Abstract
Previously we established a family of macrocyclic peptide triazoles (cPTs) that inactivate the Env protein complex of HIV-1, and identified the pharmacophore that engages Env’s receptor binding pocket. Here, we examined the hypothesis that the side chains of both components of the triazole Pro - Trp segment of cPT pharmacophore work in tandem to make intimate contacts with two proximal subsites of the overall CD4 binding site of gp120 to stabilize binding and function. Variations of the triazole Pro R group, which previously had been significantly optimized, led to identification of a derivative, MG-II-20, containing a pyrazole substitution. MG-II-20 has improved functional properties over previously examined cPTs, with KD for gp120 in the nM range. In contrast, new modifications of the Trp indole side chain, with either methyl- or bromo- components appended, had disruptive effects on gp120 binding, reflecting the sensitivity of function to changes in this component of the encounter complex. Plausible in silico models of cPT:gp120 complex structures were obtained that are consistent with the overall hypothesis of occupancy by the triazole Pro and Trp side chains, respectively, into the β20/21 and Phe43 sub-cavities. The overall results strengthen the definition of the cPT-Env inactivator binding site and provide a new lead composition (MG-II-20) as well as structure-function findings to guide future HIV-1 Env inactivator design.
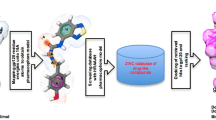
Graphical Abstract






Similar content being viewed by others
References
Margolis DM, Archin NM, Cohen MS, Eron JJ, Ferrari G, Garcia JV, et al. Curing HIV: seeking to target and clear persistent infection. Cell. 2020;181:189–206. https://doi.org/10.1016/j.cell.2020.03.005.
Thomas J, Ruggiero A, Paxton WA, Pollakis G. Measuring the success of HIV-1 cure strategies. Front Cell Infect Microbiol. 2020;10:134. https://doi.org/10.3389/fcimb.2020.00134.
Kim J, Vasan S, Kim JH, Ake JA. Current approaches to HIV vaccine development: a narrative review. J Int AIDS Soc. 2021;24:e25793. https://doi.org/10.1002/jia2.25793.
Wang Q, Finzi A, Sodroski J. The conformational states of the HIV-1 envelope glycoproteins. Trends Microbiol. 2020;28:655–67. https://doi.org/10.1016/j.tim.2020.03.007.
Vézina D, Gong SY, Tolbert WD, Ding S, Nguyen D, Richard J, et al. Stabilizing the HIV-1 envelope glycoprotein State 2A conformation. J Virol. 2020;95:e01620–20. https://doi.org/10.1128/JVI.01620-20.
Zou S, Zhang S, Gaffney A, Ding H, Lu M, Grover JR, et al. Long-acting BMS-378806 analogues stabilize the state-1 conformation of the human immunodeficiency virus type 1 envelope glycoproteins. J Virol. 2020;94:e00148–20. https://doi.org/10.1128/JVI.00148-20.
Gopi H, Cocklin S, Pirrone V, McFadden K, Tuzer F, Zentner I, et al. Introducing metallocene into a triazole peptide conjugate reduces its off-rate and enhances its affinity and antiviral potency for HIV-1 gp120. J Mol Recognit. 2009;22:169–74.
Umashankara M, McFadden K, Zentner I, Schon A, Rajagopal S, Tuzer F, et al. The active core in a triazole peptide dual-site antagonist of HIV-1 gp120. ChemMedChem. 2010;5:1871–9. https://doi.org/10.1002/cmdc.201000222.
Kamanna K, Aneja R, Duffy C, Kubinski P, Rodrigo Moreira D, Bailey LD, et al. Non-natural peptide triazole antagonists of HIV-1 envelope gp120. ChemMedChem. 2013;8:322–8. https://doi.org/10.1002/cmdc.201200422.
Emileh A, Duffy C, Holmes AP, Rosemary Bastian A, Aneja R, Tuzer F, et al. Covalent conjugation of a peptide triazole to HIV-1 gp120 enables intramolecular binding site occupancy. Biochemistry. 2014;53:3403–14. https://doi.org/10.1021/bi500136f.
Carter EP, Ang CG, Chaiken IM. Peptide triazole inhibitors of HIV-1: hijackers of env metastability. Current protein & peptide science. 2022. https://doi.org/10.2174/1389203723666220610120927.
Rashad AA, Kalyana Sundaram RV, Aneja R, Duffy C, Chaiken I. Macrocyclic envelope glycoprotein antagonists that irreversibly inactivate HIV-1 before host cell encounter. J Med Chem. 2015;58:7603–8. https://doi.org/10.1021/acs.jmedchem.5b00935.
Rashad AA, Acharya K, Haftl A, Aneja R, Dick A, Holmes AP, et al. Chemical optimization of macrocyclic HIV-1 inactivators for improving potency and increasing the structural diversity at the triazole ring. Org Biomol Chem. 2017;15:7770–82. https://doi.org/10.1039/C7OB01448A.
