Abstract
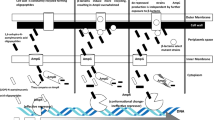
Plasmid-mediated colistin-resistance genes have been reported in human origin clinical samples worldwide which raises its threats to human infections. Notably, mcr-1, mcr-3, mcr-8, and mcr-10 have been reported isolated directly from clinical samples which creates more seriously threaten to human health than other mcr gene types. A multiplex polymerase chain reaction (Multi-PCR) protocol was developed to detect and genotype mobile colistin-resistance genes (mcr-1, mcr-3, mcr-8, mcr-10) in Enterobacteria for clinical laboratory purposes. We first designed four pairs of new primers for the amplification of mcr-1, mcr-3, mcr-8, and mcr-10 gene respectively to achieve stepwise separation of amplicons between 216 and 241 bp, and complete this Multi-PCR system with the assistance of another pair of universal primer. Among which the forward primers for mcr-8 and mcr-10 amplicons were identical. The protocol was validated by testing 11 clinical isolates of Escherichia coli and 3 clinical isolates of Klebsiella from human origin, each well characterized and prospectively validated. The Multi-PCR assay showed full concordance with whole-genome sequence data and displayed higher sensitivity and 100% specificity. The assay could detect all variants of the various mcr alleles described. The Multi-PCR assay successfully genotyped of mcr alleles described in one test.





Similar content being viewed by others
Availability of data and materials
All data and materials are available from first author and corresponding author.
References
AbuOun M, Stubberfield EJ, Duggett NA et al (2017) mcr-1 and mcr-2 variant genes identified in Moraxella species isolated from pigs in Great Britain from 2014 to 2015. J Antimicrob Chemother 72:2745–2749. https://doi.org/10.1093/jac/dkx286
Borowiak M, Fischer J, Hammerl JA, Hendriksen RS, Szabo I, Malorny B (2017) Identification of a novel transposon-associated phosphoethanolamine transferase gene, mcr-5, conferring colistin resistance in d-tartrate fermenting Salmonella enterica subsp. enterica serovar Paratyphi B. J Antimicrob Chemother 72:3317
Budel T, Clement M, Bernasconi OJ, Principe L, Perreten V, Luzzaro F, Endimiani A (2019) Evaluation of EDTA- and DPA-based microdilution phenotypic tests for the detection of MCR-mediated colistin resistance in enterobacteriaceae. Microb Drug Resist 25:494–500. https://doi.org/10.1089/mdr.2018.0275
Carattoli A, Villa L, Feudi C et al (2017) Novel plasmid-mediated colistin resistance mcr-4 gene in Salmonella and Escherichia coli, Italy 2013, Spain and Belgium, 2015 to 2016. Eurosurveillance 22:5
Catry B, Cavaleri M, Baptiste K et al (2015) Use of colistin-containing products within the European Union and European Economic Area (EU/EEA): development of resistance in animals and possible impact on human and animal health. Int J Antimicrob Agents 46:297–306
Daniels JB, Campbell D, Boyd S et al (2019) Development and validation of a clinical laboratory improvement amendments-compliant multiplex real-time PCR assay for detection of mcr genes. Microb Drug Resist 25:6. https://doi.org/10.1089/mdr.2018.0417
Dortet L, Bonnin RA, Pennisi I et al (2018) Rapid detection and discrimination of chromosome-and MCR-plasmid-mediated resistance to polymyxins by MALDI-TOF MS in Escherichia coli: the MALDIxin test. J Antimicrob Chemotherapy 73:3359–3367. https://doi.org/10.1093/jac/dky330
