Abstract
The Chinese equine infectious anemia virus (EIAV) virulent strain EIAVLN40 is derived from a naturally occurring virus by continuously passing in horses for 16 generations. Its genome sequence is 23% different from that of the American strains or the Japanese strains, and the variation of envelope gp90 surface unit (SU) is as high as 41%. In this study, evolutions of the EIAVLN40 gp90 gene in four infected horses were analyzed. Results showed that new quasispecies arose in the early stage of infection in all EIAVLN40-infected horses. These quasispecies belonged to branches different from EIAVLN40 in a phylogenetic tree. In contrast, the gp90 sequences of viruses isolated after disease onset remained in the same phylogenetic branch as EIAVLN40, with some having exactly the same sequences. The glycosylation sites 191NSSN and 237NNTW in the V3 and V4 region present or absent simultaneously in most of the predicted amino acid sequences. Changes in the glycosylation sites within V3, V4, and V5 regions are usually associated with the disease status. Glycosylation sites (191NSSN, 237NNTW, and 280NDTS) within these three regions were present in EIAVLN40 and most of the quasispecies isolated after, but not before disease onset. These unique evolutionary characteristics of SU have not been reported for EIAV and other lentiviruses. Our results provide a reference for a further understanding of the mechanism underlying the persistent infection and escape from immune surveillance of EIAV.




Similar content being viewed by others
References
C. Leroux, J.K. Craigo, C.J. Issel, R.C. Montelaro, J. Virol. 75, 4570–4583 (2001)
S.J. More, I. Aznar, D.C. Bailey, J.F. Larkin, D.P. Leadon, P. Lenihan, B. Flaherty, U. Fogarty, P. Brangan, Equine Vet. J. 40, 706–708 (2008)
A. Kaiser, H.P. Meier, R. Straub, V. Gerber, Schweiz. Arch. Tierheilkd. 151, 159–164 (2009)
C. Leroux, J.L. Cadore, R.C. Montelaro, Vet. Res. 35, 485–512 (2004)
C.J. Issel, W.V. Adams Jr., L. Meek, R. Ochoa, J. Am. Vet. Med. Assoc. 180, 272–275 (1982)
J.K. Craigo, C. Leroux, L. Howe, J.D. Steckbeck, S.J. Cook, C.J. Issel, R.C. Montelaro, J. Gen. Virol. 83, 1353–1359 (2002)
M.M. Nagarajan, C. Simard, Virus Res. 129, 228–235 (2007)
J.K. Craigo, B. Zhang, S. Barnes, T.L. Tagmyer, S.J. Cook, C.J. Issel, R.C. Montelaro, Proc. Natl. Acad. Sci. USA 104, 15105–15110 (2007)
T.L. Tagmyer, J.K. Craigo, S.J. Cook, D.L. Even, C.J. Issel, R.C. Montelaro, J. Virol. 82, 4052–4063 (2008)
C. Leroux, C.J. Issel, R.C. Montelaro, J. Virol. 71, 9627–9639 (1997)
J.M. Ball, K.E. Rushlow, C.J. Issel, R.C. Montelaro, J. Virol. 66, 732–742 (1992)
T.L. Tagmyer, J.K. Craigo, S.J. Cook, C.J. Issel, R.C. Montelaro, J. Gen. Virol. 88, 1324–1336 (2007)
L. Howe, C. Leroux, C.J. Issel, R.C. Montelaro, J. Virol. 76, 10588–10597 (2002)
