Abstract
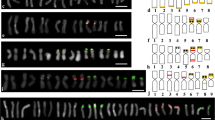
Genomic in situ hybridization offers a powerful tool for investigating genome organisation and evolution of taxa known, or suspected, to be allopolyploids. The question of the diploid progenitors of cultivated peanut (Arachis hypogaea, 2n=4x=40) has been the subject of numerous studies at cytogenetical, cytochemical, biochemical and molecular levels, but no definitive conclusions have been reached. The biotinylated total genomic DNA from potential diploidArachis species were separately hybridized in situ to root tip chromosomes ofA. hypogaea and wild speciesA. monticola (2n=4x=40) without or mixed with an excess of unlabelled DNA from the species not used as a probe. Among the range of different species combinations used, the strong and uniform signals given by labelledA. ipaensis DNA when hybridized toA. hypogaea andA. monticola in combination with unlabelledA. villosa DNA indicates that overall molecular composition of twenty chromosomes ofA. hypogaea andA. monticola is very similar toA. ipaensis chromosomes. ProbingA. hypogaea andA. monticola chromosomes with labelled genomic DNA fromA. villosa mixed with unlabelled DNA fromA. ipaensis likewise labelled strongly and uniformly the other twenty chromosomes. BarringA. ipaensis, all the diploidArachis species presently investigated had characteristic centromeric bands in the twenty chromosomes within the complement indicating a clear division ofA. ipaensis from other species. InA. hypogaea andA. monticola only twenty chromosomes showed centromeric bands. These results (i) confirm the allopolyploid nature ofA. hypogaea andA. monticola, (ii) strongly support the view that wildA. monticola and cultivatedA. hypogaea are very closely related, and (iii) indicate thatA. villosa andA. ipaensis are the diploid wild progenitors of the tetraploid species studied. The present results also reveal that the nucleolus organizing region (NOR) originating fromA. villosa alone is expressed in the two tetraploid species.
Similar content being viewed by others
References
Abdou, Y. A.-M., Gregory, W. C., Cooper, W. E., 1974: Sources and nature of resistance toCercospora arachidicola Hori andCercosporidium personatum (Beck & Curtis)Deighton inArachis species. — Peanut Sci.1: 6–11.
Anamthawat-Jónsson, K., Schwarzacher, T., Leitch, A. R., Bennett, M. D., Heslop-Harrison, J. S., 1990: Discrimination between closely relatedTriticeae species using genomic DNA as a probe. — Theor. Appl. Genet.79: 721–728.
—, 1993: Behaviour of parental genomes in the hybridHordeum vulgare xH. bulbosum. — J. Heredity84: 78–82.
Bass, M. H., Arant, F. S., 1973: Insect pests. — InSummerfield, R. J., Bunting, A. H., (Eds): Peanuts — Culture and uses, pp. 338–428. — Stillwater, OK: American Peanut Research Education Association.
Bennett, M. D., 1995: The development and use of genomic in situ hybridization (GISH) as a new tool in plant biosystematics. — InBrandham, P. E., Bennett, M. D., (Eds): Kew Chromosome Conference IV, pp. 167–183. — Richmond: Royal Botanic Gardens, Kew.
Bianchi-Hall, C. M., Keys, R. D., Stalker, H. T., Murphy, J. P., 1993: Diversity of seed storage protein patterns in wild peanut (Arachis, Fabaceae) species. — Pl. Syst. Evol.186: 1–15.
Dhesi, J. S., Stalker, H. T., 1994: Enhancing techniques for studying mitotic peanut chromosomes. — Peanut Sci.21: 92–94.
Duke, J. A., 1981: Handbook of legumes of world economic importance. — New York: Plenum Press.
Fernandez, A., Krapovickas, A., 1994: Cromosomas y evolucion enArachis (Leguminosae). — Bonplandia8: 187–220.
Foster, D. J., Stalker, H. T., Wynne, J. C., Beute, M. K., 1981: Resistance ofArachis hypogaea and wild relatives toCercospora arachidicola Hori. — Oleagineux36: 139–143.
Garren, K. H., Jackson, C. R., 1973: Peanut diseases. — InSummerfield, R. J., Bunting, A. H., (Eds): Peanuts-Culture and uses, pp. 429–494. — Stillwater, OK: American Peanut Research Education Association.
Gregory, M. P., Gregory, W. C., 1979: Exotic germplasm ofArachis L. interspecific hybrids. — J. Heredity70: 185–193.
Gregory, W. C., Gregory, M. P., 1976: GroundnutArachis hypogaea (Leguminosae-Papilionatae). — InSimmonds, N. W., (Ed.): Evolution of crop plants, pp. 151–154. — London: Longman.
—, 1980: Structure, variation, evolution, and classification inArachis. — InSummerfield, R. J., Bunting, A. H., (Eds): Advances in legume science, pp. 469–481. — Richmond: Royal Botanic Gardens, Kew.
Halward, T. M., Stalker, H. T., LaRue, E. A., Kochert, G. D., 1992: Use of single primer DNA amplifications in genetic studies of peanut (Arachis hypogaea L.). — Pl. Molec. Biol.18: 315–325.
Johnson, D. R., Wynne, J. C., Campbell, W. V., 1977: Resistance of wild species ofArachis to the twospotted spider mite,Tetranychus urticae. — Peanut Sci.4: 9–11.
Kirti, P. B., Murty, U. R., Bharati, M., Rao, N. G. P., 1982: Chromosome pairing in F1 hybridArachis hypogaea ×A. monticola Krap. etRig. — Theor. Appl. Genet.62: 139–144.
