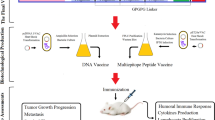
Abstract
Prostate cancer (PCa) is the second most diagnosed cancer in men and ranked fifth in overall cancer diagnosis. During the past decades, it has arisen as a significant life-threatening disease in men at an older age. At the early onset of illness when it is in localized form, radiation and surgical treatments are applied against this disease. In case of adverse situations androgen deprivation therapy, chemotherapy, hormonal therapy, etc. are widely used as a therapeutic element. However, studies found the occurrences of several side effects after applying these therapies. In current work, several immunoinformatic techniques were applied to formulate a multi-epitopic vaccine from the overexpressed antigenic proteins of PCa. A total of 13 epitopes were identified from the five prostatic antigenic proteins (PSA, PSMA, PSCA, STEAP, and PAP), after validation with several in silico tools. These epitopes were fused to form a vaccine element by (GGGGS)3 peptide linker. Afterward, 5, 6-dimethylxanthenone-4-acetic acid (DMXAA) was used as an adjuvant to initiate and induce STING-mediated cytotoxic cascade. In addition, molecular docking was performed between the vaccine element and HLA class I antigen with the low ACE value of −251 kcal/mol which showed a significant binding. Molecular simulation using normal mode analysis (NMA) illustrated the docking complex as a stable one. Therefore, this observation strongly indicated that our multi epitopes bases peptide vaccine molecule will be an effective candidate for the treatment of the PCa.










Similar content being viewed by others
References
Perera, M., Papa, N., Christidis, D., Wetherell, D., Hofman, M. S., & Murphy, D. G., et al. (2016). Sensitivity, specificity, and predictors of positive 68Ga–prostate-specific membrane antigen positron emission tomography in advanced prostate cancer: a systematic review and meta-analysis. European Urology, 70(6), 926–937.
Perera, M., Papa, N., Roberts, M., Williams, M., Udovicich, C., & Vela, I. et al. (2019). Gallium-68 prostate-specific membrane antigen positron emission tomography in advanced prostate cancer-updated diagnostic utility, sensitivity, specificity, and distribution of prostate-specific membrane antigen-avid lesions: a systematic review and meta-analysis. European Urology, 77(4), 403–417.
Siegel, R. L., Miller, K. D., & Jemal, A. (2019). Cancer statistics, 2019. CA: a Cancer Journal for Clinicians, 69(1), 7–34.
Bashir, M. N.(2015). Epidemiology of prostate cancer. Asian Pacific Journal of Cancer Prevention, 16(13), 5137–5141.
Walsh, P. C., DeWeese, T. L., & Eisenberger, M. A. (2007). Localized prostate cancer. New England Journal of Medicine, 357(26), 2696–2705.
Comiskey, M. C., Dallos, M. C., & Drake, C. G. (2018). Immunotherapy in prostate cancer: teaching an old dog new tricks. Current Oncology Reports, 20(9), 75.
Nguyen, P. L., Alibhai, S. M., Basaria, S., D’Amico, A. V., Kantoff, P. W., & Keating, N. L., et al. (2015). Adverse effects of androgen deprivation therapy and strategies to mitigate them. European Urology, 67(5), 825–836.
Ceder, Y., Bjartell, A., Culig, Z., Rubin, M. A., Tomlins, S., & Visakorpi, T. (2016). The molecular evolution of castration-resistant prostate cancer. European Urology Focus, 2(5), 506–513.
Sumanasuriya, S., & De Bono, J. (2018). Treatment of advanced prostate cancer—a review of current therapies and future promise. Cold Spring Harbor Perspectives In Medicine, 8(6), a030635.
Cheever, M. A., & Higano, C. S. (2011). PROVENGE (Sipuleucel-T) in prostate cancer: the first FDA-approved therapeutic cancer vaccine. Clinical Cancer Research, 17(11), 3520–3526.
Pardoll, D. M. (1998). Cancer vaccines. Nature Medicine, 4(5), 525.
Finn, O. J. (2003). Cancer vaccines: between the idea and the reality. Nature Reviews Immunology, 3(8), 630.