S. Yoon, S.M. Kingsman, A.J. Kingsman, S.A. Wilson, K.A. Mitrophanous, J. Gen. Virol. 81, 2189–2194 (2000)
S.L. Payne, J. Rausch, K. Rushlow, R.C. Montelaro, C. Issel, M. Flaherty, S. Perry, D. Sellon, F. Fuller, J. Gen. Virol. 75(Pt 2), 425–429 (1994)
L. Martarano, R. Stephens, N. Rice, D. Derse, J. Virol. 68, 3102–3111 (1994)
W. Maury, P.J. Wright, S. J. Virol., Bradley 77, 2385–2399 (2003)
D.L. Lichtenstein, C.J. Issel, R.C. Montelaro, J. Virol. 70, 3346–3354 (1996)
B.A. Sponseller, W.O. Sparks, Y. Wannemuehler, Y. Li, A.K. Antons, J.L. Oaks, S. Carpenter, Virology 363, 156–165 (2007)
M. Belshan, P. Baccam, J.L. Oaks, B.A. Sponseller, S.C. Murphy, J. Cornette, S. Carpenter, Virology 279, 185–200 (2001)
S. Jin, C. Chen, R.C. Montelaro, J. Virol. 79, 8793–8801 (2005)
Y.H. Zheng, T. Nakaya, H. Sentsui, M. Kameoka, M. Kishi, K. Hagiwara, H. Takahashi, Y. Kono, K. Ikuta, J. Gen. Virol. 78(Pt 4), 807–820 (1997)
Y.H. Zheng, H. Sentsui, T. Nakaya, Y. Kono, K. Ikuta, J. Virol. 71, 5031–5039 (1997)
Y.H. Zheng, H. Sentsui, Y. Kono, K. Ikuta, Virus Res. 68, 93–98 (2000)
Y.H. Zheng, H. Sentsui, M. Sugita, T. Nakaya, M. Kishi, K. Hagiwara, Y. Inoshima, C. Ishihara, Y. Kono, J.L. Lu, K. Ikuta, Virology 266, 129–139 (2000)
L. Wei, X. Fan, X. Lu, L. Zhao, W. Xiang, X. Zhang, F. Xue, Y. Shao, R. Shen, X. Wang, Virus Genes 38, 285–288 (2009)
X.Z. Cao, Y.Z. Lin, L. Li, C.G. Jiang, L.P. Zhao, X.L. Lv, J.H. Zhou, Bing Du Xue Bao 26, 128–133 (2010)
X. Gao, C.G. Jiang, X.E. Han, L.P. Zhao, R.X. Shen, W.H. Xiang, J.H. Zhou, Bing Du Xue Bao 25, 309–315 (2009)
H. Liang, X. He, R.X. Shen, T. Shen, X. Tong, Y. Ma, W.H. Xiang, X.Y. Zhang, Y.M. Shao, Arch. Virol. 151, 1387–1403 (2006)
J. Ma, C. Jiang, Y. Lin, X. Wang, L. Zhao, W. Xiang, Y. Shao, R. Shen, X. Kong, J. Zhou, Arch. Virol. 154, 867–873 (2009)
T. Shen, H. Liang, X. Tong, X. Fan, X. He, Y. Ma, W. Xiang, R. Shen, X. Zhang, Y. Shao, Vaccine 24, 738–749 (2006)
Y.B. Tu, T. Zhou, X.F. Yuan, H.J. Qiu, F. Xue, C.Q. Sun, L. Wang, D.L. Wu, J.M. Peng, X.G. Kong, G.Z. Tong, Arch. Virol. 152, 209–218 (2007)
X.F. Wang, C.G. Jiang, W. Guo, W. Xiang, X.L. Lv, L.P. Zhao, F.L. Wang, X.G. Kong, X.Y. Zhang, Y.M. Shao, J.H. Zhou, Bing Du Xue Bao 24, 443–450 (2008)
J.K. Craigo, S. Barnes, B. Zhang, S.J. Cook, L. Howe, C.J. Issel, R.C. Montelaro, Retrovirology 6, 95 (2009)
X. Qi, X. Wang, S. Wang, Y. Lin, C. Jiang, J. Ma, L. Zhao, X. Lv, R. Shen, F. Wang, X. Kong, Z. Su, J. Zhou, Virus Genes 41, 86–98 (2010)
J. Ma, N. Shi, C. Jiang, Y. Lin, X. Wang, S. Wang, X. Lv, L. Zhao, Y. Shao, X. Kong, J. Zhou, R. Shen. Virology (2011). doi:10.1016/j.virol.2010.10.032
D.L. Suarez, C.A. Whetstone, Virology 212, 728–733 (1995)
S.P. McBurney, T.M. Ross, Expert. Rev. Vaccines 7, 1405–1417 (2008)
K.J. Metzner, P. Rauch, H. Walter, C. Boesecke, B. Zollner, H. Jessen, K. Schewe, S. Fenske, H. Gellermann, H.J. Stellbrink, Aids 19, 1819–1825 (2005)
