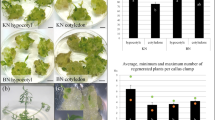
Abstract
Cellulose is one of many important polymers in plants. Cellulose is made of repeat units of the monomer glucose. Cellulose is a major industrial biopolymer in the forest products, textile, and chemical industries. It also forms a large portion of the biomass useful in the generation of energy. Moreover, cellulose-based biomass is a renewable energy source that can be used for the generation of ethanol as a fuel. Cellulose is synthesized by a variety of living organisms such as plants and algae. It is the major component of plant cell walls with secondary cell walls having a much higher content of cellulose. The relationship between cellulose and lignin biosynthesis is complicated, but it is confirmed that inhibition of lignin biosynthesis in transgenic trees will increase cellulose biosynthesis and plant growth. Cellulose accumulation may be increased by down-regulating 4-coumarate:coenzyme A ligase (4CL, EC 6.2.1.12) as shown in transgenic aspen. There is no similar reports on down-regulating 4CL in transgenic conifers. Based on our establishedAgrobacterium tumefaciens-mediated transformation system in loblolly pine, we are able to produce antisense 4-CL transgenic loblolly pine which is predicted to have increasing cellulose accumulation. The overall objective of this project is to genetically engineer forest tree species such as loblolly pine with reduced amount of lignin and increased cellulose content. The research strategy includes: (1) isolate the 4-coumarate:coenzyme A ligase gene from loblolly pine seedlings by reverse transcription-polymerase chain reaction (RT-PCR) and Rapid Amplification of cDNA Ends-Polymerase Chain Reaction (RACE-PCR) techniques from the cDNA library; (2) construct binary expression vectors with antisense 4CL coding sequences and introduce antisense constructs of the 4-coumarate:coenzyme A ligase gene cloned from loblolly pine into the loblolly pine to down regulate the 4-coumarate:coenzyme A ligase gene expression; (3) study the effect of the antisense transgene expression on lignin content, cellulose accumulation, and loblolly pine biomass; and (4) select fast growth and high cellulose accumulation transgenic loblolly pine lines for future commercial application.
Similar content being viewed by others
References
Abbott, J.C., Barakate, A., Pincon G., Legrand, M., Lapierre, C.,et al. 2002. Simultaneous suppression of multiple genes by single transgenes. Down-regulation of three unrelated lignin biosynthetic genes in tobacco [J]. Plant Physiol.,128: 844–53.
Allina, S.M., Pri-Hadash, A., Theilmann, D.A., Ellis, B.E., Douglas, C.J. 1998. 4-Coumarate: coenzymeAligase in hybrid poplar [J]. Plant Physiol.,116: 743–54.
Amor, Y., Haigler, C.H., Johnson, S., Wainscott, M., Delmer, D.P. 1995. Amembraneassociated form of sucrose synthase and its potential role in synthesis of cellulose and callose in plants [J]. Proc. Natl. Acad. Sci. USA,92: 9353–57.
Anterola, A.M., Jeon, J-H., Davin, L.B., Lewis, N.G. 2002. Transcriptional control of monolignol biosynthesis inPinus taeda. Factors affecting monolignol ratios and carbon allocation in phenylpropanoidmetabolism. J. Biol. Chem.,277: 18272–80.
Arioli, T., Peng, L.C., Betzner, A.S., Burn, J., Wittke, W.,et al. 1998. Molecular analysis of cellulose biosynthesis inArabidopsis[J]. Science,279: 717–20.
Atanassova, R., Favet, N., Martz, F., Chabbert, B., Tollier, M.T.,et al. 1995. Altered lignin composition in transgenic tobacco expressingO-methyltransferase sequences in sense and antisense orientation [J]. Plant J.,8: 465–77.
Baucher, M., Bernard-Vailh'e, M.A., Chabbert, B., Besle, J-M., Opsome, C.,et al. 1999. Down-regulation of cinnamyl alcohol dehydrogenase in transgenic alfalfa (Medicago sativa L.) and the impact on lignin composition and digestibility [J]. Plant Mol. Biol.,39: 437–47.