Doytchinova, I. A., & Flower, D. R. (2018). In silico prediction of cancer immunogens: current state of the art. BMC Immunology, 19(1), 11.
Ali, M., Pandey, R. K., Khatoon, N., Narula, A., Mishra, A., & Prajapati, V. K. (2017). Exploring dengue genome to construct a multi-epitope based subunit vaccine by utilizing immunoinformatics approach to battle against dengue infection. Scientific Reports, 7(1), 9232.
Stamey, T. A., Yang, N., Hay, A. R., McNeal, J. E., Freiha, F. S., & Redwine, E. (1987). Prostate-specific antigen as a serum marker for adenocarcinoma of the prostate. New England Journal of Medicine, 317(15), 909–916.
Chang, S. S., Reuter, V. E., Heston, W., Bander, N. H., Grauer, L. S., & Gaudin, P. B. (1999). Five different anti-prostate-specific membrane antigen (PSMA) antibodies confirm PSMA expression in tumor-associated neovasculature. Cancer Research, 59(13), 3192–3198.
Reiter, R. E., Gu Z, Watabe, T, Thomas, G, Szigeti, K, Davis, E, et al. (1998). Prostate stem cell antigen: a cell surface marker overexpressed in prostate cancer. Proceedings of the National Academy of Sciences USA, 95(4), 1735–1740.
de la Luz Garcia-Hernandez, M., Gray, A., Hubby, B., & Kast, W. M. (2007). In vivo effects of vaccination with six-transmembrane epithelial antigen of the prostate: a candidate antigen for treating prostate cancer. Cancer Research., 67(3), 1344–1351.
Rini, B. I., Weinberg, V., Fong, L., Conry, S., Hershberg, R. M., & Small, E. J. (2006). Combination immunotherapy with prostatic acid phosphatase pulsed antigen‐presenting cells (provenge) plus bevacizumab in patients with serologic progression of prostate cancer after definitive local therapy. Cancer, 107(1), 67–74.
Dannull, J., Diener, P.-A., Prikler, L., Fürstenberger, G., Cerny, T., & Schmid, U., et al. (2000). Prostate stem cell antigen is a promising candidate for immunotherapy of advanced prostate cancer. Cancer Research., 60(19), 5522–5528.
Matera, L. (2010). The choice of the antigen in the dendritic cell-based vaccine therapy for prostate cancer. Cancer Treatment Reviews, 36(2), 131–141.
Barh, D., Misra, A. N., Kumar, A., & Vasco, A. (2010). A novel strategy of epitope design in Neisseria gonorrhoeae. Bioinformation, 5(2), 77.
Su, C. T., Schönbach, C., & Kwoh, C.-K. (2013). Molecular docking analysis of 2009-H1N1 and 2004-H5N1 influenza virus HLA-B* 4405-restricted HA epitope candidates: implications for TCR cross-recognition and vaccine development. BMC Bioinformatics, 14, 1–7.
Consortium, U. (2018). UniProt: the universal protein knowledgebase. Nucleic Acids Research, 46(5), 2699.
Coordinators, N. R. (2018). Database resources of the national center for biotechnology information. Nucleic Acids Research, 46(Database issue), D8.
Potocnakova, L., Bhide, M., & Pulzova, L. B. (2016). An introduction to B-cell epitope mapping and in silico epitope prediction. Journal of Immunology Research, 6760830, 1–11.
Vita, R., Mahajan, S., Overton, J. A., Dhanda, S. K., Martini, S., & Cantrell, J. R., et al. (2018). The immune epitope database (IEDB): 2018 update. Nucleic Acids Research, 47(D1), D339–D343.
El-Manzalawy, Y., Dobbs, D., Honavar, V. (2008). Predicting flexible length linear B-cell epitopes. In: Computational systems bioinformatic, 7 (pp. 121–132). Stanford, CA: World Scientific.
El-Manzalawy, Y., Dobbs, D., & Honavar, V. (2008). Predicting linear B‐cell epitopes using string kernels. Journal of Molecular Recognition, 21(4), 243–255.
