Abstract
A proteomics approach was used to evaluate the effects of flooding stress on early symbiotic interaction between soybean roots and soil bacteria, Bradyrhizobium japonicum. Three-day-old soybean was inoculated with B. japonicum followed by flooding. The number of root hairs in seedlings, without or with flooding stress, was increased after 3 days of inoculation. Proteins were extracted from roots and separated by two-dimensional polyacrylamide gel electrophoresis. Out of 219 protein spots, 14 and 8 proteins were increased and decreased, respectively, by inoculation under flooding compared with without flooding. These proteins were compared in untreated and flooded seedlings. Increased level of 6 proteins in flooded seedlings compared with untreated seedlings was suppressed by inoculation in seedlings under flooding. They were related to disease/defense, protein synthesis, energy, and metabolism. Differential abundance of glucan endo-1,3-beta-glucosidase, phosphoglycerate kinase, and triosephosphate isomerase, based on their localization in middle and tip of root, indicated that they might be related to increase in number of root hairs. These results suggest that disease/defense, energy, and metabolism-related proteins may be particularly subjected to regulation in flooded soybean seedlings, when inoculated with B. japonicum and that this regulation may lead to increase in number of root hair during early symbiotic differentiation.





Similar content being viewed by others
Abbreviations
- 2-DE:
-
Two-dimensional polyacrylamide gel electrophoresis
- CBB:
-
Coomassie brilliant blue
- MS:
-
Mass spectrometry
- LC:
-
Liquid chromatography
- pI :
-
Isoelectric point
- IEF:
-
Isoelectric focusing
- MALDI-TOF:
-
Matrix-assisted laser desorption ionization time-of-flight
References
Bailey-Serres J, Freeling M (1990) Hypoxic stress-induced changes in ribosomes of maize seedling roots. Plant Physiol 94:1237–1243
Banks RD, Blake CCF, Evans PR, Haser R, Rice DW, Hardy GW, Merrett M, Phillips AW (1979) Sequence, structure and activity of phosphoglycerate kinase: a possible hinge-bending enzyme. Nature 279:773–777
Bevan M, Bancroft I, Bent E, Love K, Goodman H, Dean C, Bergkamp R, Dirkse W, Van Staveren M, Stiekema W, Drost L, Ridley P, Hudson SA, Patel K, Murphy G, Piffanelli P, Wedler H, Wedler E, Wambutt R, Weitzenegger T, Pohl TM, Terryn N, Gielen J, Villarroel R, De Clerck R, Van Montagu M, Lecharny A, Auborg S, Gy I, Kreis M, Lao N, Kavanagh T, Hempel S, Kotter P, Entian KD, Rieger M, Schaeffer M, Funk B, Mueller-Auer S, Silvey M, James R, Montfort A, Pons A, Puigdomenech P, Douka A, Voukelatou E, Milioni D, Hatzopoulos P, Piravandi E, Obermaier B, Hilbert H, Dusterhoft A, Moores T, Jones JDG, Eneva T, Palme K, Benes V, Rechman S, Ansorge W, Cooke R, Berger C, Delseny M, Voet M, Volckaert G, Mewes HW, Klosterman S, Schueller C, Chalwatzis N, Project EUAG (1998) Analysis of 1.9 Mb of contiguous sequence from chromosome 4 of Arabidopsis thaliana. Nature 391:485–488
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Chen M, Thelen JJ (2010) The essential role of plastidial triose phosphate isomerase in the integration of seed reserve mobilization and seedling establishment. Plant Signal Behav 5:583–585