Bastian AR, Kantharaju, McFadden K, Duffy C, Rajagopal S, Contarino MR, et al. Cell-free HIV-1 virucidal action by modified peptide triazole inhibitors of Env gp120. ChemMedChem. 2011;6:1335–9. https://doi.org/10.1002/cmdc.201100177.
Bastian AR, Contarino M, Bailey LD, Aneja R, Moreira DR, Freedman K, et al. Interactions of peptide triazole thiols with Env gp120 induce irreversible breakdown and inactivation of HIV-1 virions. Retrovirology. 2013;10:153. https://doi.org/10.1186/1742-4690-10-153.
Bailey LD, Kalyana Sundaram RV, Li H, Duffy C, Aneja R, Rosemary Bastian A, et al. Disulfide sensitivity in the env protein underlies lytic inactivation of HIV-1 by peptide triazole thiols. ACS Chem Biol. 2015;10:2861–73. https://doi.org/10.1021/acschembio.5b00381.
Ang CG, Carter E, Haftl A, Zhang S, Rashad AA, Kutzler M, et al. Peptide triazole thiol irreversibly inactivates metastable HIV-1 env by accessing conformational triggers intrinsic to virus-cell entry. Microorganisms. 2021;9:1286. https://doi.org/10.3390/microorganisms9061286.
Tuzer F, Madani N, Kamanna K, Zentner I, LaLonde J, Holmes A, et al. HIV-1 Env gp120 structural determinants for peptide triazole dual receptor site antagonism. Proteins. 2013;81:271–90. https://doi.org/10.1002/prot.24184.
Emileh A, Tuzer F, Yeh H, Umashankara M, Moreira DR, Lalonde JM, et al. A model of peptide triazole entry inhibitor binding to HIV-1 gp120 and the mechanism of bridging sheet disruption. Biochemistry. 2013;52:2245–61. https://doi.org/10.1021/bi400166b.
Aneja R, Rashad AA, Li H, Kalyana Sundaram RV, Duffy C, Bailey LD, et al. Peptide triazole inactivators of HIV-1 utilize a conserved two-cavity binding site at the junction of the inner and outer domains of env gp120. J Med Chem. 2015;58:3843–58. https://doi.org/10.1021/acs.jmedchem.5b00073.
Acharya K, Rashad AA, Moraca F, Klasse PJ, Moore JP, Abrams C, et al. Recognition of HIV-inactivating peptide triazoles by the recombinant soluble Env trimer, BG505 SOSIP.664. Proteins. 2017;85:843–51. https://doi.org/10.1002/prot.25238.
Pancera M, Druz A, Zhou T, O’Dell S, Louder M, Madani N, et al. Structure of BMS-806, a Small-molecule HIV-1 Entry Inhibitor, Bound to BG505 SOSIP.664 HIV-1 Env Trimer. AIDS research and human retroviruses. 2014;30:A151-A. https://doi.org/10.1089/aid.2014.5307c.abstract.
Jacobson MP, Friesner RA, Xiang Z, Honig B. On the role of the crystal environment in determining protein side-chain conformations. J Mol Biol. 2002;320:597–608. https://doi.org/10.1016/S0022-2836(02)00470-9.
Jacobson MP, Pincus DL, Rapp CS, Day TJF, Honig B, Shaw DE, et al. A hierarchical approach to all-atom protein loop prediction. Proteins: Struct Funct Bioinforma. 2004;55:351–67. https://doi.org/10.1002/prot.10613.
Prime. Schrödinger. Release 2021-3 ed. New York, NY: Schrödinger, LLC; 2021.
Zhang S, Wang K, Wang WL, Nguyen HT, Chen S, Lu M, et al. Asymmetric structures and conformational plasticity of the uncleaved full-length human immunodeficiency virus envelope glycoprotein trimer. J Virol. 2021;95:e0052921. https://doi.org/10.1128/jvi.00529-21.
Gopi H, Umashankara M, Pirrone V, LaLonde J, Madani N, Tuzer F, et al. Structural determinants for affinity enhancement of a dual antagonist peptide entry inhibitor of human immunodeficiency virus type-1. J Med Chem. 2008;51:2638–47. https://doi.org/10.1021/jm070814r.
Pan D, Wang W, Liu W, Yang L, Huang HW. Chain packing in the inverted hexagonal phase of phospholipids: a study by X-ray anomalous diffraction on bromine-labeled chains. J Am Chem Soc. 2006;128:3800–7. https://doi.org/10.1021/ja058045t.
Montefiori DC. Evaluating neutralizing antibodies against HIV, SIV, and SHIV in luciferase reporter gene assays. Current protocols in immunology/edited by John E Coligan [et al]. 2005;Chapter 12:Unit 12 1. https://doi.org/10.1002/0471142735.im1211s64.
Madhavi Sastry G, Adzhigirey M, Day T, Annabhimoju R, Sherman W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J Computer-Aided Mol Des. 2013;27:221–34. https://doi.org/10.1007/s10822-013-9644-8.