Chen, J., Liu, H., Yang, J., & Chou, K.-C. (2007). Prediction of linear B-cell epitopes using amino acid pair antigenicity scale. Amino Acids., 33(3), 423–428.
Jespersen, M. C., Peters, B., Nielsen, M., & Marcatili, P. (2017). BepiPred-2.0: improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Research, 45(W1), W24–W29.
Prickett, S., Rolland, J., & O’Hehir, R. (2015). Immunoregulatory T cell epitope peptides: the new frontier in allergy therapy. Clinical & Experimental Allergy, 45(6), 1015–1026.
Naz, A., Awan, F. M., Obaid, A., Muhammad, S. A., Paracha, R. Z., & Ahmad, J., et al. (2015). Identification of putative vaccine candidates against Helicobacter pylori exploiting exoproteome and secretome: a reverse vaccinology based approach. Infection Genetics and Evolution, 32, 280–291.
Patra P, Mondal N, Patra BC, Bhattacharya M. (2019) Epitope-based vaccine designing of nocardia asteroides targeting the virulence factor mce-family protein by immunoinformatics approach. International Journal of Peptide Research and Therapeutics, 1–12.
Saha, C. K., Hasan, M. M., Hossain, M. S., Jahan, M. A., & Azad, A. K. (2017). In silico identification and characterization of common epitope-based peptide vaccine for Nipah and Hendra viruses. Asian Pacific Journal of Tropical Medicine, 10(6), 529–538.
Singh, H., & Raghava, G. (2003). ProPred1: prediction of promiscuous MHC Class-I binding sites. Bioinformatics, 19(8), 1009–1014.
Singh, H., & Raghava, G. (2001). ProPred: prediction of HLA-DR binding sites. Bioinformatics, 17(12), 1236–1237.
Soria-Guerra, R. E., Nieto-Gomez, R., Govea-Alonso, D. O., & Rosales-Mendoza, S. (2015). An overview of bioinformatics tools for epitope prediction: implications on vaccine development. Journal of Biomedical Informatics, 53, 405–414.
Narula, A., Pandey, R. K., Khatoon, N., Mishra, A., & Prajapati, V. K. (2018). Excavating chikungunya genome to design B and T cell multi-epitope subunit vaccine using comprehensive immunoinformatics approach to control chikungunya infection. Infection Genetics and Evolution, 61, 4–15.
Horohov, D., Dunham, J., Liu, C., Betancourt, A., Stewart, J., & Page, A., et al. (2015). Characterization of the in situ immunological responses to vaccine adjuvants. Veterinary Immunology and Immunopathology, 164(1-2), 24–29.
Willard, L., Ranjan, A., Zhang, H., Monzavi, H., Boyko, R. F., & Sykes, B. D., et al. (2003). VADAR: a web server for quantitative evaluation of protein structure quality. Nucleic Acids Research, 31(13), 3316–3319.
Roy, A., Yang, J., & Zhang, Y. (2012). COFACTOR: an accurate comparative algorithm for structure-based protein function annotation. Nucleic Acids Research, 40(W1), W471–W7.
Peng, J., & Xu, J. (2011). RaptorX: exploiting structure information for protein alignment by statistical inference. Proteins: Structure, Function, and Bioinformatics, 79(S10), 161–171.
Khoury, G. A., Smadbeck, J., Kieslich, C. A., & Floudas, C. A. (2014). Protein folding and de novo protein design for biotechnological applications. Trends in Biotechnology, 32(2), 99–109.
Källberg, M, Margaryan, G, Wang, S, Ma, J, Xu, J. (2014). RaptorX server: a resource for template-based protein structure modeling. In: Protein structure prediction (pp. 17–27). New York, NY: Humana Press.
Webb, B, Sali, A. (2014). Protein structure modeling with MODELLER. Protein structure prediction (pp. 1–15). Springer.
Rodriguez, J., Parrilla, P., Moreno, A., Sola, J., Pinero, A., & Sa, Ortiz, et al. (1998). Insular carcinoma: an infrequent subtype of thyroid cancer. Journal of the American College of Surgeons, 187(5), 503–508.
Wiederstein, M., & Sippl, M. J. (2007). ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Research, 35(suppl_2), W407–W10.
