Abstract
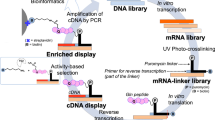
A large number of substrate proteins for tissue transglutaminase (TGase 2) have been identified in vivo and in vitro. Preference in primary sequence or secondary structure around the reactive glutamine residues in the substrate governs the reactivity for TGase 2. We established a screening system to identify preferable sequence as a glutamine-donor substrate using a phage-displayed peptide library. The results showed that several peptide sequences have higher reactivity and specificity to TGase 2 than those of preferable sequences previously reported. By analysis of the most reactive 12-amino acid sequence, T26 (HQSYVDPWMLDH), residues crucial to the enzymatic reaction were investigated. The following review summarizes the screening system and also the preference in substrate sequences that were obtained by this method and those previously reported.





Similar content being viewed by others
Abbreviations
- Bio-Cd:
-
Biotinylated cadaverine
- GST:
-
Glutathione-S-transferase
- MDC:
-
Monodansylcadaverine
- TGase 2:
-
Tissue-specific transglutaminase
References
Chen JSK, Mehta K (1999) Tissue transglutaminase: an enzyme with a split personality. Int J Biochem Cell Biol 31:817–836
Clackson T, Lowman HB (2004) Phage display. Oxford University Press, UK
Esposito C, Caputo I (2005) Mammalian transglutaminases: identification of substrates as a key to physiological function and physiological relevance. FEBS J 272:615–631
Facchiano AM, Facchiano A, Facchiano F (2003) Active sequences collection (ASC) database: a new tool to assign functions to protein sequences. Nucleic Acids Res 31:379–382
Fesus L, Piacentini M (2002) Transglutaminase 2: an enigmatic enzyme with diverstic functions. Trends Biochem Sci 27:534–539
Fleckenstein B, Molberg Ø, Qiao SW, Schmid DG, von der Mulbe F, Elgstoen K, Jung G, Sollid LM (2002) Gliadin T cell epitope selection by tissue transglutaminase in celiac disease. J Biol Chem 277:34109–34116
Gorman JJ, Folk JE (1984) Structural features of glutamine substrates for transglutaminases: role of extended interactions in the specificity of human plasma factor XIIIa and of the guinea pig liver enzyme. J Biol Chem 259:9007–9010
Griffin M, Casadio R, Bergamini CM (2002) Transglutaminases: nature’s biological glues. Biochem J 368:377–396
Grootjans JJ, Groenen PJ, de Jong WW (1995) Substrate requirements for transglutaminases. J Biol Chem 270:22855–22858
Groenen PJ, Bloemendal H, de Jong WW (1992) The carboxy-terminal lysine αB-crystalline is an amine donor substrate for tissue transglutaminase. Eur J Biochem 205:671–674
Hettasch JM, Peoples KA, Greenberg CS (1997) Analysis of factor XIII substrate specificity using recombinant human factor XIII and tissue transglutaminase chimeras. J Biol Chem 272:25149–25156
Ichinose A, Tamaki T, Aoki N (1983) Factor XIII-mediated cross-linking of NH2-terminal peptide of α2-plasmin inhibitor to fibrin. FEBS lett 153:369–371
Kamiya N, Doi S, Tominaga J, Ichinose H, Goto M (2005) Transglutaminase-mediated protein immobilization to casein nanolayers created on a plastic surface. Biomacromolecules 6:35–38
Keresztessy Z, Csosz E, Harsfalvi J, Csomos K, Gray J, Lightowlers RN, Lakey JH, Balajthy Z, Fesus L (2006) Phage display selection of efficient glutamine-donor substrate peptides for transglutaminase 2. Protein Sci 9:2466–2480
Lorand L, Parameswaran KN, Velasco PT (1991) Sorting-out of acceptor-donor relationship in the transglutaminase catalyzed cross-linking of crystallines by the enzyme-directed labeling of potential sites. Proc Natl Acad Sci USA 88:82–83