Dos Santos L, Furlan J, Ramos M, Gallo I, de Freitas L, Stehling E (2020) Co-occurrence of mcr-1, mcr-3, mcr-7 and clinically relevant antimicrobial resistance genes in environmental and fecal samples. Arch Microbiol. https://doi.org/10.1007/s00203-020-01890-3
Feng Y (2018) New multiplex PCR primer comprising five primer pairs having specified base pair sequences, useful for detecting polymyxin resistance genes, preferably mobilized colistin resistance genes, and distinguishing corresponding resistance genes. CN108384851-A
Francesca MC, Ludovica C, Andrea L et al (2019) Mobile colistin resistance genes in Escherichia coli from pigs affected by colibacillosis. Int J Antimicrob Agents 52(5):744–746. https://doi.org/10.1016/j.ijantimicag.2018.08.008
Garciagraells C, Scj DK, Vanneste K et al (2017) Detection of plasmid-mediated colistin resistance, mcr-1 and mcr-2 Genes, in Salmonella spp. Isolated from Food at Retail in Belgium from 2012 to 2015. Foodborne Pathog Dis 15:114–117
Germ J, Seme K, Cerar Kišek T et al (2019) Evaluation of a novel epidemiological screening approach for detection of colistin resistant human Enterobacteriaceae isolates using a selective SuperPolymyxin medium. J Microbiol Methods 160:117–123. https://doi.org/10.1016/j.mimet.2019.03.011
He T, Wang R, Liu D et al (2019) Emergence of plasmid-mediated high-level tigecycline resistance genes in animals and humans. Nature Microbiol 4:1450–1456. https://doi.org/10.1038/s41564-019-0445-2
Hu S, Yu Y, Wu X, Xia X, Hui W (2017) Simultaneous detection and identification of pathogenic Cronobacter species by high-resolution melting analysis in powdered infant formulas. Int J Dairy Technol. https://doi.org/10.1111/1471-0307.12410
Imirzalioglu C, Falgenhauer L, Schmiedel J et al (2017) Evaluation of a Loop-Mediated Isothermal Amplification-Based Assay for the Rapid Detection of Plasmid-Encoded Colistin Resistance Gene mcr-1 in Enterobacteriaceae Isolates. Antimicrob Agents Chemother 61(4):AAC.02326–02316. https://doi.org/10.1128/AAC.02326-16
Jousset AB, Bernabeu S, Bonnin RA et al (2019) Development and validation of a multiplex polymerase chain reaction assay for detection of the five families of plasmid-encoded colistin resistance. Int J Antimicrob Agents 53:302–309. https://doi.org/10.1016/j.ijantimicag.2018.10.022
Kahlmeter G, Brown D, Goldstein FW, Macgowan AP, Mouton JW, Odenholt I et al (2010) European committee on antimicrobial susceptibility testing (eucast) technical notes on antimicrobial susceptibility testing. Clin Microbiol Infect 12(6):501–503. https://doi.org/10.1111/j.1469-0691.2006.01454.x
Kieffer N, Royer G, Decousser J et al (2019) mcr-9, an Inducible Gene Encoding an Acquired Phosphoethanolamine Transferase in Escherichia coli, and Its Origin. Antimicrob Agents Chemother 63:e00965-00919. https://doi.org/10.1128/aac.00965-19
Lescat M, Poirel L, Nordmann P (2018) Rapid multiplex polymerase chain reaction for detection of mcr-1 to mcr-5 genes. Diagn Microbiol Infect Dis 92:267–269. https://doi.org/10.1016/j.diagmicrobio.2018.04.010
Li J, Cao J, Zhu YG et al (2018) Global survey of antibiotic resistance genes in air. Environm Sci Technol 52(19):10975–10984. https://doi.org/10.1021/acs.est.8b02204
Lin Z, Zhao X, Huang J et al (2019) Rapid screening of colistin-resistant Escherichia coli, Acinetobacter baumannii and Pseudomonas aeruginosa by the use of Raman spectroscopy and hierarchical cluster analysis. Analyst 144:2803–2810. https://doi.org/10.1039/C8AN02220H