Chen J, Chen C, Sung FJM (1992) Flooding effect on photosynthesis in soybean leaves. J Agric 41:17–26
Graham PH, Vance CP (2003) Legumes: importance and constraints to greater use. Plant Physiol 131:872–877
Hashiguchi A, Sakata K, Komatsu S (2009) Proteome analysis of early-stage soybean seedlings under flooding stress. J Proteome Res 8:2058–2069
Horton P, Park K-J, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, Nakai K (2007) WoLF PSORT: protein localization predictor. Nucleic Acids Res 35:W585–W587
Jindal HK, Vishwanatha JK (1990) Functional identity of a primer recognition protein as phosphoglycerate kinase. J Biol Chem 265:6540–6543
Komatsu S, Wada T, Abalea Y, Nouri M-Z, Nanjo Y, Nakayama N, Shimamura S, Yamamoto R, Nakamura T, Furukawa K (2009a) Analysis of plasma membrane proteome in soybean and application to flooding stress response. J Proteome Res 8:4487–4499
Komatsu S, Yamamoto R, Nanjo Y, Mikami Y, Yunokawa H, Sakata K (2009b) A comprehensive analysis of the soybean genes and proteins expressed under flooding stress using transcriptome and proteome techniques. J Proteome Res 8:4766–4778
Komatsu S, Sugimoto T, Hoshino T, Nanjo Y, Furukawa K (2010) Identification of flooding stress responsible cascades in root and hypocotyl of soybean using proteome analysis. Amino Acids 38:729–738
Lerouge P, Roche P, Faucher C, Maillet F, Truchet G, Prome JC, Denarie J (1990) Symbiotic host-specificity of Rhizobium meliloti is determined by a sulphated and acylated glucosamine oligosaccharide signal. Nature 344:781–784
Libault M, Farmer A, Brechenmacher L, Drnevich J, Langley RJ, Bilgin DD, Radwan O, Neece DJ, Clough SJ, May GD, Stacey G (2010) Complete transcriptome of the soybean root hair cell, a single-cell model, and its alteration in response to Bradyrhizobium japonicum infection. Plant Physiol 152:541–552
Minchin FR, Summerfield RJ (1976) Symbiotic nitrogen-fixation and vegetative growth of cowpea (Vigna-unguiculata (L) Walp) in waterlogged conditions. Plant Soil 45:113–127
Mitra RM, Long SR (2004) Plant and bacterial symbiotic mutants define three transcriptionally distinct stages in the development of the Medicago truncatula/Sinorhizobium meliloti symbiosis. Plant Physiol 134:595–604
Muszynski A, Laus M, Kijne JW, Carlson RW (2011) Structures of the lipopolysaccharides from Rhizobium leguminosarum RBL5523 and its UDP-glucose dehydrogenase mutant (exo5). Glycobiol 21:55–68
Nanjo Y, Skultety L, Ashraf Y, Komatsu S (2010) Comparative proteomic analysis of early-stage soybean seedlings responses to flooding by using gel and gel-free techniques. J Proteome Res 9:3989–4002
Nap J, Bisseling T (1990) Nodulin function and nodulin gene regulation in root nodule development. In: Gresshoff PM (ed) The molecular biology of symbiotic nitrogen fixation. CRC Press, Boca Raton, pp 181–229
Nelson DL, Cox MM (2004a) Protein metabolism. In: Macmillan P (ed) Lehninger principles of biochemistry. Basingstoke, Hampshire, pp 1034–1080
Nelson DL, Cox MM (2004b) Bioenergetics and metabolism. In: Macmillan P (ed) Lehninger principles of biochemistry. Basingstoke, Hampshire, pp 480–891
Ogino T, Iwama M, Kinouchi J, Shibagaki Y, Tsukamoto T, Mizumoto K (1999) Involvement of a cellular glycolytic enzyme, phosphoglycerate kinase, in Sendai virus transcription. J Biol Chem 274:35999–36008
