Abstract
Hepatocellular carcinoma (HCC) is one of the most extensive and most deadly cancers in the world. Biomarkers for early diagnosis of HCC are still lacking, and noninvasive and effective biomarkers are urgently needed. Metabolomics is committed to studying the changes of metabolites under stimulation, and provides a new approach for discovery of potential biomarkers. In the current work, 1H nuclear magnetic resonance (NMR) metabolomics approach was utilized to explore the potential biomarkers in HCC progression, and the biomarker panel was evaluated by receiver operating characteristic (ROC) curve analyses. Our results revealed that a biomarker panel consisting of hippurate, creatinine, putrescine, choline, and taurine might be involved in HCC progression. Functional pathway analysis showed that taurine and hypotaurine metabolism is markedly involved in the occurrence and development of HCC. Furthermore, our results indicated that the TPA activity and the level and expression of PKM2 were gradually increased in HCC progression. This research provides a scientific basis for screening potential biomarkers of HCC.
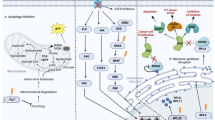
Graphical abstract












Similar content being viewed by others
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- NMR:
-
Nuclear magnetic resonance
- ROC:
-
Receiver operating characteristic
- AFP:
-
Alpha-fetoprotein
- DEN:
-
Diethylnitrosamine
- LC:
-
Liver cirrhosis
- AKP:
-
Alkaline phosphatase
- AST:
-
Aspartate aminotransferase
- ALT:
-
Alanine aminotransferase
- γ-GT:
-
Gamma-glutamyl transpeptidase
- ELISA:
-
Enzyme-linked immunosorbent assay
- TPA:
-
Taurine-pyruvate aminotransferase
- SPF:
-
Specific pathogen-free
- HE:
-
Hematoxylin and eosin
- PCA:
-
Principal component analysis
- PLS-DA:
-
Partial least-squares discriminant analysis
- OPLS-DA:
-
Orthogonal partial least-squares discriminant analysis
- FDR:
-
False discovery rate
- AUC:
-
Area under the ROC curve
References
Njei B, Rotman Y, Ditah I, Lim JK. Emerging trends in hepatocellular carcinoma incidence and mortality. Hepatology. 2015;61:191–9.
Ma C, Han MJ, Heinrich B, Fu Q, Zhang QF, Sandhu M, et al. Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science. 2018;360:eaan5931.
Wang XJ, Zhang AH, Sun H. Power of metabolomics in diagnosis and biomarker discovery of hepatocellular carcinoma. Hepatology. 2013;57:2072–7.
Shao YP, Zhu B, Zheng RY, Zhao XJ, Yin PY, Lu X, et al. Development of urinary pseudotargeted LC-MS-based metabolomics method and its application in hepatocellular carcinoma biomarker discovery. J Proteome Res. 2015;14:906–16.
Marrero JA, Lok AS. Newer markers for hepatocellular carcinoma. Gastroenterology. 2004;127:S113–9.
Chen XL, Zhou L, Yang J, Shen FK, Zhao SP, Wang YL. Hepatocellular carcinoma-associated protein markers investigated by MALDI-TOF MS. Mol Med Rep. 2010;3:589–96.
Luo P, Yin PY, Hua R, Tan YX, Li ZF, Qiu GK, et al. A large-scale, multicenter serum metabolite biomarker identification study for the early detection of hepatocellular carcinoma. Hepatology. 2018;67:662–75.
Caviglia JM, Schwabe RF. Schwabe, mouse models of liver cancer. Methods Mol Biol. 2015;1267:165–83.
Simonetti RG, Cammà C, Fiorello F, Politi F, D'Amico G, Pagliaro L. Hepatocellular carcinoma. A worldwide problem and the major risk factors. Dig Dis Sci. 1991;36:962–72.
Nie CY, Han T, Zhang L, Li Y, Liu H, Xiao SX, et al. Cross-sectional and dynamic change of serum metabolite profiling for hepatitis B-related acute-on-chronic liver failure by UPLC/MS. J Viral Hepat. 2014;21:53–63.
Yang YX, Li CL, Nie X, Feng XS, Chen WX, Yue Y, et al. Metabonomic studies of human hepatocellular carcinoma using high-resolution magic-angle spinning 1H NMR spectroscopy in conjunction with multivariate data analysis. J Proteome Res. 2007;6:2605–14.
Kimhofer T, Fye H, Taylor-Robinson S, Thursz M, Holmes E. Proteomic and metabonomic biomarkers for hepatocellular carcinoma: a comprehensive review. Br J Cancer. 2015;112:1141–56.
Tavallaie R, De Almeida SR, Gooding JJ. Toward biosensors for the detection of circulating microRNA as a cancer biomarker: an overview of the challenges and successes. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2015;7:580–92.
Liang Q, Liu H, Wang C, Li BB. Phenotypic characterization analysis of human Hepatocarcinoma by urine metabolomics approach. Sci Rep. 2016;6:19763.
Shariff MI, Gomaa AI, Cox IJ, Patel M, Williams HR, Crossey MM, et al. Urinary metabolic biomarkers of hepatocellular carcinoma in an Egyptian population: a validation study. J Proteome Res. 2011;10:1828–36.
Chen TL, Xie GX, Wang XY, Fan J, Qiu YP, Zheng XJ, et al. Serum and urine metabolite profiling reveals potential biomarkers of human hepatocellular carcinoma. Mol Cell Proteomics. 2011;10:M110.004945.
Gao HC, Lu Q, Liu X, Cong H, Zhao LC, Wang HM, et al. Application of 1H NMR-based metabonomics in the study of metabolic profiling of human hepatocellular carcinoma and liver cirrhosis. Cancer Sci. 2009;100:782–5.