Liu YY, Wang Y, Walsh TR et al (2016) Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: a microbiological and molecular biological study. Lancet Infect Dis 16:161–168
Macesic N, Kahn S, Giddins MJ et al (2019) Escherichia coli harboring mcr-1 in a cluster of liver transplant recipients: detection through active surveillance and whole genome sequencing. Antimicrob Agents Chemother 63(6):e02680–02618. https://doi.org/10.1128/aac.02680-18
Mcphee JLS, Rew H (2010) Cationic antimicrobial peptides activate a two-component regulatory system, PmrA-PmrB, that regulates resistance to polymyxin B and cationic antimicrobial peptides in Pseudomonas aeruginosa. Mol Microbiol 50:205–217
Mlynarcik P, Kolar M (2019) Molecular mechanisms of polymyxin resistance and detection of mcr genes. Biomed Pap-Olomouc 163:28–38. https://doi.org/10.5507/bp.2018.070
Monte D, Nelson V, Cerdeira L et al (2019) Corrigendum: Multidrug- and colistin-resistant Salmonella enterica: Multidrug- and colistin-resistant 4,[5], 12:i:- sequence type 34 carrying the gene on the IncHI2 plasmid recovered from a human. J Med Microbiol 68:1694–1694. https://doi.org/10.1099/jmm.0.001057
NIH (2021) https://www.ncbi.nlm.nih.gov/pathogens/isolates#/refgene/gene_family:(mcr-1)
Poirel L, Jayol A, Nordmann P (2017) Polymyxins: antibacterial activity, susceptibility testing, and resistance mechanisms encoded by plasmids or chromosomes. Clin Microbiol Rev 30:557–596
Rebelo AR, Bortolaia V, Kjeldgaard JS et al (2018) Multiplex PCR for detection of plasmid-mediated colistin resistance determinants, mcr-1, mcr-2, mcr-3, mcr-4 and mcr-5 for surveillance purposes. Euro Surveillance 23:17–00672. https://doi.org/10.2807/1560-7917.es.2018.23.6.17-00672
Rhouma M, Beaudry F, Thériault W, Letellier A (2016) Colistin in pig production: chemistry, mechanism of antibacterial action, microbial resistance emergence, and one health perspectives. Front Microbiol 7:1789
Roberts KD, Azad MAK, Wang J et al (2015) Antimicrobial activity and toxicity of the major lipopeptide components of polymyxin B and colistin: last-line antibiotics against multidrug-resistant gram-negative bacteria. ACS Infect Dis 1:568–575. https://doi.org/10.1021/acsinfecdis.5b00085
Salloum T, Panossian B, Bitar I, Hrabak J, Araj G, Tokajian S (2020) First report of plasmid-mediated colistin resistance mcr-8.1 gene from a clinical Klebsiella pneumoniae isolate from Lebanon. Antimicrob Resist Inf Control 9:94. https://doi.org/10.1186/s13756-020-00759-w
Shabnam H, Mohammadreza M, Nariman S, Azizollah E (2018) Application of high-resolution melting-curve analysis on pvp A gene for detection and classification of Mycoplasma gallisepticum strains. Microb Pathogen 124:365–371. https://doi.org/10.1016/j.micpath.2018.06.03232
Shanmugakani RK, Akeda Y, Sugawara Y et al (2019) PCR-dipstick-oriented surveillance and characterization of mcr-1- and carbapenemase-carrying enterobacteriaceae in a thai hospital. Front Microbiol 10:11. https://doi.org/10.3389/fmicb.2019.00149
Shen Y, Zhou H, Xu J et al (2018) Anthropogenic and environmental factors associated with high incidence of mcr-1 carriage in humans across China. Nature Microbiol 3:1054–1062. https://doi.org/10.1038/s41564-018-0205-8
Sun J, Zhang H, Liu YH, Feng Y (2018a) Towards understanding MCR-like colistin resistance. Trends Microbiol 26:794–807