Russell DA, Wong DML, Sachs MM (1990) The anaerobic response of soybean. Plant Physiol 92:401–407
Salavati A, Bushehri AAS, Taleei A, Hiraga S, Komatsu S (2012) A comparative proteomic analysis of the early response to compatible symbiotic bacteria in the roots of a supernodulating soybean variety. J Proteomics 75:819–832
Sallam A, Scott HD (1987) Effects of prolonged flooding on soybean at the R2 growth stage. Dry matter and N and P accumulation. J Plant Nutr 10:567–592
Sanchez F, Padilla JE, Perez H, Lara M (1991) Control of nodulin genes in root-nodule development and metabolism. Annu Rev Plant Physiol Plant Mol Biol 42:507–528
Sanchez C, Tortosa G, Granados A, Delgado A, Bedmar EJ, Delgado MJ (2011) Involvement of Bradyrhizobium japonicum denitrification in symbiotic nitrogen fixation by soybean plants subjected to flooding. Soil Biol Biochem 43:212–217
Schmutz J, Cannon SB, Schlueter J, Ma J, Mitros T, Nelson W, Hyten DL, Song Q, Thelen JJ, Cheng J, Xu D, Hellsten U, May GD, Yu Y, Sakurai T, Umezawa T, Bhattacharyya MK, Sandhu D, Valliyodan B, Lindquist E, Peto M, Grant D, Shu S, Goodstein D, Barry K, Futrell-Griggs M, Abernathy B, Du J, Tian Z, Zhu L, Gill N, Joshi T, Libault M, Sethuraman A, Zhang X-C, Shinozaki K, Nguyen HT, Wing RA, Cregan P, Specht J, Grimwood J, Rokhsar D, Stacey G, Shoemaker RC, Jackson SA (2010) Genome sequence of the palaeopolyploid soybean. Nature 463:178–183
Sung FJM (1993) Waterlogging effects on nodule nitrogenase and leaf nitrate reductase activities in soybean. Field Crops Res 35:183–189
Tenhaken R, Thulke O (1996) Cloning of an enzyme that synthesizes a key nucleotide-sugar precursor of hemicellulose biosynthesis from soybean: UDP-glucose dehydrogenase. Plant Physiol 112:1127–1134
Tesfaye M, Silverstein KA, Bucciarelli B, Samac DA, Vance CP (2006) The affymetrix medicago genechip array is applicable for transcript analysis of alfalfa (Medicago sativa). Funct Plant Biol 33:783–788
Ward ER, Payne GB, Moyer MB, Williams SC, Dincher SS, Sharkey KC, Beck JJ, Taylor HT, Ahl-Goy P, Meins F, Ryals JA (1991) Differential regulation of β-1,3-glucanase messenger rnas in response to pathogen infection. Plant Physiol 96:390–397
Zhao Z, Assmann SM (2011) The glycolytic enzyme, phosphoglycerate mutase, has critical roles in stomatal movement, vegetative growth, and pollen production in Arabidopsis thaliana. J Exp Bot 62:5179–5518
Acknowledgments
The authors thank the students exchange programme between Kohat University of Science and Technology, Pakistan, and University of Tsukuba, Japan, for providing scholarship. They are grateful to the National Institute of Crop Science of Japan for all experimental support during this project. The authors thank Dr. Yohei Nanjo and Dr. Keito Nishazawa for their valuable discussion. They are also thankful to Dr. Kentaro Kawaguchi and Dr. Md. Emdadul Haque for support during microscopic analysis. This work was supported by the grants from National Agriculture and Food Research Organization, Japan. Bacterial strain, B. japonicum MAFF 211342 was obtained from the Genebank at National Institute of Agrobiological Sciences (Tsukuba, Japan).
