Abstract
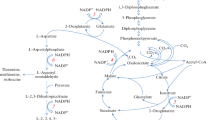
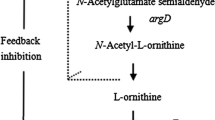
Most Pseudomonas aeruginosa PAO mutants which were unable to utilize l-arginine as the sole carbon and nitrogen source (aru mutants) under aerobic conditions were also affected in l-ornithine utilization. These aru mutants were impaired in one or several enzymes involved in the conversion of N2-succinylornithine to glutamate and succinate, indicating that the latter steps of the arginine succinyltransferase pathway can be used for ornithine catabolism. Addition of aminooxyacetate, an inhibitor of the N2-succinylornithine 5-aminotransferase, to resting cells of P. aeruginosa in ornithine medium led to the accumulation of N2-succinylornithine. In crude extracts of P. aeruginosa an ornithine succinyltransferase (l-ornithine:succinyl-CoA N2-succinyltransferase) activity could be detected. An aru mutant having reduced arginine succinyltransferase activity also had correspondingly low levels of ornithine succinyltransferase. Thus, in P. aeruginosa, these two activities might be due to the same enzyme, which initiates aerobic arginine and ornithine catabolism.
Similar content being viewed by others
Abbreviations
- OAT:
-
ornithine 5-aminotransferase
- SOAT:
-
N2-succinylornithine 5-aminotransferase
- Oru:
-
ornithine utilization
- Aru:
-
arginine utilization
References
Früh R, Haas D, Leisinger T (1985) Altered control of glutamate dehydrogenase in ornithine utilization mutant of Pseudomonas aeruginosa. Arch Microbiol 141:170–176
Haas D, Holloway BW, Schamböck A, Leisinger T (1977) The genetic organisation of arginine biosynthesis in Pseudomonas aeruginosa. Mol Gen Genet 154:7–22
Haas D, Matsumoto H, Moretti P, Stalon V, Mercenier A (1984) Arginine degradation in Pseudomonas aeruginosa mutants blocked in two arginine catabolic pathways. Mol Gen Genet 193:437–444
Haas D, Jann A, Reimmann C, Lüthi E, Leisinger T (1987) Chromosome organization in Pseudomonas aeruginosa. Clustering and scattering of genes specifying four arginine catabolic pathways. Antibiot Chemother 39:256–263
Hoshino T, Nishio K (1982) Isolation and characterization of a Pseudomonas aeruginosa PAO mutant defective in the structural gene for the LIVAT-binding protein. J Bacteriol 151:729–786
Jann A, Stalon V, Vander Wauven C, Leisinger T, Haas D (1986) N2-Succinylated intermediates in an arginine catabolic pathway of Pseudomonas aeruginosa. Proc Natl Acad Sci USA 83:4937–4941
Krishna RV, Beilstein P, Leisinger T (1979) Biosynthesis of proline in Pseudomonas aeruginosa. Properties of γ-glutamylphosphate reductase and 1-pyrroline-5-carboxylate reductase. Biochem J 181:223–230
Leisinger T, O'Sullivan C, Haas D (1974) Arginine analogues: effect on growth of the first two enzymes of the arginine pathway in Pseudomonas aeruginosa. J Gen Microbiol 84:253–260
Meile L, Soldati L, Leisinger T (1982) Regulation of proline catabolism in Pseudomonas aeruginosa PAO. Arch Microbiol 132:189–193
Mercenier A, Simon JP, Haas D, Stalon V (1980) Catabolism of l-arginine by Pseudomonas aeruginosa. J Gen Microbiol 116:381–389
O'Hoy K, Krishnapillai V (1987) Recalibration of the Pseudomonas aeruginosa strain PAO chromosome map in time units using high-frequency of recombination donors. Genetics 115:611–618
Rahman M, Laverack PD, Clarke PH (1980) The catabolism of arginine by Pseudomonas aeruginosa. J Gen Microbiol 116:371–380
Reimmann C, Haas D (1986) IS21 insertion in the trfA replication control gene of chromosomally integrated plasmid RP1: a property of stable Pseudomonas aeruginosa Hfr strains. Molec Gen Genet 203:511–519
Soldati L, Leisinger T, Haas D (1982) Mapping of genes for proline and ornithine utilization in Pseudomonas aeruginosa. Experientia 38:1379
Stalon V, Ramos F, Piérard A, Wiame JW (1967) The occurrence of a catabolic and an anabolic ornithine carbamoyltransferase in Pseudomonas. Biochim Biophys Acta 139:91–97
Stalon V, Vander Wauven C, Momin P, Legrain C (1987) Catabolism of arginine, citrulline and ornithine by Pseudomonas and related bacteria. J Gen Microbiol 133:2487–2495
Tsuda M, Iino T (1983) Ordering of the flagellar genes in Pseudomonas aeruginosa by insertion of mercury transposon Tn501. J Bacteriol 153:1008–1017
Vander Wauven C, Stalon V (1985) Occurrence of succinyl derivatives in the catabolism of arginine in Pseudomonas cepacia. J Bacteriol 164:882–886
Vander Wauven C, Piérard A, Kley-Raymann M, Haas D (1984) Pseudomonas aeruginosa mutants affected in anaerobic growth on arginine: evidence for a four-gene cluster encoding the arginine deiminase pathway. J Bacteriol 160:928–934
Voellmy R, Leisinger T (1975) The dual role for acetylornithine 5-amino transferase from Pseudomonas aeruginosa in arginine biosynthesis and arginine catabolism. J Bacteriol 122:799–809
Voellmy R, Leisinger T (1976) Role of 4-aminobutyrate aminotransferase in the arginine metabolism of Pseudomonas aeruginosa. J Bacteriol 128:722–729
Voellmy R, Leisinger T (1978) Regulation of enzyme synthesis in the arginine metabolism of Pseudomonas aeruginosa. J Gen Microbiol 109:25–35
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Vander Wauven, C., Jann, A., Haas, D. et al. N2-Succinylornithine in ornithine catabolism of Pseudomonas aeruginosa . Arch. Microbiol. 150, 400–404 (1988). https://doi.org/10.1007/BF00408314
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00408314