Abstract
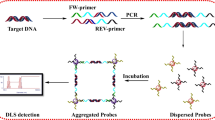
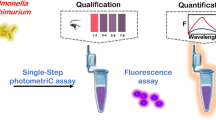
Colorimetry has emerged as a promising option for pathogenic detection. Herein, a novel method based on saltatory rolling circle amplification (SRCA) combined with photosensitization colorimetric assay (SRCA-C) was developed for rapid and sensitive detection of Salmonella in food. The target identification and signal amplification are realized by SRCA to produce a large amount of double-stranded DNA. Subsequently, SYBR Green I as photosensitizer and 3,3',5,5"-tetramethylbenzidine (TMB) as colorimetric oxidase substrate are added to the colorimetric reaction system. The photosensitization reaction occurs under cyan LED irradiation, and TMB is oxidized to convert the double-stranded DNA signal into absorbance signal, thus realizing the detection of Salmonella. This colorimetric method benefits from convenience (label-free and no requirement of costly and complex equipment), short detection time (DNA amplification for 50 min and color development for 15 min) and high sensitivity (detected as low as 13 CFU/mL of Salmonella in pure culture). Five target strains and 10 nontarget strains were used to validate the specificity of the method. The recovery of Salmonella in the spiked milk samples ranged from 98.4% to 118.4%. Compared with the standard culture method, this method was 100% sensitivity, 98.2% specificity, and 98.3% accuracy for the detection of the real samples. Overall, this method highlights the importance for convenient, rapid, and sensitive detection of Salmonella in food.






Similar content being viewed by others
Data availability
All data included in this manuscript are available upon request by contacting with the corresponding author.
References
Veltman B, Harpaz D, Cohen Y, Poverenov E, Eltzov E (2022) Characterization of the selective binding of modified chitosan nanoparticles to Gram-negative bacteria strains. Int J Biol Macromol 194:666–675. https://doi.org/10.1016/j.ijbiomac.2021.11.111
Sun MN, Ma N, Shi HX, Cheong LZ, Yang WG, Qiao ZH (2022) A HCR based multivalent aptamer amplifier for ultrasensitive detection of Salmonella. Sens Actuators B Chem 375:132860. https://doi.org/10.1016/j.snb.2022.132860
Sampson GL, Ruelle SB, Phan L, Hill DW, Hellberg RS (2023) Effectiveness of selected pre-enrichment broths for the detection of Salmonella spp. in meat analogs. Food Control 143:109282. https://doi.org/10.1016/j.foodcont.2022.109282
Bai LL, Wang L, Huang SQ, Bai R, Lv XC, Sun LP, Zhang F, Xu XH (2022) Rapid, visual, and sequence-specific detection of Salmonella in egg liquid with vis-NEAA, a CRISPR/Cas12 empowered new strategy. J Agric Food Chem 70:2401–2409. https://doi.org/10.1021/acs.jafc.1c06715
Yu S, Xu Q, Huang J, Yi B, Aguilar ZP, Xu HY (2021) Rapid and sensitive detection of Salmonella in milk based on hybridization chain reaction and graphene oxide fluorescence platform. J Dairy Sci 104:12295–12302. https://doi.org/10.3168/jds.2021-20713
Cardoso MJ, Nicolau AI, Borda D, Nielsen L, Maia RL, Moretro T, Ferreira V, Knochel S, Langsrud S, Teixeira P (2021) Salmonella in eggs: from shopping to consumption - a review providing an evidence-based analysis of risk factors. Compr Rev Food Sci Food Saf 20:2716–2741. https://doi.org/10.1111/1541-4337.12753
Li ST, He YS, Mann DA, Deng XY (2021) Global spread of Salmonella enteritidis via centralized sourcing and international trade of poultry breeding stocks. Nat Commun 12:5109. https://doi.org/10.1038/s41467-021-25319-7
Jiang R, Lu L, Cao XY, Sun C, Xia JF, Wang ZH (2022) A low-background aptasensor based on enzyme-linked multifunctional carbon nanosheets for the detection of Salmonella. Sens Actuators B Chem 370:132412. https://doi.org/10.1016/j.snb.2022.132412
