Abstract
BRI2 is a type II transmembrane protein ubiquitously expressed whose physiological function remains poorly understood. Although several recent important advances have substantially impacted on our understanding of BRI2 biology and function, providing valuable information for further studies on BRI2. These findings have contributed to a better understanding of BRI2 biology and the underlying signaling pathways involved. In turn, these might provide novel insights with respect to neurodegeneration processes inherent to BRI2-related pathologies, namely Familial British and Danish dementias, Alzheimer’s disease, ITM2B-related retinal dystrophy, and multiple sclerosis. In this review, we provided a state-of-the-art outline of BRI2 biology, both in physiological and pathological conditions, and discuss the proposed molecular underlying mechanisms. Overall, the BRI2 knowledge here reviewed is of extreme importance and may contribute to propose BRI2 and/or BRI2 proteolytic fragments as novel therapeutic targets for neurodegenerative diseases, such as Alzheimer’s disease.


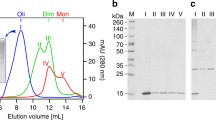
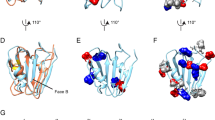
modified by N-glycosylation (N) and ubiquitination (K) at the sites indicated. B Schematic representation of BRI2 processing. BRI2 (immature BRI2 or imBRI2) is cleaved by a proprotein convertase, namely furin, to release a secreted C-terminal 23-residue peptide (BRI2-23). The remaining membrane-bound N-terminal fragment is the mature form of BRI2 (mBRI2). BRI2 undergoes an additional cleavage in its ectodomain exerted by ADAM10 which results in the secretion of a soluble peptide containing the BRICHOS domain (BRI2-BRICHOS) and in a membrane-bound N-terminal fragment (BRI2 NTF). The NTF is subsequently subjected to intramembrane proteolysis cleavage by the SPPL2/b, generating a small, secreted, BRI2 C-terminal peptide (BRI2CTF) and an intracellular domain (BRI2ICD)


Similar content being viewed by others
Data availability
Not applicable.
Code availability
Not applicable.
References
Holton JL, Lashley T, Ghiso J et al (2002) Familial Danish dementia: a novel form of cerebral amyloidosis associated with deposition of both amyloid-Dan and amyloid-beta. J Neuropathol Exp Neurol 61:254–267
Vidal R, Calero M, Révész T et al (2001) Sequence, genomic structure and tissue expression of Human BRI3, a member of the BRI gene family. Gene 266:95–102
Rostagno A, Tomidokoro Y, Lashley T et al (2005) Chromosome 13 dementias. Cell Mol Life Sci 62:1814–1825. https://doi.org/10.1007/s00018-005-5092-5
Pittois K, Wauters J, Bossuyt P et al (1999) Genomic organization and chromosomal localization of the Itm2a gene. Mamm Genome 10:54–56
Deleersnijder W, Hong G, Cortvrindt R et al (1996) Isolation of markers for chondro-osteogenic differentiation using cDNA library subtraction. Molecular cloning and characterization of a gene belonging to a novel multigene family of integral membrane proteins. J Biol Chem 271:19475–19482
Van den Plas D, Merregaert J (2004) In vitro studies on Itm2a reveal its involvement in early stages of the chondrogenic differentiation pathway. Biol Cell 96:463–470. https://doi.org/10.1016/j.biolcel.2004.04.007
Van den Plas D, Merregaert J (2004) Constitutive overexpression of the integral membrane protein Itm2A enhances myogenic differentiation of C2C12 cells. Cell Biol Int 28:199–207. https://doi.org/10.1016/j.cellbi.2003.11.019
Kirchner J, Bevan MJ (1999) ITM2A is induced during thymocyte selection and T cell activation and causes downregulation of CD8 when overexpressed in CD4(+)CD8(+) double positive thymocytes. J Exp Med 190:217–228
Martins F (2017) Functional characterization of novel BRI2 and BRI3 complexes Retrieved from RIA. PhD thesis. The Institutional repository of the University of Aveiro. http://hdl.handle.net/10773/21229
