Abstract
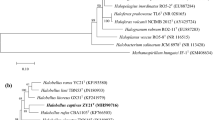
A novel halophilic archaeon, strain MBLA0160T, was isolated from a solar saltern in Sorae, Republic of Korea. The cells are deep-red pigmented, Gram-negative, rod shaped, motile, and lysed in distilled water. The strain MBLA0160T grew at 25–45 °C (optimum 37 °C), in 15–30% (w/v) NaCl (optimum 20%) and 0.1–1.0 M MgCl2 (optimum 0.3–0.5 M) at pH 5.0–9.0 (optimum 7.0). Phylogenetic analysis based on the 16S rRNA sequence showed that this strain was related to two species within the genus Halobellus (Hbs.), with 98.4% and 95.8% similarity to Hbs. salinus CSW2.24.4 T and Hbs. clavatus TNN18T, respectively. The major polar lipids of the strain MBLA160T were phosphatidylglycerol, phosphatidylglycerol sulfate, and phosphatidylglycerol phosphate methyl ester. The genome size, G + C content, and N50 value of MBLA0160T were 3.49 Mb, 66.5 mol%, and 620,127 bp, respectively. According to predicted functional proteins of strain MBLA0160T, the highest category was amino acid transport and metabolism. Genome rapid annotation showed that amino acid and derivatives was the most subsystem feature counts. Pan-genomic analysis showed that strain MBLA0160T had 97 annotated unique KEGG, which were mainly included metabolism and environmental information processing. Ortholog average nucleotide identities (OrthoANI) and in silico DNA-DNA hybridization (isDDH) values between the strain MBLA0160T and other strains of the genus Halobellus were under 84,4% and 28.1%, respectively. The genome of strain MBLA0160T also contain the biosynthetic gene cluster for C50 carotenoid as secondary metabolite. Based on the phylogenetic, phenotypic, chemotaxonomic properties, and comparative genomic analyses, strain MBLA0160T is considered to represent a novel species of the genus Halobellus, for which the name Halobellus ruber sp. nov. is proposed. The type strain is MBLA0160T (= KCTC 4291 T = JCM 34172 T).




Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST Server: rapid annotations using subsystems technology. BMC Genom 9:75
Bardavid RE, Mana L, Oren A (2007) Haloplanus natans gen. nov., sp. nov., an extremely halophilic, gas-vacuolate archaeon isolated from Dead Sea-Red Sea water mixtures in experimental outdoor ponds. Int J Syst Evol Microbiol 57:780–783
Bauer AW, Kirby MM, Sherris JC, Truck M (1966) Antibiotic susceptibility testing by a standardized single disk method. Am J Clin Pathol 45:493–496
Blin K, Shaw S, Steinke K, Villebro R, Ziemert N, Lee SY, Medema MH, Weber T (2019) antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucl Acids Res 47:W81–W87
Borowitzka MA (2010) Carotenoid production using microorganisms. In: Cohen Z, Ratledge C (eds) Single cell oils: microbial and algal oils. AOCS Press, Urbana, pp 225–240
Brettin T, Davis JJ, Disz T, Edwards RA, Gerdes S, Olsen GJ, Olson R, Overbeek R, Parrello B, Pusch GD, Shukla M, Thomason JA III, Stevens R, Vonstein V, Wattam AR, Xia F (2015) RASTtk: a modular and extensible implementation of the RAST algorithm for building custom annotation pipelines and annotating batches of genomes. Sci Rep 5:8365
Burns DG, Janssen PH, Itoh T, Kamekura M, Li Z, Jensen G, Rodriguez-Valera F, Bolhuis H, Dyall-Smith ML (2007) Haloquadratum walsbyi gen. nov., sp. nov., the square haloarchaeon of Walsby, isolated from saltern crystallizers in Australia and Spain. Int J Syst Evol Microbiol 57:387–392
Calegari-Santos R, Diogo RA, Fontana JD, Bordin Bonfim TM (2016) Carotenoid production by halophilic archaea under different culture conditions. Curr Microbiol 72:641–651