Sun QL, Hu YY, Zhou HW, Shu LB, Wang HY, Huang ZX, Zhang R (2018) Alkaline peptone water-based enrichment method for mcr-3 from acute diarrheic outpatient gut samples. Front Med 5:6. https://doi.org/10.3389/fmed.2018.00099
Sun R, Ke B, Fang L et al (2020) Global clonal spread of mcr-3-carrying MDR ST34 Salmonella enterica serotype Typhimurium and monophasic 1,4, [5], 12:i:-variants from clinical isolates. J Antimicrob Chemother 75:1756–1765. https://doi.org/10.1093/jac/dkaa115
Thiry D, Berrah A, Evrard J, Duprez J-N, Mainil JG, Saulmont M (2019) Assessment of two selective agar media to isolate colistin-resistant bovine Escherichia coli: Correlation with minimal inhibitory concentration and presence of mcr genes. J Microbiol Methods 159:174–178. https://doi.org/10.1016/j.mimet.2019.03.004
Tolosi R, Apostolakos I, Laconi A, Carraro L, Grilli G, Cagnardi P, Piccirillo A (2020) Rapid detection and quantification of plasmid-mediated colistin resistance genes (mcr-1 to mcr-5) by real-time PCR in bacterial and environmental samples. J Appl Microbiol. https://doi.org/10.1111/jam.14738
Wang Y, Tian GB, Zhang R et al (2017) Prevalence, risk factors, outcomes, and molecular epidemiology of mcr-1-positive Enterobacteriaceae in patients and healthy adults from China: an epidemiological and clinical study. Lancet Infect Dis 17:390–399. https://doi.org/10.1016/s1473-3099(16)30527-8
Wang X, Wang Y, Zhou Y et al (2018) Emergence of a novel mobile colistin resistance gene, mcr-8, in NDM-producing Klebsiella pneumoniae. Emerg Microb Infect 7(1):122–130. https://doi.org/10.1038/s41426-018-0124-z
Wang HH, Chen Y, Strich JR et al (2019) Rapid detection of colistin resistance protein MCR-1 by LC-MS/MS. Clin Proteom 16:10. https://doi.org/10.1186/s12014-019-9228-2
Wang Y, Liu F, Hu Y, Zhang G, Zhu B, Gao G (2019) Detection of mobile colistin resistance gene mcr-9 in carbapenem-resistant Klebsiella pneumoniae strains of human origin in Europe. J Infect 80:3. https://doi.org/10.1016/j.jinf.2019.12.016
Wang Z, Fu Y, Schwarz S et al (2019) Genetic environment of colistin resistance genes mcr-1 and mcr-3 in Escherichia coli from one pig farm in China. Veterin Microbiol 230:56–61. https://doi.org/10.1016/j.vetmic.2019.01.011
Wang C, Feng Y, Liu L, Wei L, Kang M, Zong Z (2020) mcr-10Identification of novel mobile colistin resistance gene. Emerg Microb Infect 9:508–516. https://doi.org/10.1080/22221751.2020.1732231
Wang Y, Xu C, Zhang R et al (2020) Changes in colistin resistance and mcr-1 abundance in Escherichia coli of animal and human origins following the ban of colistin-positive additives in China: an epidemiological comparative study. Lancet Infect Dis 20:1161–1171. https://doi.org/10.1016/S1473-3099(20)30149-3
Xavier BB, Lammens C, Ruhal R, Kumar-Singh S, Butaye P, Goossens H, Malhotra-Kumar S (2016) Identification of a novel plasmid-mediated colistin-resistance gene, mcr-2, in Escherichia coli, Belgium, June 2016. Eurosurveillance 21:07
Xian-Quan C, Hai-Qiong Y, Zhou-Xi R et al (2013) Rapid detection and simultaneous genotyping of Cronobacter spp. (formerly Enterobacter sakazakii) in powdered infant formula using real-time PCR and high resolution melting (HRM) analysis. Plos One 8:67082
Yang YQ, Li YX, Lei CW, Zhang AY, Wang HN (2018) Novel plasmid-mediated colistin resistance gene mcr-7.1 in Klebsiella pneumoniae. J Antimicrob Chemother 73:1791–1795
