Abstract
Nucleic acid modifications have attracted increasing attention in recent years since they have been found to be related to a number of diseases including cancer. Previous studies have shown that the early development of endometrial cancer (EC) is often accompanied by changes in methylation levels of related genes, and the expression of related proteins that regulate reactive oxygen species (ROS) shows significant differences in EC cells and tissues. However, it has not been reported whether nucleic acid modifications related to methylation or ROS can serve as biomarkers for EC. Accurate quantification of these nucleic acid modifications still has challenges because their amounts in urine are very low and the interferences in urine are complicated. In this study, a novel dispersive solid-phase extraction (DSPE) method based on chitosan-carbon nanotube-Al2O3 (CS-CNT-Al2O3) has been established for the analysis of 5-hydroxymethyluracil (5 mU), 5-methyl-2′-deoxycytidine (5-mdC), 5-hydroxymethyl-2′-deoxycytidine (5-hmdC), 5-formyl-2′-deoxycytidine (5-fdC), and 8-hydroxy-2′-deoxyguanosine (8-OHdG) in EC patient urine samples coupled with UHPLC-QE-Orbitrap-MS/MS and HPLC–UV. Firstly, the synthesis of the CS-CNT-Al2O3 nanocomposite was conducted by a sono-coprecipitation method and was characterized by scanning electron microscope (SEM), energy dispersive spectrometer (EDS), and Fourier transform infrared (FTIR). Under the optimal extraction conditions of DSPE, we successfully quantified 5 mU, 5-mdC, 5-hmdC, 5-fdC, and 8-OHdG in urine samples from 37 EC patients and 39 healthy controls. The results showed that there were significant differences in the levels of 5-mdC, 5-hmdC, 5-fdC, and 8-OHdG in EC patients compared to the healthy control group. The receiver operator characteristic (ROC) curve analysis was carried out to evaluate the potential of 5-mdC, 5-hmdC, 5-fdC, and 8-OHdG to distinguish EC patients from healthy volunteers. The area under the curve (AUC) for 5-mdC, 5-hmdC, 5-fdC, and 8-OHdG was 0.7412, 0.667, 0.8438, and 0.7981, respectively. It indicated that 5-mdC, 5-hmdC, 5-fdC, and 8-OHdG had certain potential in distinguishing between EC patients and healthy volunteers and they could act as potential non-invasive biomarkers for early diagnosis of EC. Moreover, the present study would stimulate investigations of the effects of nucleic acid modifications on the initiation and progression of EC.
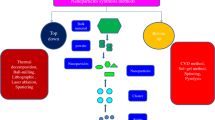
Graphical Abstract






Similar content being viewed by others
Change history
21 March 2024
A Correction to this paper has been published: https://doi.org/10.1007/s00216-024-05247-3
References
Merritt MA, Cramer DW. Molecular pathogenesis of endometrial and ovarian cancer. Cancer Biomark. 2010;9(1–6):287–305.
Mutter GL, Lin MC, Fitzgerald JT, Kum JB, Baak JP, Lees JA, et al. Altered PTEN expression as a diagnostic marker for the earliest endometrial precancers. J Natl Cancer Inst. 2000;92(11):924–30.
Bian J, Sun X, Li B, Ming L. Clinical significance of serum HE4, CA125, CA724, and CA19-9 in patients with endometrial cancer. Technol Cancer Res Treat. 2017;16(4):435–9.
Moore RG, Brown AK, Miller MC, Badgwell D, Lu Z, Allard WJ, et al. Utility of a novel serum tumor biomarker HE4 in patients with endometrioid adenocarcinoma of the uterus. Gynecol Oncol. 2008;110(2):196–201.
Hsieh CH, Changchien CC, Lin H, Huang EY, Huang CC, Lan KC, et al. Can a preoperative CA 125 level be a criterion for full pelvic lymphadenectomy in surgical staging of endometrial cancer? Gynecol Oncol. 2002;86(1):28–33.
Makabe T, Arai E, Hirano T, Ito N, Fukamachi Y, Takahashi Y, et al. Genome-wide DNA methylation profile of early-onset endometrial cancer: its correlation with genetic aberrations and comparison with late-onset endometrial cancer. Carcinogenesis. 2019;40(5):611–23.
Wentzensen N, Bakkum-Gamez JN, Killian JK, Sampson J, Guido R, Glass A, et al. Discovery and validation of methylation markers for endometrial cancer. Int J Cancer. 2014;135(8):1860–8.
Ying J, Xu T, Wang Q, Ye J, Lyu J. Exploration of DNA methylation markers for diagnosis and prognosis of patients with endometrial cancer. Taylor & Francis. 2018;13(5):490–504.
Sharapova T, Talaty N, Buck WR, Fossey S, Liguori MJ, Van Vleet TR. Reduced hepatic global hydroxymethylation in mice treated with non-genotoxic carcinogens is transiently reversible with a methyl supplemented diet. Toxicol Appl Pharmacol. 2021;415:115439.
Sjöström M, Zhao SG, Levy S, Zhang M, Ning Y, Shrestha R, et al. The 5-hydroxymethylcytosine landscape of prostate cancer. Can Res. 2022;82(21):3888–902.
