Abstract
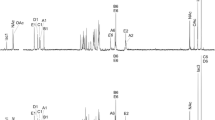
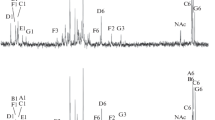
The lipopolysaccharides (LPS) of a rough (R) and a smooth (S) strain of Pseudomonas syringae pv. phaseolicola were analysed. The S-LPS revealed markedly more rhamnose and fucose, but less glucose, than the R-LPS. The presence of 3-O-methyl-rhamnose (acofriose) in the S-LPS was confirmed by cochromatography with authentic acofriose. SDS polyacrylamide gel electrophoresis of the S-LPS demonstrated a cluster of regularly spaced high molecular weight fractions, which was almost lacking in the R-LPS. The main fatty acids of the lipid A of both LPS species were 3-OH-10:0,3-OH-12:0,2-OH-12:0, and 12:0. Two N-linked diesters were demonstrated: 3-O(12:0)-12:0 and 3-O(2-OH-12:0)-12:0. S-LPS was subjected to mild hydrolysis and the “degraded polysaccharide” separated into three fractions by gel permeation chromatography on a Fractogel TSK HW-50 column. Fraction I, representing nearly only the O-specific side chain, consisted of rhamnose and fucose in a molar ratio of 4:1, with 4% of the rhamnose being 3-O-methylated (acofriose). Fraction II, representing mostly core material, was composed of glucose, rhamnose, heptose, glucosamine, galactosamine, alanine, and a still unidentified amino compound, in an approximate molar ratio of 3:1:1:1:1:1:1, and KDO. Fraction III consisted of released monomers and salts. The LPS was highly phosphorylated (3.28% phosphorus in the “core fraction”). The thus characterized composition of the LPS O-chain seems to be unique for the pathovar phaseolicola of P. syringae, although many similarities exist to other pathovars as well as to other bacterial species.
Similar content being viewed by others
Abbreviations
- LPS:
-
lipopolysacchairdes
- GC/MS:
-
combined gas liquid chromatography-mass spectrometry
- HVE:
-
high voltage electrophoresis
- KDO:
-
2-keto-3-deoxyoctonic acid
- PAGE:
-
polyacrylamide gel electrophoresis
- SDS:
-
sodium dodecylsulfate
References
Adam DB, Pugsley AT (1934) “Smooth-rough” variation in Phytomonas medicaginis phaseolicola Burk. Austr J Exp Biol Med Sci 12:193–201
Anderson AJ (1984) Differences between lipopolysaccharide compositions of plant pathogenic and saprophytic Pseudomonas species. Appl Env Microbiol 48:31–35
Anderson PJ (1966) A sensitive reagent for detecting 2-deoxysugars and 3-deoxypolyols. J Chromatogr 21:163–164
Baker CJ, Neilson MJ, Sequeira L, Keegstra KG (1984) Chemical characterization of the lipopolysaccharide of Pseudomonas solanacearum. Appl Env Microbiol 47:1096–1100
Bartlett GR (1959) Phosphorus assay in column chromatography. J Biol Chem 234:466–468
Barton-Willis PA, Wang MC, Holliday MJ, Long MR, Keen NT (1984) Purification and composition of lipopolysaccharides from Pseudomonas syringae pv. glycinea. Physiol Plant Pathol 25:387–398
Barton-Willis PA, Wang MC, Staskawicz B, Keen NT (1987) Structural studies on the O-chain polysaccharides of lipopolysaccharides from Pseudomonas syringae pv. glycinea. Physiol Mol Plant Pathol 30:187–197
Basu S, Radziejewska-Lebrecht J, Mayer H (1986) Lipopolysaccharides of Providencia rettgeri. Arch Microbiol 144:213–218
DeVos P, DeLey J (1983) Intra- and intergenic similarities of Pseudomonas and Xanthomonas ribosomal ribonucleic acid cistrons. Int J Syst Bacteriol 33:487–509
Drewry DT, Symes KC, Gray GW, Wilkinson SG (1975) Studies of polysaccharide fractions from the lipopolysaccharide of Pseudomonas aeruginosa NCTC 1999. Biochem J 149:93–106
Evans LR, Linker A (1973) Production and characterization of the slime polysaccharide of Pseudomonas aeruginosa. J Bacteriol 116:915–924
Fett WF, Osman SF, Fishman ML, Siebles TS (1986) Alginate production by plant-pathogenic pseudomonads. Appl Env Microbiol 52:466–473
Galanos C, Lüderitz O, Westphal O (1969) A new method for the extraction of R-lipopolysaccharides. Eur J Biochem 9:245–249
Gerwe PG, Rudolph K, Köhn S (1987) Vergleich eines “glatten” und eines “rauhen” Stammes von Pseudomonas syringae pv. phaseolicola. Phytopath 118:326–334
Gross M, Rudolph K (1987) Demonstration of levan and alginate in bean plants (Phaseolus vulgaris) infected by Pseudomonas syringae pv. phaseolicola. J Phytopathol 120:9–19