Vidal R, Frangione B, Rostagno A et al (1999) A stop-codon mutation in the BRI gene associated with familial British dementia. Nature 399:776–781. https://doi.org/10.1038/21637
Choi SC, Kim J, Kim TH et al (2001) Cloning and characterization of a type II integral transmembrane protein gene, Itm2c, that is highly expressed in the mouse brain. Mol Cells 12:391–397
Akiyama H, Kondo H, Arai T et al (2004) Expression of BRI, the normal precursor of the amyloid protein of familial British dementia, in human brain. Acta Neuropathol 107:53–58. https://doi.org/10.1007/s00401-003-0783-1
Garringer HJ, Sammeta N, Oblak A et al (2017) Amyloid and intracellular accumulation of BRI2. Neurobiol Aging 52:90–97. https://doi.org/10.1016/j.neurobiolaging.2016.12.018
Del Campo M, Hoozemans JJM, Dekkers L-L et al (2014) BRI2-BRICHOS is increased in human amyloid plaques in early stages of Alzheimer’s disease. Neurobiol Aging 35:1596–1604. https://doi.org/10.1016/j.neurobiolaging.2014.01.007
Choi S-I, Vidal R, Frangione B, Levy E (2004) Axonal transport of British and Danish amyloid peptides via secretory vesicles. FASEB J 18:373–375. https://doi.org/10.1096/fj.03-0730fje
Martins F, Rebelo S, Santos M et al (2016) BRI2 and BRI3 are functionally distinct phosphoproteins. Cell Signal 28:130–144. https://doi.org/10.1016/j.cellsig.2015.10.012
Martin L, Fluhrer R, Reiss K et al (2008) Regulated intramembrane proteolysis of Bri2 (Itm2b) by ADAM10 and SPPL2a/SPPL2b. J Biol Chem 283:1644–1652. https://doi.org/10.1074/jbc.M706661200
Kim SH, Wang R, Gordon DJ et al (1999) Furin mediates enhanced production of fibrillogenic ABri peptides in familial British dementia. Nat Neurosci 2:984–988. https://doi.org/10.1038/14783
Matsuda S, Matsuda Y, Snapp EL, D’Adamio L (2011) Maturation of BRI2 generates a specific inhibitor that reduces APP processing at the plasma membrane and in endocytic vesicles. Neurobiol Aging 32:1400–1408. https://doi.org/10.1016/j.neurobiolaging.2009.08.005
Tsachaki M, Ghiso J, Efthimiopoulos S (2008) BRI2 as a central protein involved in neurodegeneration. Biotechnol J 3:1548–1554. https://doi.org/10.1002/biot.200800247
Martin L, Fluhrer R, Haass C (2009) Substrate requirements for SPPL2b-dependent regulated intramembrane proteolysis. J Biol Chem 284:5662–5670. https://doi.org/10.1074/jbc.M807485200
Schneppenheim J, Huttl S, Mentrup T et al (2014) The intramembrane proteases signal peptide peptidase-like 2a and 2b have distinct functions in vivo. Mol Cell Biol. https://doi.org/10.1128/mcb.00038-14
Fluhrer R, Martin L, Klier B et al (2012) The α-helical content of the transmembrane domain of the British dementia protein-2 (Bri2) determines its processing by signal peptide peptidase-like 2b (SPPL2b). J Biol Chem 287:5156–5163. https://doi.org/10.1074/jbc.M111.328104
Mentrup T, Häsler R, Fluhrer R et al (2015) A cell-based assay reveals nuclear translocation of intracellular domains released by SPPL proteases. Traffic 16:871–892. https://doi.org/10.1111/tra.12287
Artavanis-Tsakonas S, Rand MD, Lake RJ (1999) Notch signaling: cell fate control and signal integration in development. Science 284:770–776
Rebelo S, Domingues SC, Santos M et al (2013) Identification of a novel complex AβPP:Fe65:PP1 that regulates AβPP Thr668 phosphorylation levels. J Alzheimers Dis 35:761–775. https://doi.org/10.3233/JAD-130095
Martins F, Serrano J, Muller T et al (2017) BRI2 processing and its neuritogenic role are modulated by protein phosphatase 1 complexing. J Cell Biochem 118:2752–2763. https://doi.org/10.1002/jcb.25925
Lashley T, Revesz T, Plant G et al (2008) Expression of BRI2 mRNA and protein in normal human brain and familial British dementia: its relevance to the pathogenesis of disease. Neuropathol Appl Neurobiol 34:492–505. https://doi.org/10.1111/j.1365-2990.2008.00935.x
