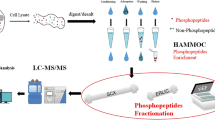
Abstract
Technological advances in liquid chromatography and tandem mass spectrometry (LC-MS/MS) have enabled comprehensive analyses of proteins and their post-translational modifications from cell culture and tissue samples. However, sample complexity necessitates offline prefractionation via a chromatographic method that is orthogonal to online reversed-phase high-performance liquid chromatography (RP-HPLC). This additional fractionation step improves target identification rates by reducing the complexity of the sample as it is introduced to the instrument. A commonly employed offline prefractionation method is high pH reversed-phase (Hi-pH RP) chromatography. Though highly orthogonal to online RP-HPLC, Hi-pH RP relies on buffers that interfere with electrospray ionization. Thus, samples that are prefractionated using Hi-pH RP are typically desalted prior to LC-MS/MS. In the present work, we evaluate an alternative offline prefractionation method, pentafluorophenyl (PFP)-based reversed-phase chromatography. Importantly, PFP prefractionation results in samples that are dried prior to analysis by LC-MS/MS. This reduction in sample handling relative to Hi-pH RP results in time savings and could facilitate higher target identification rates. Here, we have compared the performances of PFP and Hi-pH RP in offline prefractionation of peptides and phosphopeptides that have been isolated from human cervical carcinoma (HeLa) cells. Given the prevalence of isobaric mass tags for peptide quantification, we evaluated PFP chromatography of peptides labeled with tandem mass tags. Our results suggest that PFP is a viable alternative to Hi-pH RP for both peptide and phosphopeptide offline prefractionation.





Similar content being viewed by others
References
Schwanhausser B, Busse D, Li N, Dittmar G, Schuchhardt J, Wolf J, et al. Global quantification of mammalian gene expression control. Nature. 2011;473:337–42.
Yates JR. Mass spectrometry and the age of the proteome. J Mass Spectrom. 1998;33:1–19.
Zhang Y, Fonslow BR, Shan B, Moon-Chang B, Yates JR. Protein analysis by shotgun/bottom-up proteomics. Chem Rev. 2013;113(4):2343–94.
Malmstrom J, Lee H, Aebersold R. Advances in proteomic workflows for systems biology. Curr Opin Biotechnol. 2007;18(4):378–84.
Michalski A, Cox J, Mann M. More than 100,000 detectable peptide species elute in single shotgun proteomics runs but the majority is inaccessible to data-dependent LC-MS/MS. J Proteome Res. 2011;10:1785–93.
McAlister GC, Huttlin EL, Haas W, Ting L, Jedrychowski MP, Rogers JC, et al. Increasing the multiplexing capacity of TMTs using reporter ion isotopologues with isobaric masses. Anal Chem. 2012;84:7469–78.
Anagnostopoulos AK, Stravopodis DJ, Tsangaris GT. Yield of 6,000 proteins by 1D nLC-MS/MS without pre-fractionation. J Chromatogr B. 2016;1–5.
Gilar M, Olivova P, Daly AE, Gebler JC. Orthogonality of separation in two-dimensional liquid chromatography. Anal Chem. 2005;77:6426–34.
Paulo JA, Gaun A, Gygi SP. Global analysis of protein expression and phosphorylation levels in nicotine-treated pancreatic stellate cells. J Proteome Res. 2015;14:4246–56.
Baths TS, Olsen JV. Offline hi pH reversed-phase peptide fractionation for deep phosphoproteome coverage. Methods Mol Biol. 2016;1355:179–92.
Wang Y, Yang F, Gritsenko MA, Wang Y, Clauss T, Liu T, et al. Reversed-phase chromatography with multiple fraction concatenation strategy for proteome profiling of human MCF10A cells. Proteomics. 2011;11:2019–26.
Magdeldin S, Moresco JJ, Yamamoto T, Yates JR. Off-line multidimensional liquid chromatography and auto sampling result in sample loss in LC/LC-MS/MS. J Proteome Res. 2014;13(8):3826–36.