Wiuff C, Jauho ES, Stryhn H, Andresen LO, Thaulov K, Boas U, Jakobsen MH, Heegaard PMH (2000) Evaluation of a novel enzyme-linked immunosorbent assay for detection of antibodies against Salmonella, employing a stable coating of lipopolysaccharide-derived antigens covalently attached to polystyrene microwells. J Vet Diagn Invest 12:130–135. https://doi.org/10.1177/104063870001200205
Eijkelkamp JM, Aarts HJM, Klerx HJ (2009) Suitability of rapid detection methods for Salmonella in poultry slaughterhouses. Food Anal Methods 2:1–13. https://doi.org/10.1007/s12161-008-9040-5
Brinkman E, Beurden R, Mackintosh R, Beumer R (1995) Evaluation of a new dip-stick test for the rapid detection of Salmonella in food. J Food Prot 58:1023–1027. https://doi.org/10.4315/0362-028X-58.9.1023
Lee KM, Runyon M, Herrman TJ, Phillips R, Hsieh J (2015) Review of Salmonella detection and identification methods: aspects of rapid emergency response and food safety. Food Control 47:264–276. https://doi.org/10.1016/j.foodcont.2014.07.011
Shi XQ, Yu L, Lin C, Li K, Chen JH, Qin H (2021) Biotin exposure–based immunomagnetic separation coupled with sodium dodecyl sulfate, propidium monoazide, and multiplex real-time PCR for rapid detection of viable Salmonella typhimurium, Staphylococcus aureus, and Listeria monocytogenes in milk. J Dairy Sci 104:6588–6597. https://doi.org/10.3168/jds.2020-19887
Fan W, Gao XY, Li HN, Guo WP, Li YY, Wang SW (2022) Rapid and simultaneous detection of Salmonella spp., Escherichia coli O157:H7, and Listeria monocytogenes in meat using multiplex immunomagnetic separation and multiplex real-time PCR. Eur Food Res Technol 248:869–879. https://doi.org/10.1007/s00217-021-03933-5
Jenkins DM, Kubota R, Dong J, Li Y, Higashiguchi D (2011) Handheld device for real-time, quantitative, LAMP-based detection of Salmonella enterica using assimilating probes. Biosens Bioelectron 30:255–260. https://doi.org/10.1016/j.bios.2011.09.020
Zhang HQ, Xu Y, Fohlerova Z, Chang HL, Iliescu C, Neuzil P (2019) LAMP-on-a-chip: revising microfluidic platforms for loop-mediated DNA amplification. TrAC Trends Anal Chem 113:44–53. https://doi.org/10.1016/j.trac.2019.01.015
Xie GY, Zhan ZX, Ye Y, Zhou BQ, Tong P, Aguilar ZP, Xu HY (2022) Hybrid RCA-DLS assay combined with aPCR for sensitive Salmonella enteritidis detection. Anal Biochem 646:114647. https://doi.org/10.1016/j.ab.2022.114647
Hao LL, Gu HJ, Duan N, Wu SJ, Ma XY, Xia Y, Wang HT, Wang ZP (2017) A chemiluminescent aptasensor based on rolling circle amplification and Co2+/N-(aminobuty1)-N-(ethylisolumino1) functional flowerlike gold nanoparticles for Salmonella typhimurium detection. Talanta 164:275–282. https://doi.org/10.1016/j.talanta.2016.11.053
Yang Q, Yang HY, Yuan N, Zuo SN, Zhang YZ, Zhang W (2022) Closed-tube saltatory rolling circle amplification with hydroxynaphthol blue for visual on-site detection of peanut as an allergenic food. Food Chem 393:133408. https://doi.org/10.1016/j.foodchem.2022.133408
Guo W, Yang Q, Zhang YZ, Lu X, Wu CC, Tan JX, Zhang W (2022) Rapid and visual detection of viable Staphylococcus aureus in pork and pork products by PMA and saltatory rolling circle amplification. Eur Food Res Technol 248:1625–1634. https://doi.org/10.1007/s00217-022-03990-4
Zyrina NV, Zheleznaya LA, Dvoretsky EV, Vasiliev VD, Chernov A, Matvienko NI (2007) N.BspD6I DNA nickase strongly stimulates template-independent synthesis of non-palindromic repetitive DNA by Bst DNA polymerase. Biol Chem 388:367–372. https://doi.org/10.1515/BC.2007.043