Del Campo M, Teunissen CE (2014) Role of BRI2 in dementia. J Alzheimers Dis 40:481–494. https://doi.org/10.3233/JAD-131364
Tsachaki M, Serlidaki D, Fetani A et al (2011) Glycosylation of BRI2 on asparagine 170 is involved in its trafficking to the cell surface but not in its processing by furin or ADAM10. Glycobiology 21:1382–1388. https://doi.org/10.1093/glycob/cwr097
Ullrich S, Münch A, Neumann S et al (2010) The novel membrane protein TMEM59 modulates complex glycosylation, cell surface expression, and secretion of the amyloid precursor protein. J Biol Chem 285:20664–20674. https://doi.org/10.1074/jbc.M109.055608
Demirkan G, Yu K, Boylan JM et al (2011) Phosphoproteomic profiling of in vivo signaling in liver by the mammalian target of rapamycin complex 1 (mTORC1). PLoS ONE 6:e21729. https://doi.org/10.1371/journal.pone.0021729
Sharma K, D’Souza RCJ, Tyanova S et al (2014) Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep 8:1583–1594. https://doi.org/10.1016/j.celrep.2014.07.036
Wilhelm M, Schlegl J, Hahne H et al (2014) Mass-spectrometry-based draft of the human proteome. Nature 509:582–587. https://doi.org/10.1038/nature13319
Oliveira J, Costa M, De Almeida MSC et al (2017) Protein phosphorylation is a key mechanism in Alzheimer’s Disease. J Alzheimer’s Dis 58(4):953–978. https://doi.org/10.3233/JAD-170176
Rebelo S, Vieira SI, Esselmann H et al (2007) Tyr687 dependent APP endocytosis and Abeta production. J Mol Neurosci. https://doi.org/10.1007/s12031-007-0001-z
Rebelo S, Vieira SI, Esselmann H et al (2007) Tyrosine 687 phosphorylated Alzheimer’s amyloid precursor protein is retained intracellularly and exhibits a decreased turnover rate. Neurodegener Dis. https://doi.org/10.1159/000101831
Vieira SI, Rebelo S, Domingues SC et al (2009) S655 phosphorylation enhances APP secretory traffic. Mol Cell Biochem 328:145–154. https://doi.org/10.1007/s11010-009-0084-7
Oliveira JM, Henriques AG, Martins F et al (2015) Amyloid-β modulates both AβPP and Tau phosphorylation. J Alzheimers Dis. https://doi.org/10.3233/JAD-142664
Udeshi ND, Mani DR, Eisenhaure T et al (2012) Methods for quantification of in vivo changes in protein ubiquitination following proteasome and deubiquitinase inhibition. Mol Cell Proteomics 11:148–159. https://doi.org/10.1074/mcp.M111.016857
Shi Y, Chan DW, Jung SY et al (2011) A data set of human endogenous protein ubiquitination sites. Mol Cell Proteomics 10(M110):002089. https://doi.org/10.1074/mcp.M110.002089
Ciechanover A (1994) The ubiquitin-proteasome proteolytic pathway. Cell 79:13–21
Glickman MH, Ciechanover A (2002) The ubiquitin-proteasome proteolytic pathway: destruction for the sake of construction. Physiol Rev 82:373–428. https://doi.org/10.1152/physrev.00027.2001
Yasukawa T, Tsutsui A, Tomomori-Sato C et al (2020) NRBP1-containing CRL2/CRL4A regulates Amyloid β production by targeting BRI2 and BRI3 for degradation. Cell Rep. https://doi.org/10.1016/j.celrep.2020.02.059
Martins F, Marafona AM, Pereira CD et al (2018) Identification and characterization of the BRI2 interactome in the brain. Sci Rep. https://doi.org/10.1038/s41598-018-21453-3
Tsachaki M, Ghiso J, Rostagno A, Efthimiopoulos S (2010) BRI2 homodimerizes with the involvement of intermolecular disulfide bonds. Neurobiol Aging 31:88–98. https://doi.org/10.1016/j.neurobiolaging.2008.03.004
Tamayev R, Giliberto L, Li W et al (2010) Memory deficits due to familial British dementia BRI2 mutation are caused by loss of BRI2 function rather than amyloidosis. J Neurosci 30:14915–14924. https://doi.org/10.1523/JNEUROSCI.3917-10.2010