Cha IT, Yim KJ, Song HS, Lee HW, Hyun DW, Kim KN, Seo MJ, Kim D, Choi JS, Lee SJ, Bae JW, Rhee SK, Choi HJ, Rhee JK, Nam YD, Roh SW (2014) Halobellus rufus sp. nov., an extremely halophilic archaeon isolated from non-purified solar salt. Antonie Van Leeuwenhoek 105:925–932
Chaudhari NM, Gupta VK, Dutta C (2016) BPGA- an ultra-fast pan-genome analysis pipeline. Sci Rep 6:24373
Chen S, Liu HC, Zhou J, Xiang H (2016) Haloparvum sediment gen. nov., sp. nov., a member of the family Haloferacaceae. Int J Syst Evol Microbiol 66:2327–2334
Chen S, Suns S, Xu Y, Chen F, Liu J (2020) Halobellus captivus sp. nov., an extremely halophilic archaeon isolated from a subterranean salt mine. Antonie Van Leeuwenhoek 113:221–231
Chun J, Oren A, Ventosa A, Christensen H, Arahal DR, da Costa MS, Rooney AP, Yi H, Xu XW, De Meyer S, Trujillo ME (2018) Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int J Syst Evol Microbiol 68:461–466
Cortés-Albayay C, Jarmusch SA, Trusch F, Ebel R, Andrews BA, Jaspars M, Asenjo JA (2020) Downsizing class II lasso peptides: genome mining-guided isolation of huascopeptin containing the first Gly1–Asp7 macrocycle. J Org Chem 85:1661–1667
Cui HL, Zhou PJ, Oren A, Liu SJ (2009) Intraspecific polymorphism of 16S rRNA genes in two halophilic archaeal genera, Haloarcula and Halomicrobium. Extremophiles 13:31–37
Cui HL, Gao X, Sun FF, Dong Y, Xu XW, Zhou YG, Liu HC, Oren A, Zhou PJ (2010a) Halogranum rubrum gen. nov., sp. nov., a halophilic archaeaon isolated from a marine solar saltern. Int J Syst Evol Microbiol 60:1366–1371
Cui HL, Li XY, Gao X, Xu XW, Zhou YG, Liu HC, Oren A, Zhou PJ (2010b) Halopelagius inordinatus gen. nov., sp. nov., a new member of the family Halobacteriaceae isolated from a marine solar saltern. Int J Syst Evol Microbiol 60:2089–2093
Cui HL, Yang X, Gao X, Xu XW (2011) Halobellus clavatus gen. nov., sp. nov. and Halorientalis regularis gen. nov., sp. nov., two new members of the family Halobacteriaceae. Int J Syst Evol Microbiol 61:2682–2689
Cui HL, Yang X, Zhou YG, Liu HC, Zhou PJ, Dyall-Smith ML (2012) Halobellus limi sp. nov. and Halobellus salinus sp. nov., isolated from two marine solar salterns. Int J Syst Evol Microbiol 62:1307–1313
David LW, Deanna MC, Ron E, Scott F, Wolfgang H, Thomas LM, Joan UP, Gregory DS, Lynn MS, Edwin S, Tugba OS, Tatiana AT, Lukas W (2004) Database resources of the National Center for Biotechnology Information: update. Nucleic Acids Res 32:35–40
Deole R, Hoff WD (2020) A potassium chloride to glycine betaine osmoprotectant switch in the extreme halophile Halorhodospira halophila. Sci Rep 10:1–10
Dussault HP (1955) An improved technique for staining red halophilic bacteria. J Bacteriol 70:484–485
Fathepure BZ (2014) Recent studies in microbial degradation of petroleum hydrocarbons in hypersaline environments. Front Microbiol 5:173
Flores N, Hoyos S, Venegas M, Galetović A, Zúñiga LM, Fábrega F, Paredes B, Vilo Salazar-Ardiles C, C, Ascaso C, (2020) Haloterrigena sp. Strain SGH1, a Bacterioruberin-Rich, Perchlorate-Tolerant Halophilic Archaeon Isolated From Halite Microbial Communities, Atacama Desert. Chile Front Microbiol. 11:324
Gonzalez C, Gutierrez C, Ramirez C (1978) Halobacterium vallismortis sp. nov. An amylolytic and carbohydrate-metabolizing, extremely halophilic bacterium. Can J Microbiol 24:710–715
Gupta RS, Naushad S, Baker S (2015) Phylogenomic analyses and molecular signatures for the class Halobacteria and its two major clades: a proposal for division of the class Halobacteria into an emended order Halobacteriales and two new orders, Haloferacales ord. nov. and Natrialbales ord. nov., containing the novel families Haloferacaceae fam. nov. and Natrialbaceae fam. nov. Int J Syst Evol Microbiol 65:1050–1069