Laskowski, R, MacArthur, M, Thornton, J. (2006). PROCHECK: validation of protein-structure coordinates.
Bhatt, D., Saxena, S. C., Jain, S., Dobriyal, A. K., Majee, M., & Arora, S. (2013). Cloning, expression and functional validation of drought inducible ascorbate peroxidase (Ec-apx1) from Eleusine coracana. Molecular Biology Reports, 40(2), 1155–1165.
Sippl, M. J. (1995). Knowledge-based potentials for proteins. Current Opinion in Structural Biology, 5(2), 229–235.
Laskowski, R. A., MacArthur, M. W., Moss, D. S., & Thornton, J. M. (1993). PROCHECK: a program to check the stereochemical quality of protein structures. Journal of Applied Crystallography, 26(2), 283–291.
Piplani, S., Saini, V., Niraj, R. R. K., Pushp, A., & Kumar, A. (2016). Homology modelling and molecular docking studies of human placental cadherin protein for its role in teratogenic effects of anti-epileptic drugs. Computational Biology and Chemistry, 60, 1–8.
Srivastava, P. N., Jain, R., Dubey, S. D., Bhatnagar, S., & Ahmad, N. (2016). Prediction of epitope-based peptides for vaccine development from coat proteins GP2 and VP24 of Ebola virus using immunoinformatics. International Journal of Peptide Research and Therapeutics, 22(1), 119–133.
Rahman, M. S., Rahman, M. K., Saha, S., Kaykobad, M., & Rahman, M. S. (2019). Antigenic: an improved prediction model of protective antigens. Artificial Intelligence in Medicine, 94, 28–41.
Doytchinova, I. A., & Flower, D. R. (2007). VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinformatics., 8(1), 4.
Bonakdar, A., Sahebazzamani, F., Rasaee, M. J., Hosseinkhani, S., Rahbarizadeh, F., & Mahboudi, F., et al. (2019). In silico design and in vitro characterization of a recombinant antigen for specific recognition of NMP22. International Journal of Biological Macromolecules, 140, 69–77.
Nezafat, N., Eslami, M., Negahdaripour, M., Rahbar, M. R., & Ghasemi, Y. (2017). Designing an efficient multi-epitope oral vaccine against Helicobacter pylori using immunoinformatics and structural vaccinology approaches. Molecular BioSystems., 13(4), 699–713.
Saha, S., & Raghava, G. (2006). AlgPred: prediction of allergenic proteins and mapping of IgE epitopes. Nucleic Acids Research, 34(suppl_2), W202–W9.
Khatoon, N., Ojha, R., Mishra, A., & Prajapati, V. K. (2018). Examination of antigenic proteins of Trypanosoma cruzi to fabricate an epitope-based subunit vaccine by exploiting epitope mapping mechanism. Vaccine., 36(42), 6290–6300.
Dimitrov, I., Bangov, I., Flower, D. R., & Doytchinova, I. (2014). AllerTOP v. 2—a server for in silico prediction of allergens. Journal of Molecular Modeling, 20(6), 2278.
Pagadala, N. S., Syed, K., & Tuszynski, J. (2017). Software for molecular docking: a review. Biophysical Reviews, 9(2), 91–102.
Schneidman-Duhovny, D., Inbar, Y., Nussinov, R., & Wolfson, H. J. (2005). PatchDock and SymmDock: servers for rigid and symmetric docking. Nucleic Acids Research, 33(suppl_2), W363–W7.
Yadav, S., Pandey, S. K., Singh, V. K., Goel, Y., Kumar, A., & Singh, S. M. (2017). Molecular docking studies of 3-bromopyruvate and its derivatives to metabolic regulatory enzymes: Implication in designing of novel anticancer therapeutic strategies. PLoS ONE., 12(5), e0176403.
Lankapalli, A. R., & Kannabiran, K. (2013). Interaction of marine Streptomyces compounds with selected cancer drug target proteins by in silico molecular docking studies. Interdisciplinary Sciences: Computational. Life Sciences, 5(1), 37–44.
Guo, F., Li, S. C., Wang, L., & Zhu, D. (2012). Protein-protein binding site identification by enumerating the configurations. BMC Bioinformatics, 13(1), 158.