Zhang Z, Jin Y, Zhang W, Chu C, Zhang K, Gao X, et al. Values of 5mC, 5hmC, and TET2 for identifying the presence and progression of breast precancerous lesion. J Clin Lab Anal. 2020;34(5):e23162.
Seo JS, Choi YH, Moon JW, Kim HS, Park SH. Hinokitiol induces DNA demethylation via DNMT1 and UHRF1 inhibition in colon cancer cells. BMC Cell Biol. 2017;18(1):14.
Storebjerg TM, Strand SH, Høyer S, Lynnerup AS, Borre M, Ørntoft TF, et al. Dysregulation and prognostic potential of 5-methylcytosine (5mC), 5-hydroxymethylcytosine (5hmC), 5-formylcytosine (5fC), and 5-carboxylcytosine (5caC) levels in prostate cancer. Clin Epigenetics. 2018;10(1):105.
Boccaletto P, Machnicka MA, Purta E, Piatkowski P, Baginski B, Wirecki TK, et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 2018;46(1):303–7.
Sood AJ, Viner C, Hoffman MM. DNAmod: the DNA modification database. J Chem Inform. 2019;11(1):30.
Sies H, Jones DP. Reactive oxygen species (ROS) as pleiotropic physiological signalling agents. Nat Rev Mol Cell Biol. 2020;21(7):363–83.
Kuran SB, Iplik ES, Cakmakoglu B, Kahraman OT, Iyibozkurt AC, Koc A, et al. Relation of MPO, MnSOD, NQO1 gene variants in endometrial carcinoma in the line of PCR-RFLP methods. Cell Mol Biol (Noisy-le-grand). 2018;64(4):78–82.
Sims N, Rice J, Kasprzyk-Hordern B. An ultra-high-performance liquid chromatography tandem mass spectrometry method for oxidative stress biomarker analysis in wastewater. Anal Bioanal Chem. 2019;411(11):2261–71.
Scholz R, Palatzky P, Matysik FM. Simulation of oxidative stress of guanosine and 8-oxo-7,8-dihydroguanosine by electrochemically assisted injection-capillary electrophoresis-mass spectrometry. Anal Bioanal Chem. 2014;406(3):687–94.
Ataseven D, Öztürk A, Özkaraca M, Joha Z. Anticancer activity of lycopene in HT-29 colon cancer cell line. Med Oncol. 2023;40(5):127.
Saito A, Hamano M, Kataoka H. Simultaneous analysis of multiple urinary biomarkers for the evaluation of oxidative stress by automated online in-tube solid-phase microextraction coupled with negative/positive ion-switching mode liquid chromatography-tandem mass spectrometry. J Sep Sci. 2018;41(13):2743–9.
Nontawong N, Ngaosri P, Chunta S, Jarujamrus P, Nacapricha D, Lieberzeit PA, et al. Smart sensor for assessment of oxidative/nitrative stress biomarkers using a dual-imprinted electrochemical paper-based analytical device. Anal Chim Acta. 2022;1191:339363.
Kumar N, Rosy, Goyal RN. A melamine based molecularly imprinted sensor for the determination of 8-hydroxydeoxyguanosine in human urine. Talanta. 2017;166:215–22.
Chen Q, Hu Y, Fang Z, Ye M, Li J, Zhang S, et al. Elevated levels of oxidative nucleic acid modification markers in urine from gastric cancer patients: quantitative analysis by ultra performance liquid chromatography-tandem Mass Spectrometry. Front Chem. 2020;8:606495.
Guo C, Chen Q, Chen J, Yu J, Hu Y, Zhang S, et al. 8-Hydroxyguanosine as a possible RNA oxidative modification marker in urine from colorectal cancer patients: evaluation by ultra performance liquid chromatography-tandem mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci. 2020;1136:121931.
Ren L, Fang J, Liu G, Zhang J, Zhu Z, Liu H, et al. Simultaneous determination of urinary parabens, bisphenol A, triclosan, and 8-hydroxy-2’-deoxyguanosine by liquid chromatography coupled with electrospray ionization tandem mass spectrometry. Anal Bioanal Chem. 2016;408(10):2621–9.
Varodi C, Pogacean F, Coros M, Rosu MC, Stefan-van Staden RI, Gal E, et al. Detection of 8-hydroxy-2’-deoxyguanosine biomarker with a screen-printed electrode modified with graphene. Sensors. 2019;19(19):4297.
Han P, Hu K, Wang Y, Li L, Wang P, Zhu W, et al. Dispersive solid-phase extraction of non-steroidal anti-inflammatory drugs in water and urine samples using a magnetic ionic liquid hypercrosslinked polymer composite. J Chromatogr A. 2023;1689:463745.
Zong W, Chen J, Han W, Chen J, Wang Y, Wang W, et al. Preparation of chitosan/amino multiwalled carbon nanotubes nanocomposite beads for bilirubin adsorption in hemoperfusion. J Biomed Mater Res B Appl Biomater. 2018;106(1):96–103.