Gutiérrez C, González C (1972) Method for simultaneous detection of proteinase and esterase activities in extremely halophilic bacteria. Appl Microbiol 24:516–517
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Holding A, Collee J (1971) Chapter I Routine biochemical tests. Methods Microbiol 6:1–32
Huerta-Cepas J, Forslund K, Coelho LP, Szklarczyk D, Jensen LJ, vonMering C, Bork P (2017) Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol Biol Evol 34:2115–2122
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16(2):111–120
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Lee I, Kim YO, Park SC, Chun J (2016) OrthoANI: an improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Lee I, Chalita M, Ha SM, Na SI, Yoon SH, Chun J (2017) ContEst16S: analgorithm that identifies contaminated prokaryotic genomes using 16S RNAgene sequences. Int J Syst Evol Microbiol 67:2053–2057
Lefort V, Desper R, Gascuel O (2015) FastME 2.0: a comprehensive, accurate, and fast distance-based phylogeny inference program. Mol Biol Evol 32:2798–2800
Liu BB, Narsing Rao MP, Yin XQ, Li X, Salam N, Zhang Y, Alkhalifah DHM, Hozzein WN, Li WJ (2019) Description of Halegenticoccus soli gen. nov., sp. nov., a halophilic archaeon isolated from a soil sample of Ebi lake. Extremophiles 23:521–528
Maksimov MO, Pan SJ, Link AJ (2012) Lasso peptides: structure, function, biosynthesis, and engineering. Nat Prod Rep 29:996–1006
Meier-Kolthoff JP, Göker M (2019) TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat Commun 10:2182
Meier-Kolthoff JP, Auch AF, Klenk HP, Goker M (2013) Genome sequence-based species delimitation with confidence intervals and improved distance functions. BMC Bioinform 14:60
Minnikin DE, O’Donnell AG, Goodfellow M, Alderson G, Athalye M, Schaal A, Parlett JH (1984) An integrated procedure for the extraction of bacterial isoprenoid quinones and polar lipids. J Microbiol Methods 2:233–241
Montalvo-Rodriguez R, Vreeland RH, Oren A, Kessel M, Betancourt C, Lopez-Garriga J (1998) Halogeometricum borinquense gen. nov., sp. nov., a novel halophilic archaeon from Puerto Rico. Int J Syst Bacteriol 48:1305–1312
Oren A (2002) Halophilic Microorganisms and their Environments. Kluwer, London, UK, pp 233–278
Oren A, Ventosa A, Grant WD (1997) Proposed minimal standards for description of new taxa in the order Halobacteriales. Int J Syst Bacteriol 47:233–238
Pérez-Davó A, Aguilera M, González-Paredes A, Luján Jiménez-Pranteda M, Monteoliva-Sánchez M (2015) Halobellus ramosii sp. nov., an extremely halophilic archaeon isolated from a saline-wetland wildfowl reserve. Int J Syst Evol Microbiol 65:3847–3852
Qiu XX, Mou YZ, Zhao ML, Zhang WJ, Han D, Ren M, Cui HL (2013) Halobellus inordinatus sp. nov., from a marine solar saltern and an inland salt lake of China. Int J Syst Evol Microbiol 63:3975–3980
Rebuffat S, Blond A, Destoumieux-Garzon D, Goulard C, Peduzzi J (2004) Microcin J25, from the macrocyclic to the lasso structure: implications for biosynthetic, evolutionary and biotechnological perspectives. Curr Protein Pept Sci 5:383–391
Roh SW, Sung Y, Nam YD, Chang HW, Kim KH, Yoon JH, Jeon CO, Oh HM, Bae JW (2008) Arthrobacter soli sp. nov., a novel bacterium isolated from wastewater reservoir sediment. J Microbiol 46:40–44
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Bio Evol 4:406–425
Savage KN, Krumholz LR, Oren A, Elshahed MS (2008) Halosarcina pallida gen. nov., sp. nov., a halophilic archaeon from a low-salt, sulphide-rich spring. Int J Syst Evol Microbiol 58:856–860
Scalise M, Indiveri C (2020) Amino acids Transport and Metabolism 2.0. Int J Mol Sci 21:1212