Tamayev R, Matsuda S, Fà M et al (2010) Danish dementia mice suggest that loss of function and not the amyloid cascade causes synaptic plasticity and memory deficits. Proc Natl Acad Sci U S A 107:20822–20827. https://doi.org/10.1073/pnas.1011689107
Béïque J-C, Andrade R (2003) PSD-95 regulates synaptic transmission and plasticity in rat cerebral cortex. J Physiol 546:859–867. https://doi.org/10.1113/jphysiol.2002.031369
El-Husseini AE, Schnell E, Chetkovich DM et al (2000) PSD-95 involvement in maturation of excitatory synapses. Science 290:1364–1368
Niethammer M, Kim E, Sheng M (1996) Interaction between the C terminus of NMDA receptor subunits and multiple members of the PSD-95 family of membrane-associated guanylate kinases. J Neurosci 16:2157–2163
Kornau HC, Schenker LT, Kennedy MB, Seeburg PH (1995) Domain interaction between NMDA receptor subunits and the postsynaptic density protein PSD-95. Science 269:1737–1740
Vitale M, Renzone G, Matsuda S et al (2012) Proteomic characterization of a mouse model of familial Danish dementia. J Biomed Biotechnol 2012:728178. https://doi.org/10.1155/2012/728178
Yao W, Yin T, Tambini MD, D’Adamio L (2019) The Familial dementia gene ITM2b/BRI2 facilitates glutamate transmission via both presynaptic and postsynaptic mechanisms. Sci Rep. https://doi.org/10.1038/s41598-019-41340-9
Schwenk J, Pérez-Garci E, Schneider A et al (2016) Modular composition and dynamics of native GABAB receptors identified by high-resolution proteomics. Nat Neurosci. https://doi.org/10.1038/nn.4198
Dinamarca MC, Raveh A, Schneider A et al (2019) Complex formation of APP with GABA B receptors links axonal trafficking to amyloidogenic processing. Nat Commun. https://doi.org/10.1038/s41467-019-09164-3
Fotinopoulou A, Tsachaki M, Vlavaki M et al (2005) BRI2 interacts with amyloid precursor protein (APP) and regulates amyloid beta (Abeta) production. J Biol Chem 280:30768–30772. https://doi.org/10.1074/jbc.C500231200
Matsuda S, Giliberto L, Matsuda Y et al (2005) The familial dementia BRI2 gene binds the Alzheimer gene amyloid-beta precursor protein and inhibits amyloid-beta production. J Biol Chem 280:28912–28916. https://doi.org/10.1074/jbc.C500217200
Matsuda S, Giliberto L, Matsuda Y et al (2008) BRI2 inhibits amyloid beta-peptide precursor protein processing by interfering with the docking of secretases to the substrate. J Neurosci 28:8668–8676. https://doi.org/10.1523/JNEUROSCI.2094-08.2008
Tsachaki M, Fotinopoulou A, Slavi N et al (2013) BRI2 interacts with BACE1 and reduces its cellular levels by reducing the levels of BACE1 mRNA and inducing its degradation through the lysosomal pathway. Curr Alzheimer Res 10:532–541
Kim J, Miller VM, Levites Y et al (2008) BRI2 (ITM2b) inhibits Abeta deposition in vivo. J Neurosci 28:6030–6036. https://doi.org/10.1523/JNEUROSCI.0891-08.2008
Peng S, Fitzen M, Jörnvall H, Johansson J (2010) The extracellular domain of Bri2 (ITM2B) binds the ABri peptide (1–23) and amyloid beta-peptide (Abeta1-40): implications for Bri2 effects on processing of amyloid precursor protein and Abeta aggregation. Biochem Biophys Res Commun 393:356–361. https://doi.org/10.1016/j.bbrc.2009.12.122
Willander H, Presto J, Askarieh G et al (2012) BRICHOS domains efficiently delay fibrillation of amyloid β-peptide. J Biol Chem 287:31608–31617. https://doi.org/10.1074/jbc.M112.393157
Kilger E, Buehler A, Woelfing H et al (2011) BRI2 protein regulates β-amyloid degradation by increasing levels of secreted insulin-degrading enzyme (IDE). J Biol Chem 286:37446–37457. https://doi.org/10.1074/jbc.M111.288373
Morelli L, Llovera RE, Alonso LG et al (2005) Insulin-degrading enzyme degrades amyloid peptides associated with British and Danish familial dementia. Biochem Biophys Res Commun. https://doi.org/10.1016/j.bbrc.2005.05.020