Fire A, Xu SQ (1995) Rolling replication of short DNA circles. Proc Natl Acad Sci USA 92:4641–4645. https://doi.org/10.1073/pnas.92.10.4641
Liu J, Xie GY, Lv SD, Xiong Q, Xu HY (2023) Recent applications of rolling circle amplification in biosensors and DNA nanotechnology. TrAC Trends Anal Chem 160:116953. https://doi.org/10.1016/j.trac.2023.116953
Zhang YZ, Yang Q, Li C, Yuan YW, Zhang W (2019) Sensitive and visual detection of Cronobacter spp. in powdered infant formula by saltatory rolling circle amplification method. LWT Food Sci Technol 107:41–48. https://doi.org/10.1016/j.lwt.2019.02.050
Lee JE, Mun H, Kim SR, Kim MG, Chang JY, Shim WB (2020) A colorimetric loop-mediated isothermal amplification (LAMP) assay based on HRP-mimicking molecular beacon for the rapid detection of Vibrio parahaemolyticus. Biosens Bioelectron 151:111968. https://doi.org/10.1016/j.bios.2019.111968
Qi YY, Chen YT, He JH, Xiu FR (2020) A colorimetric sensor for DNA detection: combination of synergistic coupling catalysis and significant distinction in the dimensional structure of DNA. Microchem J 159:105546. https://doi.org/10.1016/j.microc.2020.105546
Zhang XF, Huang CP, Xu SX, Chen JB, Zeng Y, Wu P, Hou XD (2015) Photocatalytic oxidation of TMB with the double stranded DNA-SYBR Green I complex for label-free and universal colorimetric bioassay. Chem Commun 51:14465–14468. https://doi.org/10.1039/10.1039/c5cc06105a
Song SX, Wang XY, Xu KE, Xia GM, Yang XB (2019) Visualized detection of Vibrio parahaemolyticus in food samples using dual-functional aptamers and cut-assisted rolling circle amplification. J Agric Food Chem 67:1244–1253. https://doi.org/10.1021/acs.jafc.8b04913
Wang LY, Bai H, Liu XM, Xiao XL, Yu YG, Li XF (2022) Colorimetric sensor based on peroxidase-like activity of chitosan coated on magnetic nanoparticles for rapid detection of the total bacterial count in raw milk. Eur Food Res Technol 248:1321–1333. https://doi.org/10.1007/s00217-022-03970-8
Wang YY, Hu H, Dong TY, Mansour H, Zhang XF, Li F, Wu P (2021) Double-stranded DNA matrix for photosensitization switching. CCS Chem 3:2394–2404. https://doi.org/10.31635/ccschem.020.202000543
Huang LQ, Yuan N, Guo W, Zhang YZ, Zhang W (2023) An electrochemical biosensor for the highly sensitive detection of Staphylococcus aureus based on SRCA-CRISPR/Cas12a. Talanta 252:123821. https://doi.org/10.1016/j.talanta.2022.123821
Zhai SS, Yang Y, Wu YH, Li J, Li YJ, Wu G, Liang JG, Gao HF (2023) A visual CRISPR/dCas9-mediated enzyme-linked immunosorbent assay for nucleic acid detection with single-base specificity. Talanta 257:124318. https://doi.org/10.1016/j.talanta.2023.124318
Jeber JN, Hassan RF, Hammood MK, Al-Jeilawi OHR (2021) Sensitive and simple colorimetric methods for visual detection and quantitative determination of semicarbazide in flour products using colorimetric reagents. Sens Actuators B Chem 341:130009. https://doi.org/10.1016/j.snb.2021.130009
Amin SS, Khoubnasabjafari M, Gharamaleki VJ, Rahimpour E, Jouyban A (2023) Propofol-induced in-situ formation of silver nanoparticles: A sensing colorimetric method. J Pharm Biomed Anal 229:115377. https://doi.org/10.1016/j.jpba.2023.115377
Liu J, Zhan ZX, Liang TB, Xie GY, Aguilar ZP, Xu HY (2020) Dual-signal amplification strategy: universal asymmetric tailing-PCR triggered rolling circle amplification assay for fluorescent detection of Cronobacter spp. in milk. J Dairy Sci 103:3055–3065. https://doi.org/10.3168/jds.2019-17590
Bruno JG (2022) Syringe filter-based DNA aptamer-enzyme-linked colorimetric assay of Salmonella on lettuce. J Microbiol Methods 193:106406. https://doi.org/10.1016/j.mimet.2022.106406