Silver S, Walderhaung M (1992) Gene regulation of plasmid and chromosome determined inorganic ion transport in bacteria. Microbiol Rev 56:195–228
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P (ed) Methods for general and molecular bacteriology. American Society for Microbiology, Washington DC, pp 607–654
Stock AM, Robinson VL, Goudreau PN (2000) Two-component signal transduction. Annu Rev Biochem 69:183–215
Sullivan MJ, Petty NK, Beatson SA (2011) Easyfig: a genome comparison visualizer. Bioinformatics 27:1009–1010
Tatusov RL, Galperin MY, Natale DA, Koonin EV (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28:33–36
Thombre RS, Shinde VD, Oke RS, Dhar SK, Shouche YS (2016) Biology and survival of extremely halophilic archaeon Haloarcula marismortui RR12 isolated from Mumbai salterns, India in response to salinity stress. Sci Rep 6:25642
Tittsler RP, Sandholzer LA (1936) The use of semi-solid agar for the detection of bacterial motility. J Bacteriol 31:575–580
Tobias AV, Arnold FH (2006) Biosynthesis of novel carotenoid families based on unnatural carbon backbones: a model for diversification of natural product pathways. Biochim Biophys Acta 1761:235–246
Torreblanca M, Rodriguez-Valera F, Juez G, Ventosa A, Kamekura M, Kates M (1986) Classification of non-alkaliphilic halobacteria based on numerical taxonomy and polar lipid composition, and description of Haloarcula gen. nov. and Haloferax gen. nov. Syst Appl Microbiol 8:89–99
Yang Y, Yatsunami R, Ando A, Miyoko N, Fukui T, Takaichi S, Nakamura S (2015) Complete biosynthetic pathway of the C50 carotenoid bacterioruberin from lycopene in the extremely halophilic archaeon Haloarcula japonica. J Bacteriol 197:1614–1623
Yatsunami R, Ando A, Yang Y (2014) Identification of carotenoids from the extremely halophilic archaeon Haloarcula japonica. Front Microbiol 5:100
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA gene sequences and whole-genome assemblies. Int J Syst Evol Microbiol 67:1613–1617
Zalazar L, Pagola P, Miró MV, Churio MS, Cerletti M, Martínez C, Iniesta-Cuerda M, Soler AJ, Cesari A, De Castro R (2019) Bacterioruberin extracts from a genetically modified hyperpigmented Haloferax volcanii strain: antioxidant activity and bioactive properties on sperm cells. J Appl Microbiol 126:796–810
Zhang WJ, Han D, Qiu XX, Zhao ML, Mou YZ, Cui HL, Li ZR (2013) Halobellus rarus sp. nov., a halophilic archaeon from an inland salt lake of China. Antonie Van Leeuwenhoek 104:377–384
Zhang G, Gu J, Zhang R, Rashid M, Haroon MF, Xun W, Ruan Z, Dong X, Stingl U (2017) Haloprofundus marisrubri gen. nov., sp. nov., an extremely halophilic archaeon isolated from a brine-seawater interface. Int J Syst Evol Microbiol 67:9–16
Zhao ML, Qiu XX, Zhang WJ, Han D, Cui HL, Li ZR (2014) Halobellus litoreus sp. nov., a halophilic archaeon isolated from a Chinese marine solar saltern. Curr Microbiol 68:156–160
Funding
This work was supported by an Incheon National University Research Grant in 2018.
Author information
Authors and Affiliations
Contributions
C Y H and E-S C performed all the experiments and drafted the manuscript. D J Y contributed to isolate the strains and analyze the data. M-J S designed all the experiments and supervised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that there is no conflict of interest.
Ethical approval
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hwang, C.Y., Cho, ES., Yoon, D.J. et al. Halobellus ruber sp. nov., a deep red-pigmented extremely halophilic archaeon isolated from a Korean solar saltern. Antonie van Leeuwenhoek 114, 997–1011 (2021). https://doi.org/10.1007/s10482-021-01571-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01571-1