Blount, J.W., Masoud, S., Sumner, L.W., Huhman, D., Dixon, R.A. 2002. Overexpression of cinnamate 4-hydroxylase leads to increased accumulation of acetosyringone in elicited tobacco cellsuspension cultures [J]. Planta,214: 902–10.
Boerjan, W., Ralph, J., Baucher, M. 2003. Lignin biosynthesis [J]. Annu Rev Plant Biol.,54: 519–46.
Bolwell, G.P., Gramer, C.L., Lamb, C.J., Schuch, W., Dixon, R.A. 1986. L-Phenylalanine ammonia-lyase fromPhaseolus vulgaris. Modulation of the levels of active enzyme bytrans-cinnamic acid [J]. Planta,169: 97–107.
Brown, R.M.Jr. 1996. The biosynthesis of cellulose [J]. J. Macromol. Sci. Pure Appl. Chem.,A33(10): 1345–73.
Brown, R.M. Jr, Montezinos, D. 1976. Cellulose microfibrils: visualization of biosynthetic and orienting complexes in association with the plasma membrane [J]. Proc. Natl. Acad. Sci. USA,73: 143–147.
Brown, R.M. Jr, Saxena, I.M., Kudlicka, K. 1997. Cellulose biosynthesis in higher plants [J]. Trends Plant Sci.,1: 149–156.
Brummell, D.A., Catala, C., Lashbrook, C.C., Bennett, A.B. 1997. A membrane-anchored E-type endo-1,4-beta-glucanase is localized on Golgi and plasma membranes of higher plants [J]. Proc. Natl. Acad. Sci. USA,94: 4794–4799.
Carpita, N.C. 1996. Structure and biogenesis of the cell walls of grasses[J]. Annu. Rev. Plant Physiol. Plant Mol. Biol.,47: 445–476.
Carpita, N., Vergara, C. 1998. A recipe for cellulose [J]. Science,279: 672–673.
Chabannes, M., Barakate, A., Lapierre, C., Marita, J.M., Ralph, J.,et al. 2001a. Strong decrease in lignin content without significant alteration of plant development is induced by simultaneous downregulation of cinnamoyl CoA reductase (CCR) and cinnamyl alcohol dehydrogenase (CAD) in tobacco plants [J]. Plant J.,28: 257–70.
Chabannes, M., Ruel, K., Yoshinaga, A., Chabbert, B., Jauneau, A.,et al. 2001b.In situ analysis of lignins in transgenic tobacco reveals a differential impact of individual transformations on the spatial patterns of lignin deposition at the cellular and subcellular levels [J]. Plant J.,28: 271–282.
Chanzy, H., Henrissat, B. 1985. Unidirectional degradation of Valonia cellulose microcrystals subjected to cellulase action [J]. FEBS Lett.184: 285–288.
Chapple C. 1998. Molecular-genetic analysis of plant cytochrome P450-dependent monooxygenases.Annu. Rev. Plant Physiol. Plant Mol. Biol. 49: 311–43.
Chen, C., Meyermans, H., Burggraeve, B., De Rycke, R.M., Inoue, K.,et al. 2000. Cell-specific and conditional expression of caffeoyl-CoAO-methyltransferase in Poplar [J]. Plant Physiol.,123: 853–867.
Chen, F., Yasuda, S., Fukushima, K. 1999. Evidence for a novel biosynthetic pathway that regulates the ratio of syringyl to guaiacyl residues in lignin in the differentiating xylem ofMagnolia kobus DC [J]. Planta,207: 597–603.
Christensen, J.H., Baucher, M., O'Connell, A.P., Van Montagu, M., Boerjan, W. 2000. Control of lignin biosynthesis [C]. In: SM Jain, SC Minocha (eds),Molecular Biology of Woody Plants, Volume1, For. Sci., 64: 227–267. Dordrecht: Kluwer. 520 pp.
Christensen, J.H., Overmey, S., Rohde, A., Ardiles Diaz, W., Bauw, G.,et al. 2001. The syringaldazine-oxidizing peroxidase PXP 3–4 from poplar xylem: cDNA isolation, characterization and expression [J]. Plant Mol. Biol.,47: 581–93.