Liu Y, Hong ZY, Tan GG, Dong X, Yang GJ, Zhao L, et al. NMR and LC/MS-based global metabolomics to identify serum biomarkers differentiating hepatocellular carcinoma from liver cirrhosis. Int J Cancer. 2014;135:658–68.
Casadei-Gardini A, Del Coco L, Marisi G, Conti F, Rovesti G, Ulivi P, et al. (1)H-NMR based serum metabolomics highlights different specific biomarkers between early and advanced hepatocellular carcinoma stages. Cancers (Basel). 2020;12:241.
Carloni V, Lulli M, Madiai S, Mello T, Hall A, Luong TV, et al. CHK2 overexpression and mislocalisation within mitotic structures enhances chromosomal instability and hepatocellular carcinoma progression. Gut. 2018;67:348–61.
Wang KX, Du GH, Qin XM, Gao L. Compound Kushen injection intervenes metabolic reprogramming and epithelial-mesenchymal transition of HCC via regulating β-catenin/c-Myc signaling. Phytomedicine. 2021;153781:93.
Beyoğlu D, Imbeaud S, Maurhofer O, Bioulac-Sage P, Zucman-Rossi J, Dufour JF, et al. Tissue metabolomics of hepatocellular carcinoma: tumor energy metabolism and the role of transcriptomic classification. Hepatology. 2013;58:229–38.
Van der Sluis R, Ungerer V, Nortje C. A, van Dijk A, Erasmus E. New insights into the catalytic mechanism of human glycine N-acyltransferase. J Biochem Mol Toxicol. 2017;31:11.
Coude FX, Coude M, Grimber G, Pelet A, Charpentier C. Potentiation by piridoxilate of the synthesis of hippurate from benzoate in isolated rat hepatocytes. An approach to the determination of new pathways of nitrogen excretion in inborn errors of urea synthesis. Clin Chim Acta. 1984;136:211–7.
Papalazarou V, Zhang T, Paul NR, Juin A, Cantini M, Maddocks ODK, et al. The creatinine-phosphagen system is mechanoresponsive in pancreatic adenocarcinoma and fuels invasion and metastasis. Nat Metab. 2020;2:62–80.
Bardócz S, Grant G, Brown DS, Ralph A, Pusztai A. Polyamines in food-implications for growth and health. J Nutr Biochem. 1993;4:66–71.
Wang J, Zhang S, Li ZF, Yang J, Huang C, Liang RR, et al. (1)H-NMR-based metabolomics of tumor tissue for the metabolic characterization of rat hepatocellular carcinoma formation and metastasis. Tumour Biol. 2011;32:223–31.
Skill NJ, Scott RE, Wu J, Maluccio MA. Hepatocellular carcinoma associated lipid metabolism reprogramming. J Surg Res. 2011;169:51–6.
Moreno A, Rey M, Montane JM, Alonso J, Arús C. 1H NMR spectroscopy of colon tumors and normal mucosal biopsies; elevated taurine levels and reduced polyethyleneglycol absorption in tumors may have diagnostic significance. NMR Biomed. 1993;6:111–8.
Petrick JL, Florio AA, Koshiol J, Pfeiffer RM, Yang B, Yu K, et al. Prediagnostic concentrations of circulating bile acids and hepatocellular carcinoma risk: REVEAL-HBV and HCV studies. Int J Cancer. 2020;147:2743–53.
Alves AP, Mamede AC, Alves MG, Oliveira PF, Rocha SM, Botelho MF, et al. Glycolysis inhibition as a strategy for hepatocellular carcinoma treatment? Curr Cancer Drug Targets. 2019;19:26–40.
Mazurek S. Pyruvate kinase type M2: a key regulator within the tumour metabolome and a tool or metabolic profiling of tumours. Ernst Schering Found Symp Proc. 2007;4:99–124.
Buddington RK, Sangild PT. Companion animals symposium: development of the mammalian gastrointestinal tract, the resident microbiota, and the role of diet in early life. J Anim Sci. 2011;89:1506–19.
Dyar KA, Lutter D, Artati A, Ceglia NJ, Liu Y, Armenta D, et al. Atlas of circadian metabolism reveals system-wide coordination and communication between clocks. Cell. 2018;174:1571–85.
Funding
This project was supported by the Base Program of Joint training graduate student of Shanxi Province (no. 2016JD05), Science and Technology Innovation Team of Shanxi Province (no. 201605D131045–18), and the Key laboratory of Effective Substances Research and Utilization in TCM of Shanxi Province, China (201705D111008–21).
Author information
Authors and Affiliations
Contributions
X.-M. Qin and L. Gao provided the concept and designed the study. K.-X. Wang performed the experiments and drafted the manuscript. K-X. Wang and L. Gao participated in data analysis. X.-M. Qin, G.-H. Du and L. Gao provided oversight. L. Gao contributed to revising and proofreading the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This study was approved in accordance with the ethical standards and approved by SXU Ethics Committee. All the recommendations in the National Institutes of Health Guidelines for Care and Use of Laboratory Animals were strictly followed in this study.
Conflict of interest
The authors have no conflicts of interest, either real or potential, associated with this work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 990 kb)
Rights and permissions
About this article
Cite this article
Wang, Kx., Du, Gh., Qin, Xm. et al. 1H-NMR-based metabolomics reveals the biomarker panel and molecular mechanism of hepatocellular carcinoma progression. Anal Bioanal Chem 414, 1525–1537 (2022). https://doi.org/10.1007/s00216-021-03768-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-021-03768-9