Abstract
We provide details on the shared-memory parallelization for manycore architectures of the molecular dynamics framework ls1-mardyn, including an optimization of the SIMD vectorization for multi-centered molecules. The novel shared-memory parallelization scheme allows to retain Newton’s third law optimization and exhibits very good scaling on many-core devices such as a full Xeon Phi card running 240 threads. The Xeon Phi can thus be exploited and delivers comparable performance as IvyBridge nodes in our experiments.
You have full access to this open access chapter, Download conference paper PDF
Similar content being viewed by others
Keywords
- Interaction Site
- Force Calculation
- Specific Computer System
- Floating Point Performance
- Scatter Operation
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
1 Introduction
Molecular Dynamics (MD) simulations are an established tool in life sciences [5] and, more recently, in chemical and process engineering [8]. The latter requires the simulation of a large number of particles, reaching the order of millions, or even more. In particular, the software framework ls1-mardyn [11] has specialized on this field, providing efficient treatment for short-range, non-bonded, 12-6 Lennard-Jones interactions of rigid multi-centered molecules via the linked-cell method.
Due to the compute intensive nature of MD, much research has been and currently is conducted to exploit modern HPC architectures such as GPUs [1] or the Xeon Phi [6, 12].
This paper presents enhancements and extensions of an original shared-memory parallelized Xeon Phi implementation of ls1-mardyn [7]. In [6, 7], it has been shown that a gather-scatter-based implementation is a promising approach to efficiently run MD simulations on Xeon Phi and potentially on the upcoming AVX-512 standard to Xeon. This is due to the standard “masking” SIMD approach becoming less efficient than the gather-scatter one for the vector width of eight (double-precision) elements [6]. We introduce an improvement in the gather-scatter SIMD vectorization, which renders the implementation efficient for bigger numbers of interaction sites per molecule.
We particularly focus on a new shared-memory implementation, which scales up to 240 threads on 60 Xeon Phi cores, thereby retaining Newton’s third law optimization. It is based on linked-cell coloring with eight synchronization stages, independent of the number of threads and linked-cells. It avoids the pitfalls of load-balancing and halo-layer overhead compared to domain decomposition variants using OpenMP [9, 12]. Tuned for scalability on the Xeon Phi, the shared-memory implementation also exhibits excellent performance on Xeon. We further investigate hyperthreading performance, as it plays a crucial role on Xeon Phi.
The optimized sequential and parallel performance allow us to draw significant comparisons between the performance on Xeon and Xeon Phi. It is demonstrated that Xeon Phi has the potential to keep up with or even outperform Xeon, and thus allows to double the performance of the complete host-accelerator system. As all presented improvements concern the force calculation kernel, we restrict this discussion to it. It represents the most time consuming part of MD simulations (60 % to \(\ge \) 90 % of total simulation time) as it locally exhibits a \(\mathcal {O}(N^2)\) behavior, while remaining operations are handled in \(\mathcal {O}(N)\).
We introduce the short-range MD problem in Sect. 2. The sequential and parallel algorithms and respective implementations are detailed in Sect. 3. Performance evaluations are provided in Sect. 4, including detailed single-core and full-node analysis for both Xeon Phi (MIC) and IVB nodes. We conclude and give an outlook to future work in Sect. 5.
2 Short-Range Molecular Dynamics
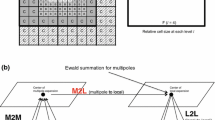
In ls1-mardyn, Newton’s equations of motion are solved for all molecules. Each molecule is modeled as rigid body and consists of \(N\ge 1\) interaction sites, cf. Fig. 1(a). The computationally most expensive part is given by the force accumulation: each interaction site of molecule i interacts with the interaction sites of all other molecules \(j\ne i\), \(i,j\in \{1,...,N\}\). For many target applications of ls1-mardyn, it is sufficient to only consider short-range interactions: molecules are sorted into linked-cells with mesh size \(\ge r_c\) and only molecules within a cut-off distance \(r_c\)—that is molecules inside a certain linked-cell, or its neighbored cells—interact with molecule i, cf. Fig. 1(b). Due to Newton’s third law (N3), each site-site interaction force acts identically (but with opposed signs) on each site. Each site-site pair thus needs to be evaluated only once [5]. This N3 optimization allows a reduction of computation time by a factor of 2. It, however, introduces race conditions if neighbored linked-cells are processed concurrently by differing threads, which represents a major obstacle to shared-memory parallelization [2]. For further details on the simulation core, we refer to [11].
