact, it was pointed out that when different stabilising mutations accumulate along different lineages, the maximum-likelihood reconstruction could incorrectly incorporate all of the stabilising mutations in the same ancient sequence, resulting in an over-stabilised ancestor (Wheeler et al. 2016). An indication that this is not the case here is that in ancient reconstructed sequences, we do not observe an increase of hydrophobic residues, which are expected to be the most stabilising ones. Anyway, we observe various types of behaviour, with families becoming more stable and families becoming less stable along evolution. In the literature, the large majority of proteins reconstructed by maximum-likelihood algorithms and studied biochemically were shown to become less stable towards recent times (Gaucher et al. 2008; Perez-Jimenez et al. 2011; Carstensen et al. 2012; Risso et al. 2013; Akanuma et al. 2013); this behaviour was explained by an increased environmental temperature in the pre-Cambrian era, which imposed a larger stability to proteins. However, these studies involved few proteins, to be compared with our hundreds. Moreover, we have also shown that the variability in thermodynamic stability is very large even in proteins of similar ages, and consequently drawing general conclusions from a small sampling is quite dangerous.
We then focused the attention to the contacts which mostly stabilise the native state of the proteins. Their number does not seem to vary systematically along evolution. However, they form in each protein a small, highly interconnected cluster. The size of this cluster increases towards recent times, including more and more residues. This is in agreement with the observation that more recent proteins display a more highly connected core (Tiana et al. 2004b). The analysis of amino-acid mutations (Tokuriki et al. 2007) and of protein models (Tiana et al. 2004c) suggests that the stabilisation energy in proteins is not distributed uniformly but is concentrated in a small core. This analysis of reconstructed proteins suggests that the size of the core increases with time.
Also frustrated interactions, that is interactions that evolution could not optimise, are an important feature of proteins. We quantified the degree of frustration of proteins using Phil Anderson’s original definition and showed that it is equivalent to the calculation of the number of interactions with null energy, using gaps in the alignment to gauge the zero of the interactions in the derivation of the energy from coevolution (Lui and Tiana 2013). Also frustrated contacts concentrate into few small, highly connected clusters.
The present data do not support the idea that the total number of frustrated interactions is minimised by evolution, as suggested by the principle of minimal frustration (Bryngelson and Wolynes 1987), but only that they tend to clusterise more. Of course, this does not exclude that minimisation of frustration could take place in the pre-biotic period.
Data availability
Upon request to the authors (large files).
Code availability
References
Abkevich VI, Gutin AM, Shakhnovich EI (1994) Specific nucleus as the transition state for protein folding: evidence from the lattice model. Biochemistry 33:10026–10036
Akanuma S, Nakajima Y, Yokobori S-I et al (2013) Experimental evidence for the thermophilicity of ancestral life. Proc Natl Acad Sci 110:11067–11072. https://doi.org/10.1073/pnas.1308215110
Anderson PW (1978) The concept of frustration in spin glasses. J Less Common Met 62:291–294
Baldessari F, Capelli R, Carloni P, Giorgetti A (2020) Coevolutionary data-based interaction networks approach highlighting key residues across protein families: the case of the G-protein coupled receptors. Comput Struct Biotechnol J. https://doi.org/10.1016/j.csbj.2020.05.003
Bloom JD, Wilke CO, Arnold FH, Adami C (2004) Stability and the evolvability of function in a model protein. Biophys J 86:2758–2764. https://doi.org/10.1016/S0006-3495(04)74329-5
Boussau B, Blanquart S, Necsulea A et al (2008) Parallel adaptations to high temperatures in the Archaean eon. Nature 456:942–945. https://doi.org/10.1038/nature07393