Klozova, E., Turkova, V., Smartt, J., Pitterova, K., Svachulova, J., 1983: Immunochemical characterization of seed proteins of some species of the genusArachis L. — Biol. Pl.25: 201–208.
Kochert, G., Halward, T., Branch, W. D., Simpson, C. E., 1991: RFLP variability in peanut (Arachis hypogaea L.) cultivars and wild species. — Theor. Appl. Genet.81: 565–570.
—, 1996: RFLP and cytogenetic evidence on the origin and evolution of allotetraploid domesticated peanut,Arachis hypogaea (Leguminosae). — Amer. J. Bot.83: 1282–1291.
Krapovickas, A., Gregory, W. C., 1994: Taxonomical del generaArachis (Leguminosae). — Bonplandia8: 1–184.
—, 1957: Nuevas aspecies deArachis vinculadas al problema del origen del mani. — Darwiniana11: 431–455.
—, 1974: Recuperaction de la fertilidad en un hibgido interspecifico deArachis (Leguminosae). — Bonplandia3: 129–142.
Krishna, T. G., Mitra, R., 1988: The probable genome donors toArachis hypogaea L. based on arachin seed storage protein. — Euphytica37: 47–52.
Lanham, P. G., Forster, B. P., McNicol, P., Moss, J. P., Powell, W., 1994: Seed storage protein variation inArachis species. — Genome37: 487–496.
Lu, J., Pickersgill, B., 1993: Isozyme variation and species relationships in peanut and its wild relatives (Arachis L.-Leguminosae). — Theor. Appl. Genet.85: 550–560.
Mukai, Y., 1996: Multicolor in situ hybridization: A new tool for genome analysis. — InJauhar, P. P., (Ed.): Methods of genome analysis in plants, pp. 182–192. — Boca Raton: CRC Press.
Paik-Ro, O. G., Smith, R. L., Knauft, D. A., 1992: Restriction fragment length polymorphism evaluation of six peanut species within theArachis section. — Theor. Appl. Genet.84: 201–208.
Raman, V. S., Kesavan, P. C., 1962: Studies on a diploid interspecific hybrid inArachis. — Nucleus5: 123–126.
Resslar, P. M., 1980: A review of the nomenclature of the genusArachis L. — Euphytica29: 813–817.
—, 1979: A cytological study of three diploid species of the genusArachis L. — J. Heredity70: 13–16.
Saghai-Maroof, M. A., Soliman, K. M., Jorgensen, R. A., Allard, R. W., 1984: Ribosomal DNA spacer-length polymorphisms in barley: Mendelian inheritance, chromosomal location, and population dynamics. — Proc. Natl. Acad. Sci. USA81: 8014–8018.
Seetharam, A., Nayar, K. M. D., Sreekantaradhya, R., Acher, D. K. T., 1973: Cytological studies on the interspecific hybrid ofArachis hypogaea ×Arachis duranensis. — Cytologia38: 277–280.
Singh, A. K., 1986: Utilization of wild relatives in the genetic improvement ofArachis hypogaea L. 8. Synthetic amphidiploids and their importance in interspecific breeding. — Theor. Appl. Genet.72: 433–439.
—, 1988: Putative genome donors ofArachis hypogaea (Fabaceae), evidence from crosses with synthetic amphidiploids. — Pl. Syst. Evol.160: 143–151.
—, 1982: Utilization of wild relatives in genetic improvement ofArachis hypogaea L. Part 2: Chromosome complements of species in sectionArachis. — Theor. Appl. Genet.61: 305–314.
—, 1991: Phylogenetic relations in sectionArachis based on seed protein profile. — Theor. Appl. Genet.82: 593–597.
Singh, K. P., Raina, S. N., Singh, A. K., 1996: Variation in chromosomal DNA associated with the evolution ofArachis species. — Genome39: 890–897.
Smartt, J., Stalker, H. T., 1982: Speciation and cytogenetics inArachis. — InPattee, H. E., Young, C. T., (Eds): Peanut science and technology, pp. 21–49. — Yokum, TX: American Peanut Research and Education Society.
—, 1978a: The genomes ofArachis hypogaea. 1. Cytogenetic studies of putative genome donors. — Euphytica27: 665–675.
—, 1978b: The genomes ofArachis hypogaea. 2. The implications in interspecific breeding. — Euphytica27: 677–680.
Stalker, H. T., Dalmacio, R. D., 1981: Chromosomes ofArachis species, sectionArachis. — J. Heredity72: 403–408.
—, 1987: Speciation, cytogenetics and utilization ofArachis species. — Advances Agron.41: 1–38.
—, 1979: Cytology of interspecific hybrids in sectionArachis of peanuts. — Peanut Sci.6: 110–114.
—, 1994: Variation of isozyme patterns amongArachis species. — Theor. Appl. Genet.87: 746–755.
Varisai Muhammad, S., 1973: Cytogenetical investigations in the genusArachis L. II. Triploid hybrids and their derivatives. — Madras Agric. J.60: 1414–1427.
Wynne, J. C., Halward, T., 1989: Cytogenetics and genetics ofArachis. — Pl. Sci.8: 189–220.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Raina, S.N., Mukai, Y. Genomic in situ hybridization inArachis (Fabaceae) identifies the diploid wild progenitors of cultivated (A. hypogaea) and related wild (A. monticola) peanut species. Pl Syst Evol 214, 251–262 (1999). https://doi.org/10.1007/BF00985743
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00985743