Delmer, D.P. 1999 Cellulose Biosynthesis: Exciting Times for A Difficult Field of Study [J]. Annu. Rev. Plant Physiol. Plant Mol. Biol.,50: 245–276.
Delmer, D.P. 1987. Cellulose biosynthesis [J]. Annu. Rev. Plant Physiol.,38: 259–290.
Delmer, D.P., Amor, Y. 1995. Cellulose biosynthesis [J]. Plant Cell,7: 987–1000.
Dixon, R.A., Chen, F., Guo, D., Parvathi, K. 2001. The biosynthesis of monolignols: a “metabolic grid,” or independant pathways to guaiacyl and syringyl units? [J] Phytochemistry,57: 1069–1084.
Doblin, M.S., Kurek, I., Jacob-Wilk, D., Delmer, D.P. 2002. Cellulose Biosynthesis in Plants: from Genes to Rosettes [J]. Plant Cell Physiol.,43(12): 1407–1420.
Donaldson, L.A. 2001. Lignification and lignin topochemistry—an ultrastructural view [J]. Phytochemistry,57: 859–873.
Ehlting, J., B'uttner, D., Wang, Q., Douglas, C.J., Somssich, I.E., Kombrink, E. 1999. Three 4-coumarate:coenzyme A ligases inArabidopsis thaliana represent two evolutionarily divergent classes in angiosperms [J]. Plant J.,19: 9–20.
Franke R, Hemm MR, Denault JW, Ruegger MO, Humphreys JM, Chapple C. 2002. Changes in secondary metabolism and deposition of an unusual lignin in theref8 mutant ofArabidopsis [J]. Plant J.,30: 47–59.
Gardner, K.H., Blackwell, J. 1974. The structure of native cellulose [J]. Biopolymers,13: 1975–2001.
Grabber, J.H., Ralph, J., Hatfield, R.D., Quideau, S. 1997.p-hydroxyphenyl, guaiacyl, and syringyl lignins have similar inhibitory effects on wall degradability [J]. J. Agric. Food Chem.,45: 2530–2532.
Guo, D., Chen, F., Inoue, K., Blount, J.W., Dixon, R.A. 2001. Downregulation of caffeic acid 3-O-methyltransferase and caffeoyl CoA 3-O-methyltransferase in transgenic alfalfa: impacts on lignin structure and implications for the biosynthesis of G and S lignin [J]. Plant Cell,13: 73–88.
Halpin, C., Holt, K., Chojecki, J., Oliver, D., Chabbert, B.,et al. 1998.Brown-midrib maize (bml)—a mutation affecting the cinnamyl alcohol dehydrogenase gene [J]. Plant J.,14: 545–553.
Hibino, T., Takabe, K., Kawazu, T., Shibata, D., Higuchi, T. 1995. Increase of cinnamaldehyde groups in lignin of transgenic tobacco plants carrying an antisense gene for cinnamyl alcohol dehydrogenase [J]. Biosci. Biotech. Biochem.,59: 929–931.
Howles, P.A., Sewalt, V.J.H., Paiva, N.L., Elkind, Y., Bate, N.J.,et al. 1996. Overexpression of L-phenylalanine ammonialyase in transgenic tobacco plants reveals control points for flux into phenylpropanoid biosynthesis [J]. Plant Physiol.,112: 1617–1624.
Hu, W-J., Harding, S.A., Lung, J., Popko, J.L., Ralph, J.,et al. 1999. Repression of lignin biosynthesis promotes cellulose accumulation and growth in transgenic trees [J]. Nat. Biotechnol.,17: 808–812.
Humphreys, J.M., Hemm, M.R., Chapple, C. 1999. New routes for lignin biosynthesis defined by biochemical characterization of recombinant ferulate 5-hydroxylase, a multifunctional cytochrome P450-dependent monooxygenase [J]. Proc. Natl. Acad. Sci. USA,96: 10045–10050.