3 Implementation
3.1 SIMD Vectorization
ls1-mardyn features a hand-written intrinsics kernel for the force calculation [3, 7]. Prior to the force calculation, the molecule data necessary for the force calculation is converted from an Array-of-Structures (AoS) layout to a Structure-of-Arrays (SoA) layout to enforce data contiguity and alignment. If no shared-memory parallelization is employed, only a “sliding window” of approx. three linked-cell layers is temporarily converted to SoA format [4]. We include a comparison to the sliding window non-OpenMP version, which we will refer to as win.
If shared-memory parallelization is employed, all cells need to be available in SoA format simultaneously, in order to enable their parallel processing. Moreover, it was observed, that freeing the SoAs after the force calculation and allocating them anew prior to the next force calculation can present a bottleneck in some cases. Thus, in the presented implementation, SoAs are not freed until the very end of the simulation. Reusing the allocated SoAs between iterations does not remove the necessity of “AoS-to-SoA” and “SoA-to-AoS” pre- and postprocessing steps, however: molecules may move between linked-cells and, thus, may need to be moved from one SoA to another, all the while maintaining the necessary contiguity and alignment.
Cutoff Mode. In ls1-mardyn, a center-of-mass (CoM) cutoff mode is employed. The cut-off condition needs therefore to be evaluated only between the centers-of-mass of the molecules (see Fig. 1(b)). Besides, all sites of a molecule always interact with full molecules and share the same interaction partners w.r.t. the cut-off condition.
Intrinsics Kernel. Listing 1.1 shows a schematic of the (already optimized) gather-scatter kernel for MIC. For every molecule m1 of soa1, first, the cutoff-condition between m1 and all molecules from soa2 is evaluated (line 4). The indices of the sites of the molecules from soa2, with which m1 will interact, are stored within the iV vector via the packstore intrinsic function. We proceed to compute the forces between the sites of m1 and the needed sites from soa2 (lines 6–12). If m1 has only one interaction center (line 6), we “broadcast-load” the site data of m1 into __mm512d registers, “gather-load” site-data from soa2 via iV and compute the first set of LJ-forces. The forces are then accumulated in an __mm512d register for m1 and scattered to soa2 via the Newton’s third law optimization (if applied). When the iteration is complete, the accumulated forces for the site of m1 are horizontally added to the respective SoA entry. If m1 has more than one interaction center, the else-clause is entered, where the current optimization takes place. The optimization consists of a loop-interchange of the for-loops over the entries of iV and the sites of m1 in lines 8 and 10, respectively. Due to the CoM condition, all sites see identical interaction partners and, hence, we can first gather data via the index vector and then loop over the sites of the molecule. Thus, we have traded expensive gather-loads and scatter-stores for cheaper broadcast-loads.

On IVB, due to the absence of gather- and scatter operations, the entire soa2 is traversed and unnecessary force-entries are masked away [6, 7]. The introduced loop-interchange does not improve performance there, as it trades in normal loads for broadcast-loads, which have similar costs.
In [6] a theoretical upper bound of 20 % of peak performance is given for IVB. On scenarios comparable to the ones presented here, values of 11 % \(-\) 16 % are observed [3, 6]. The MIC implementation further suffers from the costly packstore in line 4, and the expensive gather- and scatter operations, further reducing the attainable percentage of peak performance.
3.2 Shared-Memory Parallelization
Mode no3 : The first mode consists of simply not using the N3 optimization between linked-cells. It stems from other accelerator architectures, where resolving the race-condition was observed to be more costly than recomputing the respective forces [3]. Processing the cells, thus, becomes an embarrassingly parallel operation, cf. Fig. 2(a). In the present no3 implementation, the N3 optimization is employed when computing interactions within a single cell.