Yin W, Li H, Shen Y et al (2017) Novel Plasmid-Mediated Colistin Resistance Gene mcr-3 in Escherichia coli. mBio 8:e00543–00517. https://doi.org/10.1128/mBio.00543-17
Yu Y, Andrey D, Yang R-S et al (2020) A Klebsiella pneumoniae strain co-harbouring mcr-1 and mcr-3 from a human in Thailand. J Antimicrob Chemother 75:1. https://doi.org/10.1093/jac/dkaa133
Yu Y, Andrey D, Yang R-S et al (2020) A Klebsiella pneumoniae strain co-harbouring mcr-1 and mcr-3 from a human in Thailand. J Antimicrob Chemother 75:3. https://doi.org/10.1093/jac/dkaa133
Yu Y, Andrey D, Yang R et al (2020) A Klebsiella pneumoniae strain co-harbouring mcr-1 and mcr-3 from a human in Thailand. J Antimicrob Chemother 75:3. https://doi.org/10.1093/jac/dkaa133
Zhang J, Chen L, Wang J et al (2018) Molecular detection of colistin resistance genes (mcr-1, mcr-2 and mcr-3) in nasal/oropharyngeal and anal/cloacal swabs from pigs and poultry. Sci Rep 8:3705
Zheng BW, Feng CY, Xu H, Yu X, Guo LH, Jiang XW, Song XH (2019) Detection and characterization of ESBL-producing Escherichia coli expressing mcr-1 from dairy cows in China. J Antimicrob Chemother 74:321–325. https://doi.org/10.1093/jac/dky446
Zhong LL, Zhou Q, Tan CY, Roberts AP, El-Sayed Ahmed MAEG, Chen G, Dai M, Yang F, Xia Y, Liao K, Liang Y, Yang Y, Feng S, Zheng X, Tian GB (2019) Multiplex loop-mediated isothermal amplification (multi-LAMP) assay for rapid detection of mcr-1 to mcr-5 in colistin-resistant bacteria. Infect Drug Resist 12:1877–1887. https://doi.org/10.2147/IDR.S210226
Acknowledgements
We appreciated Miss Changfeng Peng for her assistance improving the language of manuscript.
Funding
This work was supported by funding from the National Key Research and Development Program of China (No. 2018YFD0500303 and 2019YFC1605104), the China Postdoctoral Science Foundation (No. 2018M643202), the Sanming Project of Medicine in Shenzhen (SZSM201611068), and the Shenzhen Science and Technology Planning Project (JCYJ20170413101841798 and JCYJ20190807103003724).
Author information
Authors and Affiliations
Contributions
HSF performed microbiological and molecular experiments, analyzed the data, and wrote the manuscript. HSF, LZQ and KYB acquired laboratory data and performed bioinformatics analysis. HSF, WY, and SJZ designed the study, wrote, reviewed, and edited the manuscript.
Corresponding authors
Ethics declarations
Conflicts of interest
The authors declares that they have no conflict of interests.
Ethical approval
Ethical approval was given by Medical Ethic Committee of Shenzhen Center for Disease Control and Prevention. All procedures performed in this study involving human participants were in accordance with the ethical standards of the Medical Ethic Committee of Shenzhen Center for Disease Control and Prevention.
Consent to participate
Individual consent forms were translated into Mandarin and consent was obtained for all volunteers face to face. Informed consent from all participants were obtained in this study.
Consent for publication
Written informed consent for publication was obtained from all participants.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hu, S., Lv, Z., Wang, Y. et al. Rapid detection of human origin colistin-resistance genes mcr-1, mcr-3, mcr-8, mcr-10 in clinical fecal samples. Arch Microbiol 203, 4405–4417 (2021). https://doi.org/10.1007/s00203-021-02407-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02407-2