Fleischer A, Ayllón V, Dumoutier L et al (2002) Proapoptotic activity of ITM2B(s), a BH3-only protein induced upon IL-2-deprivation which interacts with Bcl-2. Oncogene 21:3181–3189. https://doi.org/10.1038/sj.onc.1205464
Fleischer A, Ayllon V, Rebollo A (2002) ITM2Bs regulates apoptosis by inducing loss of mitochondrial membrane potential. Eur J Immunol. https://doi.org/10.1002/1521-4141(200212)32:12%3c3498::AID-IMMU3498%3e3.0.CO;2-C
Fleischer A, Rebollo A (2004) Induction of p53-independent apoptosis by the BH3-only protein ITM2Bs. FEBS Lett. https://doi.org/10.1016/S0014-5793(03)01519-9
Rengaraj D, Gao F, Liang X-H, Yang Z-M (2007) Expression and regulation of type II integral membrane protein family members in mouse male reproductive tissues. Endocrine 31:193–201
Rengaraj D, Liang XH, Gao F et al (2008) Differential expression and regulation of integral membrane protein 2b in rat male reproductive tissues. Asian J Androl. https://doi.org/10.1111/j.1745-7262.2008.00360.x
Han C, Park I, Lee B et al (2011) Identification of heat shock protein 5, calnexin and integral membrane protein 2B as Adam7-interacting membrane proteins in mouse sperm. J Cell Physiol 226:1186–1195. https://doi.org/10.1002/jcp.22444
Mandal AK, Mount DB (2019) Interaction between ITM2B and GLUT9 links Urate transport to neurodegenerative disorders. Front Physiol. https://doi.org/10.3389/fphys.2019.01323
Davies MN, Verdi S, Burri A et al (2015) Generalised anxiety disorder—a twin study of genetic architecture, genome-wide association and differential gene expression. PLoS ONE. https://doi.org/10.1371/journal.pone.0134865
Matsuda S, Senda T (2019) BRI2 as an anti-Alzheimer gene. Med Mol Morphol 52:1–7. https://doi.org/10.1007/s00795-018-0191-1
Dolfe L, Tambaro S, Tigro H et al (2018) The Bri2 and Bri3 BRICHOS domains interact differently with Aβ42 and Alzheimer Amyloid plaques. J Alzheimer’s Dis Reports. https://doi.org/10.3233/adr-170051
Poska H, Haslbeck M, Kurudenkandy FR et al (2016) Dementia-related Bri2 BRICHOS is a versatile molecular chaperone that efficiently inhibits A 42 toxicity in Drosophila. Biochem J 473:3683–3704. https://doi.org/10.1042/BCJ20160277
Kim SH, Creemers JWM, Chu S et al (2002) Proteolytic processing of familial British dementia-associated BRI variants: evidence for enhanced intracellular accumulation of amyloidogenic peptides. J Biol Chem. https://doi.org/10.1074/jbc.M108739200
Ghiso JA, Holton J, Miravalle L et al (2001) Systemic amyloid deposits in familial British dementia. J Biol Chem 276:43909–43914. https://doi.org/10.1074/jbc.M105956200
Vidal R, Revesz T, Rostagno A et al (2000) A decamer duplication in the 3’ region of the BRI gene originates an amyloid peptide that is associated with dementia in a Danish kindred. Proc Natl Acad Sci U S A 97:4920–4925. https://doi.org/10.1073/pnas.080076097
Plant GT, Révész T, Barnard RO et al (1990) Familial cerebral amyloid angiopathy with nonneuritic amyloid plaque formation. Brain 113:721–747
Mead S, James-Galton M, Revesz T et al (2000) Familial British dementia with amyloid angiopathy: early clinical, neuropsychological and imaging findings. Brain 123:975–991
Revesz T, Holton JL, Doshi B et al (1999) Cytoskeletal pathology in familial cerebral amyloid angiopathy (British type) with non-neuritic amyloid plaque formation. Acta Neuropathol 97:170–176
Strömgren E, Dalby A, Dalby MA, Ranheim B (1970) Cataract, deafness, cerebellar ataxia, psychosis and dementia—a new syndrome. Acta Neurol Scand 46(Suppl 43):261–262
Tomidokoro Y, Lashley T, Rostagno A et al (2005) Familial Danish dementia: co-existence of Danish and Alzheimer amyloid subunits (ADan AND A{beta}) in the absence of compact plaques. J Biol Chem 280:36883–36894. https://doi.org/10.1074/jbc.M504038200
El-Agnaf O, Gibson G, Lee M et al (2004) Properties of neurotoxic peptides related to the Bri gene. Protein Pept Lett 11:207–212