Itoh T. 1990. Cellulose synthesizing complexes in some giant marine algae [J]. J. Cell Sci.,95: 309–319.
Jones L, Ennos AR, Turner SR. 2001. Cloning and characterization ofirregular xylem4 (irx4): a severely lignin deficient mutant ofArabidopsis [J]. Plant J.,26: 205–216.
Kajita, S., Hishiyama, S., Tomimura, Y., Katayama, Y., Omori, S. 1997. Structural characterization of modified lignin in transgenic tobacco plants in which the activity of 4-coumarate: coenzymeAligase is depressed [J]. Plant Physiol.,114: 871–879.
Kajita, S., Katayama, Y., Omori, S. 1996. Alterations in the biosynthesis of lignin in transgenic plants with chimeric genes for 4-coumarate:coenzyme A ligase [J]. Plant Cell Physiol.,37: 957–965.
Kawagoe, Y., Delmer, D.P. 1997. Pathways and genes involved in cellulose biosynthesis [J]. Genet. Eng.,19: 63–87.
Keller, B., Templeton, M.D., Lamb, C.J. 1989. Specific localization of a plant cell wall glycine-rich protein in protoxylem cells of the vascular system [J]. Proc. Natl. Acad. Sci. USA,86: 1529–1533.
Lee, D., Meyer, K., Chapple, C., Douglas, C.J. 1997. Antisense suppression of 4-coumarate: coenzyme A ligase activity inArabidopsis leads to altered lignin subunit composition [J]. Plant Cell,9: 1985–1998.
Li, L., Cheng, X.F., Leshkevich, J., Umezawa, T., Harding, S.A., Chiang, V.L., 2001. The last step of syringyl monolignol biosynthesis in angiosperms is regulated by a novel gene encoding sinapyl alcohol dehydrogenase [J]. Plant Cell,13: 1567–1585.
Lin, F.C., Brown, R.M. Jr. 1989. Purification of cellulose synthase from Acetobacter xylinum [C]. In: C. Scheurch (ed.), Cellulose and Wood—Chemistry and Technology. New York: Wiley, pp. 473–492
MacKay, J.J., O'Malley, D.M., Presnell, T., Booker, F.L., Campbell, M.M.,et al. 1997. Inheritance, gene expression, and lignin characterization in a mutant pine defi-cient in cinnamyl alcohol dehydrogenase [J]. Proc. Natl. Acad. Sci. USA,94: 8255–8260.
Marita, J.M., Ralph, J., Hatfield, R.D., Chapple, C. 1999. NMR characterization of lignins inArabidopsis altered in the activity of ferulate 5-hydroxylase [J]. Proc. Natl. Acad. Sci. USA,96: 12328–12332.
Meyer, K., Shirley, A.M., Cusumano, J.C., Bell-Lelong, D.A., Chapple, C. 1998. Lignin monomer composition is determined by the expression of a cytochrome P450-dependent monooxygenase inArabidopsis[J]. Proc. Natl. Acad. Sci. USA,95: 6619–6623.
Neish, A.C. 1968. Monomeric intermediates in the biosynthesis of lignin [C]. In: K. Freudenberg and A.C. Neish (eds), Constitution and Biosynthesis of Lignin, New York: Springer-Verlag, pp. 3–43.
Osakabe, K., Tsao, C.C., Li, L., Popko, J.L., Umezawa, T.,et al. 1999. Coniferyl aldehyde 5-hydroxylation and methylation direct syringyl lignin biosynthesis in angiosperms [J]. Proc. Natl. Acad. Sci. USA,96: 8955–8960.
Parvathi, K., Chen, F., Guo, D., Blount, J.W., Dixon, R.A. 2001. Substrate preferences ofO-methyltransferases in alfalfa suggest new pathways for 3-O-methylation of monolignols [J]. Plant J.,25: 193–202.
Pear, J., Kawagoe, Y., Schreckengost, W., Delmer, D.P., Stalker, D. 1996. Higher plants contain homologs of the bacterial CelA genes encoding the catalytic subunit of the cellulose synthase [J]. Proc. Natl. Acad. Sci. USA93: 12637–12642.