Hendrick CA, Sequeira L (1984) Lipopolysaccharide-defective mutants of the wilt pathogen Pseudomonas solanacearum. Appl Env Microbiol 48:94–101
Jansson PE, Lindberg AA, Lindberg B, Wollin R (1981) Structural studies on the hexose region of the core in lipopolysaccharides from Enterobacteriaceae. Eur J Biochem 115:571–577
Karkhanis YD, Zetner J, Jackson JJ, Carlo DJ (1978) A new and improved micro-assay to determine 2-keto-3-deoxyoctonate in lipopolysaccharide of Gram-negative bacteria. Anal Biochem 85:595–601
Keen NT, Williams PH (1971) Chemical and biological properties of a lipomucopolysaccharide from Pseudomonas lachrymans. Physiol Plant Pathol 1:247–264
King EO, Ward MK, Raney DE (1954) Two simple media for the demonstration of pyocyanin and fluorescin. J Lab Clin Med 44:301–307
Klement Z, Rudolph K, Ebrahim-Nesbat F (1984) Comparison of rough and smooth strains of Pseudomonas phaseolicola: Symptom development on a macroscopic and microscopic scale and multiplication rates in the leaves. Proc. 2nd Working Group on Pseudomonas syringae Pathovars. Sounion, Greece, pp 76–77
Kotelko K (1986) Proteus mirabilis: Taxonomic position, pecularities of growth, components of the cell envelope. Curr Top Microbiol Immunol 129:181–215
Lowry OH, Roberts NR, Leiner KY, Mu ML, Farr AL (1954) The quantitative histochemistry of brain. I. Chemical methods. J Biol Chem 207:1–17
Palleroni NJ (1984) Family 1. Pseudomonaceae. In: Krieg NR, Holt JG (eds) Bergey's manual of systematic bacteriology, vol 1. Williams and Wilkens, Baltimore, pp 141–219
Romanowska E, Gamian A, Dabrowski J (1986) Core region of Citrobacter lipopolysaccharide from strain PCM 1487. Structure elucidation by two-dimensional 1H-NMR spectroscopy at 500 MHz and methylation analysis/mass spectrometry. Eur J Biochem 161:557–564
Rowe PSN, Meadow PM (1983) Structure of the core oligosaccharide from the lipopolysaccharide of Pseudomonas aeruginosa PAC 1 R and its defective mutants. Eur J Biochem 132:329–337
Sawardeker JS, Sloneker JH, Jeanes A (1965) Quantitative determination of monosaccharides as their alditol acetates by gas liquid chromatography. Anal Chem 37:1602–1604
Sequeira L, Graham TL (1977) Agglutination of avirulent strains of Pseudomonas phaseolicola by potato lectin. Physiol Plant Pathol 11:43–54
Smith ARW, Zamze SE, Hignett RC (1985) Composition of lipopolysaccharide from Pseudomonas syringae pv. morsprunorum and its digestion by bacteriophage A7 J Gen Microbiol 131:963–974
Tharanathan RN, Weckesser J, Mayer H (1978) Location of O-methyl sugars in antigenic (lipo-)polysaccharides of photosynthetic bacteria and cyanobacteria. Biochem J 171:403–408
Tsai CM, Frash CE (1982) A sensitive silver stain for detecting lipopolysaccharides in polyacrylamide gels. Anal Biochem 119:115–119
Westphal O, Lüderitz O, Bister F (1952) Über die Extraktion von Bakterien mit Phenol/Wasser. Z Naturforsch 7b:148–155
Wilkinson SG (1977) Composition and structure of bacterial lipopolysaccharides. In: Sutherland I (ed) Surface carbohydrates of the procaryotic cell. Academic Press, London, pp 97–175
Whatley MH, Hunter, N, Cantrell MA, Hendrick C, Keegstra K, Sequeira L (1980) Lipopolysaccharide composition of the wilt pathogen, Pseudomonas solanacearum. Plant Physiol 65:557–559
Wollenweber HW, Broady KW, Lüderitz O, Rietschel ET (1982) The chemical structure of lipid A. Demonstration of amide-linked 3-acyloxyacyl residues in Salmonella minnesota Re lipopolysaccharide. Eur J Biochem 124:191–198
Wollenweber H-W, Seydel U, Lindner B, Lüderitz O, Rietschel ET (1984) Nature and location of amide-bound (R)-3-acyloxyacyl groups in lipid A of lipopolysaccharides from various Gramnegative bacteria. Eur J Biochem 145:265–272
Author information
Authors and Affiliations
Additional information
P.s. pv. phaseolicola is termed P. phaseolicola in the text
Rights and permissions
About this article
Cite this article
Gross, M., Mayer, H., Widemann, C. et al. Comparative analysis of the lipopolysaccharides of a rough and a smooth strain of Pseudomonas syringae pv. phaseolicola . Arch Microbiol 149, 372–376 (1988). https://doi.org/10.1007/BF00411658
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00411658