J.K. Craigo, T.J. Sturgeon, S.J. Cook, C.J. Issel, C. Leroux, R.C. Montelaro, Virology 344, 340–353 (2006)
J.K. Craigo, R.C. Montelaro, Curr. HIV Res. 8, 81–86 (2010)
L.Q. Zhang, P. MacKenzie, A. Cleland, E.C. Holmes, A.J. Brown, P. Simmonds, J. Virol. 67, 3345–3356 (1993)
T. Zhu, H. Mo, N. Wang, D.S. Nam, Y. Cao, R.A. Koup, D.D. Ho, Science 261, 1179–1181 (1993)
C.L. Kuiken, J.J. de Jong, E. Baan, W. Keulen, M. Tersmette, J. Goudsmit, J. Virol. 66, 4622–4627 (1992)
S.S. Hwang, T.J. Boyle, H.K. Lyerly, B.R. Cullen, Science 253, 71–74 (1991)
H. Li, X. Zhang, X. Fan, T. Shen, X. Tong, R. Shen, Y. Shao, AIDS Res. Hum. Retroviruses 21, 1057–1059 (2005)
T.C. McGuire, T.B. Crawford, J.B. Henson, Am. J. Pathol. 62, 283–294 (1971)
N.R. Rice, A.S. Lequarre, J.W. Casey, S. Lahn, R.M. Stephens, J. Edwards, J. Virol. 63, 5194–5200 (1989)
D.C. Sellon, S.T. Perry, L. Coggins, F.J. Fuller, J. Virol. 66, 5906–5913 (1992)
C.K. Leonard, M.W. Spellman, L. Riddle, R.J. Harris, J.N. Thomas, T.J. Gregory, J. Biol. Chem. 265, 10373–10382 (1990)
D.J. Vigerust, V.L. Shepherd, Trends Microbiol. 15, 211–218 (2007)
P. Edmonson, M. Murphey-Corb, L.N. Martin, C. Delahunty, J. Heeney, H. Kornfeld, P.R. Donahue, G.H. Learn, L. Hood, J.I. Mullins, J. Virol. 72, 405–414 (1998)
Acknowledgments
This study was funded by National Natural Science Foundation of China (30771994 to JZ and 30901349 to YL) and National Key Programs for Infectious Diseases (2008ZX1001-1010 to JZ).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
11262_2010_563_MOESM1_ESM.doc
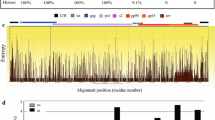
Fig. S1. Comparison of deduced amino acid sequences of SU from EIAVLN40 stock and circulating virions of EIAVLN40-infected horses (25#, 26#, 4#, and 10#). The italic, red numbers on the left indicate the consensus SU sequence of each sample. Identifications of each sequence are listed on the left. Dots indicate residues identical to the consensus sequence of EIAVLN40, and dashes indicate amino acid deletions. Underlined residues indicate potential N-glycosylation sites (NXS/T). Based on the amino acid sequence alignment of the SU sequences, eight SU variable regions (V1 to V8, indicated by boxed residues) were defined as stretches of amino acids with at least 30% divergence from the consensus sequence of the all sequences. Only the same mutations present in at least two of the analyzed clones were considered to delineate the variable regions. (DOC 354 kb)
Rights and permissions
About this article
Cite this article
Wang, X., Wang, S., Lin, Y. et al. Unique evolution characteristics of the envelope protein of EIAVLN40, a virulent strain of equine infectious anemia virus. Virus Genes 42, 220–228 (2011). https://doi.org/10.1007/s11262-010-0563-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-010-0563-7