Rauniyar N, Gao B, McClatchy DB, Yates JR. Comparison of protein expression ratios observed by sixplex and duplex TMT labeling method. J Proteome Res. 2013;12:1031–9.
Edwards A, Haas W. Multiplexed quantitative proteomics for high-throughput comprehensive proteome comparisons of human cell lines. Methods Mol Biol. 2016;1394:1–13.
Cao JY, Xu YP, Cai XZ. TMT-based quantitative proteomics analyses reveal novel defense mechanisms of Brassica napus against the devastating necrotrophic pathogen Sclerotinia scleriotorum. J Proteome. 2016;143:265–77.
Kettenbach AN, Gerber SA. Rapid and reproducible single stage phosphopeptide enrichment of complex peptide mixtures: application to general and phosphotyrosine-specific phosphoproteomics experiments. Anal Chem. 2011;83:7635–44.
Paulo JA, O’Connell JD, Everley RA, O’Brien J, Gygi MA, Gygi SP. Quantitative mass spectrometry-based multiplexing compares the abundance of 5000 S. cerevisiae proteins across 10 carbon sources. J Proteome. 2016;148:85–93.
Chick JM, Munger SC, Simecek P, Huttlin EL, Choi K, Gatti DM, et al. Defining the consequences of genetic variation on a proteome-wide scale. Nature. 2016;534:500–5.
Xie R, Oleschuk R. Photoinduced polymerization for entrapping of octadecylsilane microsphere columns for capillary electrochromatography. Anal Chem. 2007;79(4):1529–35.
Eng JK, Jahan TA, Hoopmann MR. Comet: an open-source MS/MS sequence database search tool. Proteomics. 2013;13:22–4.
Arike L, Valgepea K, Peil L, Nahku R, Adamberg K, Vilu R. Comparison and applications of label-free absolute proteome quantification methods on Escherichia coli. J Proteome. 2012;75:5437–48.
Kyte J, Doolittle R. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982;157(1):105–32.
Link AJ, Eng J, Schieltz DM, Carmack E, Mize GJ, Morris DR, et al. Direct analysis of protein complexes using mass spectrometry. Nat Biotechnol. 1999;17(7):676–92.
Acknowledgements
The authors would like to thank Hildreth R. Frost for help with statistical considerations and Richard Avonti, Jason Gilmore, and Sameh Magdeldin for help in generating Venn diagrams. The authors would like to acknowledge funding from the National Institutes of Health (R01-CA155260 and S10-OD016212) to S.A.G.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare no conflicts of interest.
Electronic supplementary material
ESM 1
(PDF 656 kb)
ESM 3
Spreadsheet of Peptides Identified in pH 8 and pH 10 Hi-pH RP Analyses (XLSX 13.1 mb)
ESM 4
Spreadsheet of Proteins Identified in pH 8 and pH 10 Hi-pH RP Analyses (XLSX 1.41 mb)
ESM 6
Spreadsheet of Peptides Identified in PFP RP Analyses (XLSX 13.2 mb)
ESM 7
Spreadsheet of Proteins Identified in PFP RP Analyses (XLSX 1.24 mb)
ESM 8
Spreadsheet of Peptides Identified in Hi-pH RP Analyses (XLSX 9.41 mb)
ESM 9
Spreadsheet of Proteins Identified in Hi-pH RP Analyses (XLSX 1.23 mb)
ESM 10
Spreadsheet of Phosphorylated Peptides Identified in PFP and Hi-pH RP Analyses (XLSX 4.34 mb)
Rights and permissions
About this article
Cite this article
Grassetti, A.V., Hards, R. & Gerber, S.A. Offline pentafluorophenyl (PFP)-RP prefractionation as an alternative to high-pH RP for comprehensive LC-MS/MS proteomics and phosphoproteomics. Anal Bioanal Chem 409, 4615–4625 (2017). https://doi.org/10.1007/s00216-017-0407-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-017-0407-6