Bryngelson JD, Wolynes PG (1987) Spin glasses and the statistical mechanics of protein folding. Proc Natl Acad Sci USA 84:7524–7528
Bryngelson J, Wolynes P (1989) Intermediates and barrier crossing in a random energy model (with applications to protein folding). J Phys Chem 93:6902–6915. https://doi.org/10.1021/j100356a007
Carstensen L, Sperl JM, Bocola M et al (2012) Conservation of the folding mechanism between designed primordial (βα) 8 -barrel proteins and their modern descendant. J Am Chem Soc 134:12786–12791. https://doi.org/10.1021/ja304951v
Contini A, Tiana G (2015) A many-body term improves the accuracy of effective potentials based on protein coevolutionary data. J Chem Phys 143:25103
Cuturello F, Tiana G, Bussi G (2020) Assessing the accuracy of direct-coupling analysis for RNA contact prediction. RNA. https://doi.org/10.1261/rna.074179.119
de la Paz JA, Nartey CM, Yuvaraj M, Morcos F (2020) Epistatic contributions promote the unification of incompatible models of neutral molecular evolution. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1913071117
dos Santos RN, Morcos F, Jana B et al (2015) Dimeric interactions and complex formation using direct coevolutionary couplings. Sci Rep 5:13652
Ekeberg M, Lövkvist C, Lan Y et al (2013) Improved contact prediction in proteins: using pseudolikelihoods to infer Potts models. Phys Rev E 87:620630
Ferreiro DU, Komives EA, Wolynes PG (2014) Frustration in biomolecules. Q Rev Biophys 47:285–363
Figliuzzi M, Jacquier H, Schug A et al (2015) Coevolutionary landscape inference and the context-dependence of mutations in beta-lactamase TEM-1. Mol Biol Evol 33:268–280
Franco G, Cagiada M, Bussi G, Tiana G (2019) Statistical mechanical properties of sequence space determine the efficiency of the various algorithms to predict interaction energies and native contacts from protein coevolution. Phys Biol 16:046007. https://doi.org/10.1088/1478-3975/ab1c15
Gaucher EA, Govindarajan S, Ganesh OK (2008) Palaeotemperature trend for Precambrian life inferred from resurrected proteins. Nature 451:704–707. https://doi.org/10.1038/nature06510
Granata D, Ponzoni L, Micheletti C, Carnevale V (2017) Patterns of coevolving amino acids unveil structural and dynamical domains. Proc Natl Acad Sci 114:E10612–E10621. https://doi.org/10.1073/pnas.1712021114
Halabi N, Rivoire O, Leibler S, Ranganathan R (2009) Protein sectors: evolutionary units of three-dimensional structure. Cell 138:774–786
Hart KM, Harms MJ, Schmidt BH et al (2014) Thermodynamic system drift in protein evolution. PLoS Biol 12:e1001994. https://doi.org/10.1371/journal.pbio.1001994
Hedges SB, Marin J, Suleski M et al (2015) Tree of life reveals clock-like speciation and diversification. Mol Biol Evol 32:835–845
Jana B, Morcos F, Onuchic JN (2014) From structure to function: the convergence of structure based models and co-evolutionary information. Phys Chem Chem Phys 16:6496–6507
Kassem MM, Christoffersen LB, Cavalli A, Lindorff-Larsen K (2018) Enhancing coevolution-based contact prediction by imposing structural self-consistency of the contacts. Sci Rep 8:11112
Lui S, Tiana G (2013) The network of stabilizing contacts in proteins studied by coevolutionary data. J Chem Phys 139:155103
Mirny LA, Shakhnovich EI (1999) Universally conserved positions in protein folds: reading evolutionary signals about stability, folding kinetics and function. J Mol Biol 291:177–196
Mirny L, Shakhnovich E (2001) Evolutionary conservation of the folding nucleus. J Mol Biol 308:123–129
Morcos F, Pagnani A, Lunt B et al (2011) Direct-coupling analysis of residue coevolution captures native contacts across many protein families. Proc Natl Acad Sci USA 108:E1293–E1301
Morcos F, Jana B, Hwa T, Onuchic JN (2013) Coevolutionary signals across protein lineages help capture multiple protein conformations. Proc Natl Acad Sci USA 110:20533–20538
Morcos F, Schafer NP, Cheng RR et al (2014) Coevolutionary information, protein folding landscapes, and the thermodynamics of natural selection. Proc Natl Acad Sci USA 111:12408–12413