Prabhakar, P. K., Srivastava, A., Rao, K. K., & Balaji, P. V. (2016). Monomerization alters the dynamics of the lid region in Campylobacter jejuni CstII: an MD simulation study. Journal of Biomolecular Structure and Dynamics, 34(4), 778–791.
López-Blanco, J. R., Aliaga, J. I., Quintana-Ortí, E. S., & Chacón, P. (2014). iMODS: internal coordinates normal mode analysis server. Nucleic Acids Research, 42(W1), W271–W276.
Hasan, M., Ghosh, P. P., Azim, K. F., Mukta, S., Abir, R. A., & Nahar, J., et al. (2019). Reverse vaccinology approach to design a novel multi-epitope subunit vaccine against avian influenza A (H7N9) virus. Microbial Pathogenesis., 130, 19–37. https://doi.org/10.1016/j.micpath.2019.02.023.
Grote, A., Hiller, K., Scheer, M., Münch, R., Nörtemann, B., & Hempel, D. C., et al. (2005). JCat: a novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Research, 33(suppl_2), W526–W531.
Pandey, R. K., & Prajapati, V. K. (2019). Exploring sand fly salivary proteins to design multiepitope subunit vaccine to fight against visceral leishmaniasis. Journal of Cellular Biochemistry, 120(2), 1141–1155.
Khatoon, N., Pandey, R. K., & Prajapati, V. K. (2017). Exploring Leishmania secretory proteins to design B and T cell multi-epitope subunit vaccine using immunoinformatics approach. Scientific Reports, 7(1), 8285.
Kamens, J. (2014). The Addgene repository: an international nonprofit plasmid and data resource. Nucleic Acids Research, 43(D1), D1152–D1157.
Sanchez-Trincado, J. L., Gomez-Perosanz, M., & Reche, P. A. (2017). Fundamentals and methods for T-and B-cell epitope prediction. Journal of Immunology Research, 2680160, 1–14.
Chen, X., Zaro, J. L., & Shen, W.-C. (2013). Fusion protein linkers: property, design and functionality. Advanced Drug Delivery Reviews, 65(10), 1357–1369.
Prantner, D., Perkins, D. J., Lai, W., Williams, M. S., Sharma, S., & Fitzgerald, K. A., et al. (2012). 5, 6-Dimethylxanthenone-4-acetic acid (DMXAA) activates stimulator of interferon gene (STING)-dependent innate immune pathways and is regulated by mitochondrial membrane potential. Journal of Biological Chemistry, 287(47), 39776–39788.
Park, J.-H., Ko, S.-M., & Park, H.-S. (2008). Toward the virtual screening of α-glucosidase inhibitors with the homology-modeled protein structure. Bulletin of the Korean Chemical Society, 29(5), 921–927.
Bianchi, V., Bulek, A., Fuller, A., Lloyd, A., Attaf, M., & Rizkallah, P. J., et al. (2016). A molecular switch abrogates glycoprotein 100 (gp100) T-cell receptor (TCR) targeting of a human melanoma antigen. The Journal of Biological Chemistry, 291(17), 8951–8959. https://doi.org/10.1074/jbc.M115.707414.
Patra, P, Ghosh, P, Patra, B. C., Bhattacharya, M. (2019) Biocomputational analysis and in silico characterization of an angiogenic protein (RNase5) in Zebrafish (Danio rerio). International Journal of Peptide Research and Therapeutics, 1–11.
Schrodinger, L (2010). The PyMOL molecular graphics system. Version 1.5.0.4.
Bauer, J. A., Pavlović, J., & Bauerová-Hlinková, V. (2019). Normal mode analysis as a routine part of a structural investigation. Molecules., 24(18), 3293.
Atapour, A., Negahdaripour, M., Ghasemi, Y., Razmjuee, D., Savardashtaki, A., & Mousavi, S. M. et al. (2019). In silico designing a candidate vaccine against breast cancer. International Journal of Peptide Research and Therapeutics, 26, 369–380.