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
726_2012_1333_MOESM2_ESM.pptx
Supplementary Fig 1. Experimental design of effects of flooding stress on early symbiotic interaction in soybean roots inoculated with compatible bacteria. Experimental design representing physiological (A) and proteomics (B) work flow. Three-day-old soybean was inoculated with B. japonicum without or with flooding stress. Untreated soybean served as control. a, b, c and d represent, no treatment, inoculation, inoculation under flooding and flooding treatment, respectively. Time of sowing, inoculation, initiation of flooding, and sampling are specified by downward triangles, inverted arrow, upward triangles, and circles, respectively. Statistical analyses were performed between the treatments as shown by the lines connecting the treatments vertically, using Student’s t test and one-way ANOVA Duncan’s multiple comparisons test when 2 or more than 2 treatments were compared, respectively. (PPTX 53 kb)
726_2012_1333_MOESM3_ESM.pptx
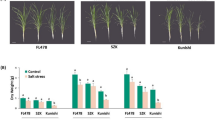
Supplementary Fig. 2. Comparision of shoot length and fresh weight of soybean seedlings between untreated seedlings, inoculated seedlings, seedlings inoculated under flooding, and flooded seedlings. Three-day-old soybean was inoculated with B. japonicum followed by flooding. Untreated soybean served as control. Lengths and fresh weights of epicotyls (E) and hypocotyl (H) were measured 0, 3 and 6 days after treatment. Each bar represents the average ± SE of 18 seedlings. White, black, squared, and dotted bars represent untreated seedlings, inoculated seedlings, inoculated seedlings under flooding, and flooded seedlings, respectively. Statistical analysis was performed by one-way ANOVA Duncan’s multiple comparisons test. Different letters above the bars indicate a statistically significant difference (P < 0.05). (PPTX 46 kb)
726_2012_1333_MOESM4_ESM.pptx
Supplementary Fig. 3. Morphological comparison of root hair growth response between untreated seedlings, inoculated seedlings, seedlings inoculated under flooding, and flooded seedlings. Three-day-old soybean was inoculated with B. japonicum and flooded. Soybean without any treatment served as control. Root samples were collected 3 and 6 days after treatment and stained with methylene blue. Number of root hairs per 200 μm2 of root was determined under light microscope (4X, 10X, 20X). (PPTX 1023 kb)
726_2012_1333_MOESM5_ESM.pptx
Supplementary Fig. 4. Comparison of 2-DE pattern of protein abundance in soybean roots between untreated seedlings, inoculated seedlings, seedlings inoculated under flooding, and flooded seedlings. Three-day-old soybean was inoculated with B. japonicum and flooded. Untreated soybean served as control. Proteins were extracted from roots 3 days after treatment, separated by 2-DE and stained by CBB. The abundance pattern of 22 protein spots, which differentially changed in soybean roots inoculated with B. japonicum under flooding, was compared with untreated and flooded seedlings. Open circles show protein spots with altered abundance. Upward and downward arrows indicate increased and decreased abundance, respectively. Spot numbers are same as in Fig. 2. (PPTX 693 kb)
726_2012_1333_MOESM6_ESM.pptx
Supplementary Fig. 5. Comparison of protein abundance in soybean roots between untreated seedlings, inoculated seedlings, seedlings inoculated under flooding, and flooded seedlings. Three-day-old soybean was inoculated with B. japonicum and flooded. Untreated soybean served as control. Three days after treatments, proteins were extracted from roots of untreated seedlings (white bars), inoculated seedlings (black bars), inoculated seedlings under flooding (squared bars), and flooded soybean seedlings (dotted bars). The differentially changed proteins were quantified using PDQuest software and plotted as the relative intensity. Results are presented as mean ± SE of relative protein intensity for gels from three biological replicates. The results were compared using Student’s t test between the treatments as shown by line with asterisks above the bars, indicating significant differences between treatments (*P < 0.05, **P < 0.01). (PPTX 73 kb)
726_2012_1333_MOESM7_ESM.pptx
Supplementary Fig. 6. Photograph of soybean seedling showing different portions of root. Three-day-old soybean was inoculated with B. japonicum followed by flooding. Middle and tip of roots were taken for differential abundance analysis of proteins related to root hair number and morphology in respective portions of roots. (PPTX 150 kb)
Rights and permissions
About this article
Cite this article
Khatoon, A., Rehman, S., Salavati, A. et al. A comparative proteomics analysis in roots of soybean to compatible symbiotic bacteria under flooding stress. Amino Acids 43, 2513–2525 (2012). https://doi.org/10.1007/s00726-012-1333-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-012-1333-8