Nguyen HC, Zecchina R, Berg J (2017) Inverse statistical problems: from the inverse Ising problem to data science. Adv Phys 66:197–261
Parra RG, Schafer NP, Radusky LG et al (2016) Protein Frustratometer 2: a tool to localize energetic frustration in protein molecules, now with electrostatics. Nucleic Acids Res 44:W356–W360
Perez-Jimenez R, Inglés-Prieto A, Zhao Z-M et al (2011) Single-molecule paleoenzymology probes the chemistry of resurrected enzymes. Nat Struct Mol Biol 18:592–596. https://doi.org/10.1038/nsmb.2020
Punta M, Coggill PC, Eberhardt RY et al (2012) The Pfam protein families database. Nucleic Acids Res 40:D290-301
Risso VA, Gavira JA, Mejia-Carmona DF et al (2013) Hyperstability and substrate promiscuity in laboratory resurrections of Precambrian β-lactamases. J Am Chem Soc 135:2899–2902
Rodrigues JV, Bershtein S, Li A et al (2016) Biophysical principles predict fitness landscapes of drug resistance. Proc Natl Acad Sci USA 113:E1470–E1478. https://doi.org/10.1073/pnas.1601441113
Russ WP, Figliuzzi M, Stocker C et al (2020) An evolution-based model for designing chorismate mutase enzymes. Science. https://doi.org/10.1126/science.aba3304
Shakhnovich EI, Gutin AM (1993a) Engineering of stable and fast-folding sequences of model proteins. Proc Natl Acad Sci USA 90:7195–7199
Shakhnovich EI, Gutin AM (1993b) A new approach to the design of stable proteins. Protein Eng 6:793–800
Sutto L, Marsili S, Valencia A, Gervasio FL (2015) From residue coevolution to protein conformational ensembles and functional dynamics. Proc Natl Acad Sci USA 112:13567–13572
Taverna DM, Goldstein RA (2002) Why are proteins marginally stable? Proteins Struct Funct Genet. https://doi.org/10.1002/prot.10016
Tian P, Boomsma W, Wang Y et al (2015) Structure of a functional amyloid protein subunit computed using sequence variation. J Am Chem Soc. https://doi.org/10.1021/ja5093634
Tiana G, Broglia RA, Roman HE et al (1998) Folding and misfolding of designed proteinlike chains with mutations. J Chem Phys 108:757–761. https://doi.org/10.1110/ps.03223804
Tiana G, Broglia RA, Shakhnovich EI (2000) Hiking in the energy landscape in sequence space: a bumpy road to good folders. Proteins Struct Funct Genet 39:244–251
Tiana G, Dokholyan NV, Broglia RA, Shakhnovich EI (2004a) The evolution dynamics of model proteins. J Chem Phys 121:2381–2389
Tiana G, Shakhnovich BE, Dokholyan NV, Shakhnovich EI (2004b) Imprint of evolution on protein structures. Proc Natl Acad Sci USA 101:2846–2851
Tiana G, Simona F, De Mori GMS et al (2004c) Understanding the determinants of stability and folding of small globular proteins from their energetics. Protein Sci 13:113–124
Tokuriki N, Stricher F, Schymkowitz J et al (2007) The stability effects of protein mutations appear to be universally distributed. J Mol Biol 369:1318–1332
Tzul FO, Vasilchuk D, Makhatadze GI (2017) Evidence for the principle of minimal frustration in the evolution of protein folding landscapes. Proc Natl Acad Sci USA 114:E1627–E1632
Van Der Spoel D, Lindahl E, Hess B et al (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718
Webb B, Sali A (2016) Comparative protein structure modeling using MODELLER. Curr Protoc Bioinform. https://doi.org/10.1002/cpbi.3
Wetlaufer DB (1973) Nucleation, rapid folding, and globular intrachain regions in proteins. Proc Natl Acad Sci 70:697–701. https://doi.org/10.1073/pnas.70.3.697
Wheeler LC, Lim SA, Marqusee S, Harms MJ (2016) The thermostability and specificity of ancient proteins. Curr Opin Struct Biol 38:37–43
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Zeldovich KB, Chen P, Shakhnovich EI (2007) Protein stability imposes limits on organism complexity and speed of molecular evolution. Proc Natl Acad Sci USA 104:16152–16157
Funding
Open Access funding provided by Università degli Studi di Milano.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Crippa, M., Andreghetti, D., Capelli, R. et al. Evolution of frustrated and stabilising contacts in reconstructed ancient proteins. Eur Biophys J 50, 699–712 (2021). https://doi.org/10.1007/s00249-021-01500-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-021-01500-0