Salvo-Comino C, Rassas I, Minot S, Bessueille F, Arab M, Chevallier V, et al. Voltammetric sensor based on molecularly imprinted chitosan-carbon nanotubes decorated with gold nanoparticles nanocomposite deposited on boron-doped diamond electrodes for catechol detection. Materials. 2020;13(3):688.
Sun Y, Xie L, Feng F, Han Q, Wei L, Tang Z, et al. Simultaneous analysis of two urinary biomarkers of oxidative damage to DNA and RNA based on packed-fiber solid phase extraction coupled with high-performance liquid chromatography. J Chromatogr B Analyt Technol Biomed Life Sci. 2020;1159:122358.
Cervinkova B, Krcmova LK, Sestakova V, Solichova D, Solich P. A fully validated bioanalytical method using an UHPLC–MS/MS system for quantification of DNA and RNA oxidative stress biomarkers. Anal Bioanal Chem. 2017;409(14):3611–21.
Li MJ, Zhang JB, Li WL, Chu QC, Ye JN. Capillary electrophoretic determination of DNA damage markers: content of 8-hydroxy-2′-deoxyguanosine and 8-nitroguanine in urine. J Chromatogr B Analyt Technol Biomed Life Sci. 2011;879(32):3818–22.
Lin HS, Jenner AM, Ong CN, Huang SH, Whiteman M, Halliwell B. A high-throughput and sensitive methodology for the quantification of urinary 8-hydroxy-2’-deoxyguanosine: measurement with gas chromatography-mass spectrometry after single solid-phase extraction. Biochem J. 2004;380(Pt 2):541–8.
Encarnación R, Leticia H, Diego G. Development, validation and application of a fast analytical methodology for the simultaneous determination of DNA- and RNA-derived urinary nucleosides by liquid chromatography coupled to tandem mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci. 2016;1019:132–9.
Hu CW, Liu HH, Li YJ, Chao MR. Direct analysis of 5-methylcytosine and 5-methyl-2′-deoxycytidine in human urine by isotope dilution LC-MS/MS: correlations with N-methylated purines and oxidized DNA lesions. Chem Res Toxicol. 2012;25(2):462–70.
Qian X, Wan Y, Wang A, Xia W, Yang Z, He Z, et al. Urinary metabolites of multiple volatile organic compounds among general, population in Wuhan, central China: inter-day reproducibility, seasonal difference, and their associations with oxidative stress biomarkers. Environ Pollut. 2021;289:117913.
Men C, Chai H, Song X, Li Y, Du H, Ren Q. Identification of DNA methylation associated gene signatures in endometrial cancer via integrated analysis of DNA methylation and gene expression systematically. J Gynecol Oncol. 2017;28(6):e83.
Huo X, Sun H, Cao D, Yang J, Peng P, Yu M, et al. Identification of prognosis markers for endometrial cancer by integrated analysis of DNA methylation and RNA-Seq data. Sci Rep. 2019;9(1):9924.
Li W, Liu M. Distribution of 5-hydroxymethylcytosine in different human tissues. J Nucleic Acids. 2011;2011:870726.
Shen B, Hao J, Lin Y, Li X, Yang X, Huang T, et al. Estrogen-induced extracellular calcium influx promotes endometrial cancer progress by regulating lysosomal activity and mitochondrial ROS. Front Med (Lausanne). 2022;9:835700.
Chung CJ, Lee HL, Chang CH, Chang H, Liu CS, Jung WT, et al. Measurement of urinary arsenic profiles and DNA hypomethylation in a case-control study of urothelial carcinoma. Arch Toxicol. 2019;93(8):2155–64.
Acknowledgements
This work was done in collaboration with Hebei Medical University and the Fourth Hospital of Hebei Medical University.
Funding
The authors received financial support provided by the Natural Science Foundation of Hebei Province (H2022206240), the Medical Science Foundation of Hebei Provine (20220080), and the Research Project of Traditional Chinese Medicine Administration of Hebei Province (2023357).
Author information
Authors and Affiliations
Contributions
Z H, Z X, J Q, L Y: CS-CNT-Al2O3-DSPE conception and design, urine sample collection, data analysis and interpretation, UHPLC-QE-MS/MS and HPLC–UV analysis, manuscript writing, and final approval of the manuscript; Z L, L L, Y H, Z W, W N: urine sample collection, creatinine detection, UHPLC-QE-Orbitrap-MS/MS and HPLC–UV analysis, manuscript writing, and final approval of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
The study protocol was approved by the Ethics Committee of the Fourth Hospital of Hebei Medical University (ethics approval number: 2022KY412). All the urine samples in our present work were obtained from the Fourth Hospital of Hebei Medical University.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: The order of the literature numbers in Table 4, column 8 was wrong, which is different from the description in the paper.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, H., Zhang, X., Zuo, L. et al. A new methodology to reveal potential nucleic acid modifications associated with the risk of endometrial cancer through dispersive solid-phase extraction coupled with UHPLC-QE-Orbitrap-MS/MS and HPLC–UV. Anal Bioanal Chem 416, 2439–2452 (2024). https://doi.org/10.1007/s00216-024-05206-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-024-05206-y