Pollock, P., Ludgate, A., & Wassersug, R. (2015). In 2124, half of all men can count on developing prostate cancer. Current Oncology, 22(1), 10.
Pilleron, S., Sarfati, D., Janssen‐Heijnen, M., Vignat, J., Ferlay, J., & Bray, F., et al. (2019). Global cancer incidence in older adults, 2012 and 2035: a population‐based study. International Journal of Cancer, 144(1), 49–58.
Carlsson, S., Assel, M., Ulmert, D., Gerdtsson, A., Hugosson, J., & Vickers, A., et al. (2017). Screening for prostate cancer starting at age 50–54 years. A population-based cohort study. European Urology, 71(1), 46–52.
Wijk, M. V., Garland, S. N., Wall, K., Sathya, J., & Thoms, J. (2019). 0825 The effect of androgen deprivation therapy on insomnia symptoms, fatigue, mood, and hot flashes in men with non-metastatic prostate cancer. Sleep., 42(Supplement_1), A331–A.
Mayer, M. J., Klotz, L. H., & Venkateswaran, V. (2017). The effect of metformin use during docetaxel chemotherapy on prostate cancer specific and overall survival of diabetic patients with castration resistant prostate cancer. The Journal of Urology, 197(4), 1068–1075.
Poeppel, T. D., Handkiewicz-Junak, D., Andreeff, M., Becherer, A., Bockisch, A., & Fricke, E., et al. (2018). EANM guideline for radionuclide therapy with radium-223 of metastatic castration-resistant prostate cancer. European Journal of Nuclear Medicine and Molecular Imaging, 45(5), 824–845.
Higano, C. S., Armstrong, A. J., Sartor, A. O., Vogelzang, N. J., Kantoff PW, McLeod DG et al. (2019). Real‐world outcomes of sipuleucel‐T treatment in PROCEED, a prospective registry of men with metastatic castration‐resistant prostate cancer. Cancer.
Namvar, A., Panahi, H. A., Agi, E., & Bolhassani, A. (2020). Development of HPV 16, 18, 31, 45 E5 and E7 peptides-based vaccines predicted by immunoinformatics tools. Biotechnology Letters., 8, 1–6.
Bhattacharya, M., Malick, R. C., Mondal, N., Patra, P., Pal, B. B., & Patra, B. C., et al. (2020). Computational characterization of epitopic region within the outer membrane protein candidate in Flavobacterium columnare for vaccine development. Journal of Biomolecular Structure and Dynamics, 38(2), 450–459.
Vishnu, U. S., Sankarasubramanian, J., Gunasekaran, P., & Rajendhran, J. (2017). Identification of potential antigens from non-classically secreted proteins and designing novel multitope peptide vaccine candidate against Brucella melitensis through reverse vaccinology and immunoinformatics approach. Infection, Genetics and Evolution, 55, 151–158.
Lockhat, H. A., Silva, J. R., Alves, C. N., Govender, T., Lameira, J., & Maguire, G. E., et al. (2016). Binding free energy calculations of nine FDA‐approved protease inhibitors against HIV‐1 subtype C I36T↑ T containing 100 amino acids per monomer. Chemical Biology & Drug Design, 87(4), 487–498.
Camacho, C. J., & Vajda, S. (2002). Protein–protein association kinetics and protein docking. Current Opinion in Structural Biology, 12(1), 36–40.
Sayed, S. B., Nain, Z., Abdullah, F., Haque, Z., Rahman, S. R., Tasmin, R., et al. (2019). Immunoinformatics-guided designing of peptide vaccine against Lassa virus with dynamic and immune simulation studies. Preprints 2019, 2019090076 https://doi.org/10.20944/preprints201909.0076.v2.
Acknowledgements
This research was supported by Hallym University Research Fund and by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2017R1A2B4012944).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary material
Rights and permissions
About this article
Cite this article
Patra, P., Bhattacharya, M., Sharma, A.R. et al. Identification and Design of a Next-Generation Multi Epitopes Bases Peptide Vaccine Candidate Against Prostate Cancer: An In Silico Approach. Cell Biochem Biophys 78, 495–509 (2020). https://doi.org/10.1007/s12013-020-00912-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-020-00912-7