Gibson G, Gunasekera N, Lee M et al (2004) Oligomerization and neurotoxicity of the amyloid ADan peptide implicated in familial Danish dementia. J Neurochem 88:281–290
Rostagno A, Ghiso J (2008) Preamyloid lesions and cerebrovascular deposits in the mechanism of dementia: lessons from non-beta-amyloid cerebral amyloidosis. Neurodegener Dis 5:173–175. https://doi.org/10.1159/000113694
Bradt BM, Kolb WP, Cooper NR (1998) Complement-dependent proinflammatory properties of the Alzheimer’s disease beta-peptide. J Exp Med 188:431–438
Yin T, Yao W, Lemenze AD, D’Adamio L (2020) Danish and British dementia ITM2b/BRI2 mutations reduce BRI2 protein stability and impair glutamatergic synaptic transmission. J Biol Chem. https://doi.org/10.1074/jbc.RA120.015679
Tamayev R, D’Adamio L (2012) Memory deficits of British dementia knock-in mice are prevented by Aβ-precursor protein haploinsufficiency. J Neurosci 32:5481–5485. https://doi.org/10.1523/JNEUROSCI.5193-11.2012
Tamayev R, Matsuda S, Giliberto L et al (2011) APP heterozygosity averts memory deficit in knockin mice expressing the Danish dementia BRI2 mutant. EMBO J 30:2501–2509. https://doi.org/10.1038/emboj.2011.161
Matsuda S, Tamayev R, D’Adamio L (2011) Increased AβPP processing in familial Danish dementia patients. J Alzheimers Dis 27:385–391. https://doi.org/10.3233/JAD-2011-110785
Biundo F, Ishiwari K, Del Prete D, D’Adamio L (2016) Deletion of the γ-secretase subunits Aph1B/C impairs memory and worsens the deficits of knock-in mice modeling the Alzheimer-like familial Danish dementia. Oncotarget 7:11923–11944. https://doi.org/10.18632/oncotarget.7389
Garringer HJ, Murrell J, D’Adamio L et al (2010) Modeling familial British and Danish dementia. Brain Struct Funct 214:235–244. https://doi.org/10.1007/s00429-009-0221-9
Cantlon A, Frigerio CS, Walsh DM (2015) Lessons from a rare familial dementia: amyloid and beyond. J Park Dis Alzheimer’s Dis 2(1):12. https://doi.org/10.13188/2376-922x.1000009
Chen WT, Lu A, Craessaerts K et al (2020) Spatial transcriptomics and in situ sequencing to study Alzheimer’s disease. Cell. https://doi.org/10.1016/j.cell.2020.06.038
Del Campo M, Oliveira CR, Scheper W et al (2015) BRI2 ectodomain affects Aβ42 fibrillation and tau truncation in human neuroblastoma cells. Cell Mol Life Sci 72:1599–1611. https://doi.org/10.1007/s00018-014-1769-y
Beers MF, Lomax CA, Russo SJ (1998) Synthetic processing of surfactant protein C by alevolar epithelial cells: the COOH terminus of proSP-C is required for post-translational targeting and proteolysis. J Biol Chem. https://doi.org/10.1074/jbc.273.24.15287
Sánchez-Pulido L, Devos D, Valencia A (2002) BRICHOS: a conserved domain in proteins associated with dementia, respiratory distress and cancer. Trends Biochem Sci 27:329–332
Hedlund J, Johansson J, Persson B (2009) BRICHOS—a superfamily of multidomain proteins with diverse functions. BMC Res Notes. https://doi.org/10.1186/1756-0500-2-180
Knight SD, Presto J, Linse S, Johansson J (2013) The BRICHOS domain, amyloid fibril formation, and their relationship. Biochemistry 52:7523–7531. https://doi.org/10.1021/bi400908x
Willander H, Hermansson E, Johansson J, Presto J (2011) BRICHOS domain associated with lung fibrosis, dementia and cancer—a chaperone that prevents amyloid fibril formation? FEBS J 278:3893–3904
Cohen SIA, Arosio P, Presto J et al (2015) A molecular chaperone breaks the catalytic cycle that generates toxic Aβ oligomers. Nat Struct Mol Biol. https://doi.org/10.1038/nsmb.2971
Chen G, Abelein A, Nilsson HE et al (2017) Bri2 BRICHOS client specificity and chaperone activity are governed by assembly state. Nat Commun 8:2081. https://doi.org/10.1038/s41467-017-02056-4