Ralph, J., MacKay, J.J., Hatfield, R.D., O'Malley, D.M., Whetten, R.W., Sederoff, R.R. 1997. Abnormal lignin in a loblolly pine mutant [J]. Science,277: 235–39.
Ranocha, P., McDougall, G., Hawkins, S., Sterjiades, R., Borderies, G.,et al. 1999. Biochemical characterization, molecular cloning and expression of laccases—a divergent gene family—in poplar [J]. Eur. J. Biochem.,259: 485–95.
Samuels, A.L., Rensing, K., Douglas, C.J., Mansfield, S., Dharmawardhana, P., Ellis, B. 2002. Cellular machinery of wood production: differentiation of secondary xylem inPinus contorta var.latifolia [J]. Planta,216: 72–82.
Sarkanen, K.V., Ludwig, C.H., ed. 1971. Lignins: Occurrence, Formation, Structure, and Reactions [M]. New York Wiley-Intersci. 916 pp.
Sederoff, R.R., MacKay, J.J., Ralph, J., Hat-field, R.D. 1999. Unexpected variation in lignin [J]. Curr. Opin. Plant Biol.,2: 145–52.
Sewalt, V.J.H., Ni W, Blount, J.W., Jung, H.G., Masoud, S.A.,et al. 1997. Reduced lignin content and altered lignin composition in transgenic tobacco down-regulated in expression of L-phenylalanine ammonialyase or cinnamate 4-hydroxylase [J]. Plant Physiol.,115: 41–50.
Tang, W., Sedoroff, R., Whetten, R. 2001. Regeneration of transgenic loblolly pine (Pinus taeda L.) from zygotic embryos transformed withAgrobacterium tumefaciens [J]. Planta.,213: 981–989.
Thelen, M.T., Delmer, D.P. 1986. Gelelectrophoretic separation, detection, and characterization of plant and bacterial UDP-glucose glucosyltransferases [J]. Plant Physiol.,81: 913–18.
VanderHart, D.L., Atalla, R.H. 1986. In Cellulose: Structure, Modification and Hydrolysis, ed. RA Young, RM Rowell, pp. 88–118. New York: Wiley-Intersci.
Voo, K.S., Whetten, R.W., O'Malley, D.M., Sederoff, R.R. 1995. 4-Coumarate:Coenzyme A ligase from loblolly pine xylem: Isolation, Characterization, and Complementary DNA Cloning [J]. Plant Physiol.,108: 85–97.
Whetten, R.W., MacKay, J.J., Sederoff, R.R. 1998. Recent advances in understanding lignin biosynthesis [J]. Annu. Rev. Plant Physiol. Plant Mol. Biol.,49: 585–609.
Zhong, R., Morrison, W.H. III, Himmelsbach, D.S., Poole, F.L. II, Ye, Z-H. 2000. Essential role of caffeoyl coenzyme AOmethyltransferase in lignin biosynthesis in woody poplar plants [J]. Plant Physiol.,124: 563–577.
Zhong, R., Morrison, W.H. III, Negrel, J., Ye, Z-H. 1998. Dual methylation pathways in lignin biosynthesis [J]. Plant Cell.,10: 2033–2046.
Zubieta, C., Kota, P., Ferrer, J-L., Dixon, R.A., Noel, J.P. 2002. Structural basis for the modulation of lignin monomer methylation by caffeic acid/5-hydroxyferulic acid 3/5-O-methyltransferase [J]. Plant Cell,14: 1265–1277.
Author information
Authors and Affiliations
Corresponding author
Additional information
Biography: TANG Wei (1964-), male, Ph. Doctor, Assistant Professor, Department of Biology, Howell Science Complex, East Carolina University, Greenville, NC 27858-4353, USA.
Responsible: editor: Chai Ruihai
Rights and permissions
About this article
Cite this article
Wei, T., Nelson, A. & Johnson, E. Increasing cellulose production and transgenic plant growth in forest tree species. Journal of Forestry Research 16, 67–72 (2005). https://doi.org/10.1007/BF02856860
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF02856860