Protein Preparation Wizard. Schrödinger. Release 2021-3 ed. New York, NY: Schrödinger, LLC; 2021.
Epik. Schrödinger. Release 2021-3 ed. New York, NY: Schrödinger, LLC; 2021.
Impact. Schrödinger. Release 2021-3 ed. New York, NY: Schrödinger, LLC; 2021.
Proceedings of the 2006 ACM/IEEE conference on Supercomputing2006; Tampa, Florida: Association for Computing Machinery.
Desmond Molecular Dynamics System. Schrödinger. Release 2021-3 ed. New York, NY: D.E. Shaw Research; 2021.
Maestro-Desmond Interoperability Tools. Schrödinger. Release 2021-3 ed. New York, NY: Schrödinger, LLC; 2021.
Halgren TA. Identifying and characterizing binding sites and assessing druggability. J Chem Inf Model. 2009;49:377–89. https://doi.org/10.1021/ci800324m.
Halgren T. New method for fast and accurate binding-site identification and analysis. Chem Biol Drug Des. 2007;69:146–8. https://doi.org/10.1111/j.1747-0285.2007.00483.x.
SiteMap. Schrödinger. Release 2021-3. New York, NY: Schrödinger, LLC; 2021.
LigPrep. Schrödinger. Release 2021-3. New York, NY: Schrödinger, LLC; 2021.
Farid R, Day T, Friesner RA, Pearlstein RA. New insights about HERG blockade obtained from protein modeling, potential energy mapping, and docking studies. Bioorg Med Chem. 2006;14:3160–73. https://doi.org/10.1016/j.bmc.2005.12.032.
Sherman W, Day T, Jacobson MP, Friesner RA, Farid R. Novel procedure for modeling ligand/receptor induced fit effects. J Med Chem. 2006;49:534–53. https://doi.org/10.1021/jm050540c.
Sherman W, Beard HS, Farid R. Use of an induced fit receptor structure in virtual screening. Chem Biol drug Des. 2006;67:83–4. https://doi.org/10.1111/j.1747-0285.2005.00327.x.
Induced Fit Docking protocol. Schrödinger. Release 2021-3 ed. New York, NY: Schrödinger, LLC; 2021.
Glide. Schrödinger. Release 2021-3 ed. New York, NY: Schrödinger, LLC; 2021.
Friesner RA, Murphy RB, Repasky MP, Frye LL, Greenwood JR, Halgren TA, et al. Extra precision glide: docking and scoring incorporating a model of hydrophobic enclosure for protein−ligand complexes. J Med Chem. 2006;49:6177–96. https://doi.org/10.1021/jm051256o.
Halgren TA, Murphy RB, Friesner RA, Beard HS, Frye LL, Pollard WT, et al. Glide: a new approach for rapid, accurate docking and scoring. 2. Enrichment factors in database screening. J Med Chem. 2004;47:1750–9. https://doi.org/10.1021/jm030644s.
Friesner RA, Banks JL, Murphy RB, Halgren TA, Klicic JJ, Mainz DT, et al. Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem. 2004;47:1739–49. https://doi.org/10.1021/jm0306430.
Meanwell NA, Krystal MR, Nowicka-Sans B, Langley DR, Conlon DA, Eastgate MD, et al. Inhibitors of HIV-1 attachment: the discovery and development of temsavir and its prodrug fostemsavir. J Med Chem. 2018;61:62–80. https://doi.org/10.1021/acs.jmedchem.7b01337.
Fritschi CJ, Anang S, Gong Z, Mohammadi M, Richard J, Bourassa C, et al. Indoline CD4-mimetic compounds mediate potent and broad HIV-1 inhibition and sensitization to antibody-dependent cellular cytotoxicity. Proc Natl Acad Sci. 2023;120:e2222073120. https://doi.org/10.1073/pnas.2222073120.
Acknowledgements
Research reported in this publication was supported by NIH GM56550/AI150471, PA Department of Health CURE grant SAP#4100083087 (2019 Formula Grant Program: Characterization of the Conformational Change Properties of the HIV-1 Envelope Protein), and an Undergraduate Research and Enrichment Program Minigrant to Monisha Gupta. S200 biosensor used in this work was supported by the Office of the Director, National Institutes of Health through NIH Award S10OD027009. We also acknowledge with gratitude the assistance of Dr. Chris Fritschi for discussions on synthetic strategy; Dr. Adel Ahmed Rashad and Aicha Bendia for compound design consultations and preliminary synthesis trial information; and Prof. Wayne Hendrickson for consultation on development of X-ray crystallographic tool compounds.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics approval
This work was performed in accordance with the funding grantor associations’ ethics requirements (NIH, PA Department of Health, and Drexel University), which did not require separate IRB review for this study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gupta, M., Canziani, G., Ang, C. et al. Pharmacophore variants of the macrocyclic peptide triazole inactivator of HIV-1 Env. Med Chem Res 32, 1497–1509 (2023). https://doi.org/10.1007/s00044-023-03092-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-023-03092-0