Dolfe L, Winblad B, Johansson J, Presto J (2016) BRICHOS binds to a designed amyloid-forming β-protein and reduces proteasomal inhibition and aggresome formation. Biochem J. https://doi.org/10.1042/BJ20150920
Tambaro S, Galan-Acosta L, Leppert A et al (2019) Blood-brain and blood-cerebrospinal fluid passage of BRICHOS domains from two molecular chaperones in mice. J Biol Chem 294:2606–2615. https://doi.org/10.1074/jbc.RA118.004538
Galan-Acosta L, Sierra C, Leppert A et al (2020) Recombinant BRICHOS chaperone domains delivered to mouse brain parenchyma by focused ultrasound and microbubbles are internalized by hippocampal and cortical neurons. Mol Cell Neurosci. https://doi.org/10.1016/j.mcn.2020.103498
Audo I, Bujakowska K, Orhan E et al (2014) The familial dementia gene revisited: A missense mutation revealed by whole-exome sequencing identifies ITM2B as a candidate gene underlying a novel autosomal dominant retinal dystrophy in a large family. Hum Mol Genet. https://doi.org/10.1093/hmg/ddt439
Nassisi M, Wohlschlegel J, Liu B et al (2020) Deep-phenotyping and further insights in ITM2B-related retinal dystrophy. Retina. https://doi.org/10.1097/IAE.0000000000002953
Giannoccaro MP, Bartoletti-Stella A, Piras S et al (2018) The first historically reported Italian family with FTD/ALS teaches a lesson on C9orf72 RE: clinical heterogeneity and Oligogenic inheritance. J Alzheimer’s Dis. https://doi.org/10.3233/JAD-170913
Harris VK, Diamanduros A, Good P et al (2010) Bri2-23 is a potential cerebrospinal fluid biomarker in multiple sclerosis. Neurobiol Dis. https://doi.org/10.1016/j.nbd.2010.06.007
Latil A, Chêne L, Mangin P et al (2003) Extensive analysis of the 13q14 region in human prostate tumors: DNA analysis and quantitative expression of genes lying in the interval of deletion. Prostate 57:39–50. https://doi.org/10.1002/pros.10272
Baron BW, Baron RM, Baron JM (2015) The ITM2B (BRI2) gene is a target of BCL6 repression: Implications for lymphomas and neurodegenerative diseases. Biochim Biophys Acta 1852:742–748. https://doi.org/10.1016/j.bbadis.2014.12.018
Lee S, Jeong J, Majewski T et al (2007) Forerunner genes contiguous to RB1 contribute to the development of in situ neoplasia. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.0701771104
Creytens D, Van Gorp J, Savola S et al (2014) Atypical spindle cell lipoma: a clinicopathologic, immunohistochemical, and molecular study emphasizing its relationship to classical spindle cell lipoma. Virchows Arch. https://doi.org/10.1007/s00428-014-1568-8
Creytens D, Mentzel T, Ferdinande L et al (2017) Atypical pleomorphic lipomatous tumor: a clinicopathologic, immunohistochemical and molecular study of 21 cases, emphasizing its relationship to atypical spindle cell lipomatous tumor and suggesting a morphologic spectrum (Atypical spindle cell/Pleomorphi. Am J Surg Pathol. https://doi.org/10.1097/PAS.0000000000000936
Acknowledgements
This work was supported by the Instituto de Biomedicina‐iBiMED (UIDB/04501/2020 and POCI‐01‐0145‐FEDER‐007628); and the Fundação para a Ciência e Tecnologia (FCT) of the Ministério da Educação e Ciência, the COMPETE program, QREN and the EU (Fundo Europeu de Desenvolvimento Regional), Stiftelsen För Gamla Tjänarinnor, and Demensfonden from Demensförbundet. Authors acknowledge support from Integrated Programme of SR&TD “pAGE–Protein aggregation across lifespan” (CENTRO‐01‐0145‐FEDER‐000003), co‐funded by Centro 2020 Program, Portugal 2020, EU, through the European Regional Development Fund.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Martins, F., Santos, I., da Cruz e Silva, O.A.B. et al. The role of the integral type II transmembrane protein BRI2 in health and disease. Cell. Mol. Life Sci. 78, 6807–6822 (2021). https://doi.org/10.1007/s00018-021-03932-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-021-03932-5