Liu ZK, Yu Y, Fotina T, Petrov R, Klishchova Z, Fotin A, Ma JY (2022) Multiplex PCR assay based on the citE2 gene and intergenic sequence for the rapid detection of Salmonella Pullorum in chickens. Poult Sci 101:101981. https://doi.org/10.1016/j.psj.2022.101981
Nuchchanart W, Pikoolkhao P, Saengthongpinit C (2023) Development of a lateral flow dipstick test for the detection of 4 strains of Salmonella spp. in animal products and animal production environmental samples based on loop-mediated isothermal amplification. Anim Biosci 36:654–670. https://doi.org/10.5713/ab.22.0151
Ge C, Yuan R, Yi L, Yang JL, Zhang HW, Li LX, Nian WQ, Yi G (2018) Target-induced aptamer displacement on gold nanoparticles and rolling circle amplification for ultrasensitive live Salmonella typhimurium electrochemical biosensing. J Electroanal Chem 826:174–180. https://doi.org/10.1016/j.jelechem.2018.07.002
Wang XQ, Luo ZW, Xie QY, Huang ZJ, Wu MF, Duan YX (2020) Toehold-mediated strand displacement reaction formation of three-way junction DNA structure combined with nicking enzyme signal amplification for highly sensitive colorimetric detection of Salmonella typhimurium. Anal Chim Acta 1139:138–145. https://doi.org/10.1016/j.aca.2020.09.023
Wang X, Xu YC, Cheng N, Zhang Q, Yang ZS, Liu BX, Wang XX, Huang KL, Luo YB (2022) Pd@Pt nanoparticles: trienzyme catalytic mechanisms, surface-interface effect with DNA and application in biosensing. Sens Actuators B Chem 364:131907. https://doi.org/10.1016/j.snb.2022.131907
Pal S, Dey S, Batabyal K, Banerjee A, Joardar SN, Samanta I, Isore DP (2017) Characterization of Salmonella gallinarum isolates from backyard poultry by polymerase chain reaction detection of invasion (invA) and Salmonella plasmid virulence (spvC) genes. Vet World 10:814–817. https://doi.org/10.14202/vetworld.2017.814-817
Ye YK, Yan WW, Liu YQ, He SD, Cao XD, Xu X, Zheng HS, Gunasekaran S (2019) Electrochemical detection of Salmonella using an invA genosensor on polypyrrole-reduced graphene oxide modified glassy carbon electrode and AuNPs-horseradish peroxidase-streptavidin as nanotag. Anal Chim Acta 1074:80–88. https://doi.org/10.1016/j.aca.2019.05.012
Acknowledgements
This work was supported by the National Natural Science Foundation of China (32172288 and 31371772), the Natural Science Foundation of Hebei Province (C2019204342), the Scientific Research Program of Hebei Education Department (QN2022073), the Advanced Talents Incubation Program of the Hebei University (521100222024), Medical Science Foundation of Hebei University (2022A06), the Central Government Guided Local Science and Technology Development Fund Projects (Basic Research Projects) (216Z5501G), the Talent Introduction Project of Hebei Province (360-0803-JSN-3YGS), and the Food Processing Discipline Group of Hebei Agricultural University (No. 2021-06).
Author information
Authors and Affiliations
Contributions
HW: Writing—Original Draft, Conceptualization, Methodology, Investigation. QY: Validation, Funding acquisition, Data curation, Writing—Review & Editing. HX: Formal analysis, Software. YZ: Resources, Visualization. XL: Data curation, Software. WZ: Conceptualization, Methodology, Supervision, Funding acquisition, Project administration.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Compliance with Ethics requirements
This article does not contain any studies with human or living animal subjects.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wang, H., Yang, Q., Xu, H. et al. Saltatory rolling circle amplification assay coupled with photosensitization method for rapid and sensitive detection of Salmonella in food. Eur Food Res Technol 249, 2067–2075 (2023). https://doi.org/10.1007/s00217-023-04278-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-023-04278-x