Lorand L, Graham RM (2003) Transglutaminases: crosslinking enzymes with pleiotropic functions. Nat Rev Mol Cell Biol 4:140–156
Parameswaran KN, Velasco PT, Wilson J, Lorand L (1990) Labeling of ε-lysine cross-linking sites in proteins with peptide substrate of factor XIIIa and transglutaminase. Proc Natl Acad Sci USA 87:8472–8475
Pastor MT, Diez A, Peretz-Paya E, Abad C (1999) Addressing substrate glutamine requirements for tissue transglutaminase using substance P analogue. FEBS Lett 451:231–234
Pucci P, Malorni A, Marino G, Metafora S, Esposito C, Porta R (1988) β-endorphin modification by transglutaminase in vitro: identification by FAB/MS of glutamine-11 and lysine-29 as acyl donor and acceptor sites. Biochem Biophys Res Commun 154:735–740
Qiao SW, Bergseng E, Molberg Ø, Jung G, Fleckenstein B, Sollid LM (2005) Refining the rules of gliadin T cell epitope binding to the disease-associated DQ2 molecule in celiac disease: importance of proline spacing and glutamine deamidation. J Immunol 175:254–261
Ruoppolo M, Orru S, D’Amato A, Francese S, Rovero P, Marino G, Esposito C (2003) Analysis of transglutaminase protein substrates by functional proteomics. Protein Sci 12:1290–1297
Schalkwijk J, Wiedow O, Hirose S (1999) The trappin gene family: proteins defined by an N-terminal transglutaminase substrate domain and a C-terminal four-disulphide core. Biochem J 340:569–577
Sollid LM (2002) Coeliac disease: dissecting a complex inflammatory disorder. Nat Rev Immunol 2:647–655
Steinert PM, Candi E, Tarcsa E, Marekov LN, Sette M, Paci M, Ciani B, Guerrieri P, Melino G (1999) Transglutaminase crosslinking and structural studies of the human small proline rich 3 protein. Cell Death Differ 6:916–930
Sugimura Y, Hosono M, Wada F, Yoshimura T, Maki M, Hitomi K (2006) Screening for the preferred substrate sequence of transglutaminase using a phage-displayed peptide library: identification of peptide substrates for TGase 2 and Factor XIIIa. J Biol Chem 281:17699–17706
Sugimura Y, Ueda H, Maki M, Hitomi K (2007) Novel site-specific immobilization of a functional protein using a preferred substrate sequence for transglutaminase 2. J Biotechnol 131:121–127
Tarcsa E, Marekov LN, Andreoli J, Idler WW, Candi E, Chung S-I, Steinert PM (1997) The fate of trichohyalin: sequential post-translational modifications by peptidyl-arginine deiminase and transglutaminase. J Biol Chem 272:27893–27901
Tanaka Y, Tsuruda Y, Nishi M, Kamiya N, Goto M (2007) Exploring enzymatic catalysis at a solid surface: a case study with transglutaminase-mediated protein immobilization. Org Biomol Chem 5:1764–1770
Vader LW, Ru A, van der Wal Y, Kooy YMC, Benckhuijsen W, Mearin ML, Drijfhout JW, van Veelen P, Koning F (2002) Specificity of tissue transglutaminase explains cereal toxicity in celiac disease. J Exp Med 195:643–649
Valdivia A, Villalonga R, Di Pierro P, Perez Y, Mariniello L, Gomez L, Porta R (2006) Transglutaminase-catalyzed site-specific glycosidation of catalase with aminated dextran. J Biotechnol 122:326–333
Zeeuwen PL, Hendriks W, de Jong WW, Schalkwijk J (1997) Identification and sequence analysis of two new members of the SKALP/elafin and SPAI-2 gene family. J Biol Chem 272:20471–20478
Acknowledgment
This work was supported by Grant-in-Aid for Scientific Research (C) 19580103 (to K.·H.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Hitomi, K., Kitamura, M. & Sugimura, Y. Preferred substrate sequences for transglutaminase 2: screening using a phage-displayed peptide library. Amino Acids 36, 619–624 (2009). https://doi.org/10.1007/s00726-008-0126-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-008-0126-6