Approaches for shared memory parallelization of pairwise force evaluations (simplified illustration for 2D case). The yellow-colored cells can be processed in parallel. Red squares indicate “write-regions” for the lower-left yellow-colored cell, arrows indicate pairwise cell handlings for the yellow-color traversal. (a) no3 : All cells can be processed in parallel, but force contributions need to be computed twice, that is once for each molecule of a pairwise interaction. (b) c18, corresponding to 6 colors in 2D case. (c) c08, corresponding to 4 colors in 2D case.
Colored Modes c18 and c08 : Both of these approaches retain the N3 optimization by introducing a suitable coloring. The first one, c18, is based on the “standard” cell traversal order found in many implementations, including ls1-mardyn. After molecule interactions within a certain linked-cell have been computed, one iterates over the neighboring linked-cells, which have an index greater than the index of the current cell in a lexicographic ordering, and computes the pairwise forces. Coloring this traversal requires a stride of 2 in the “last” dimension and strides of 3 in all previous dimensions. For 3D simulations, this results in \(2\cdot 3^{3-1}=18\) colors and, respectively, synchronization stages, see Fig. 2(b).
In this work, we introduce the more compact cell-pair traversal mode, which was originally developed for strictly local operations on grids cells in a Lattice Boltzmann context c08 [10]. Instead of only computing force contributions for pairwise interactions with the yellow-colored cells, molecule pairs are also handled for certain diagonally neighbored linked-cell pairs, see Fig. 2(b). Strides of 2 in all dimensions suffice for this scheme, resulting in only \(2^3=8\) colors/sync. stages. This reduction further implies larger amount of work per color and, hence, allows the use of a larger number of threads.
In the current implementation, the indices of the individual colors have been precomputed at startup and are processed via a dynamic scheduling, to allow for potential load-balancing. We point out that both the c18 and c08 approaches exhibit a reduced computational intensity as compared to win : after a thread has processed a “patch” of cells (of size 18 for c18 or 8 for c08 ), it needs to load another patch, which almost always will be a fully disjoint set of SoAs. On the other hand, the win mode reuses SoAs already residing in cache for up to three consecutive “patch” processings.
3.3 MPI Parallelization
The win approach employs standard domain decomposition employing a Cartesian grid topology [2, 3]. Each process is extended by halo linked-cells which are filled by respective molecular information from the neighbored processes. Forces between molecules belonging to the process-local linked-cells and halo regions are computed on each respective process. As we consider only time spent in the force calculation, the MPI implementation used for reference has a somewhat unfair advantage, as we disregard time for communication in our analysis.
4 Results and Evaluation
4.1 Test Setup
We consider combinations of the following configurations: two system sizes with 0.16 million and 1.32 million molecules (0.16M and 1.32M denoted hereafter), two cutoff radii (\(r_c=3\sigma \) and \(r_c=5\sigma \)), and different numbers of Lennard-Jones interaction sites (“1CLJ” and “4CLJ” consisting of one and four centers, respectively), corresponding to the potential \(U_{LJ}(x_j - x_i):= 4\epsilon ((\frac{\sigma }{\Vert x_j-x_i\Vert })^{12} - (\frac{\sigma }{\Vert x_j - x_i\Vert })^6)\) and force contributions \(F_{ij}=-F_{ji}=-\nabla U_{LJ}(x_j-x_i)\) with parameters \(\sigma , \epsilon \) and positions \(x_i\), \(x_j\) of the interaction sites. The simulations were executed for 51 timesteps, each starting from a configuration of molecules initially arranged on a regular grid. The presented values are the mean values collected over four repetitions. A density of \(\rho \sigma ^3 = 0.3\) was used. The SuperMIC partition of the SuperMUC clusterFootnote 1, was used. The cluster nodes consist of a dual-socket eight-core Intel® IvyBridge host processor Xeon® E5-2650@2.6 GHz (IVB) with enabled hyperthreading and two Intel® Xeon PhiTM 5110p coprocessors with 60 cores@1.1GHz.
4.2 Single Core Performance
In order to set appropriate expectations for performance on both MIC and IVB, we first analyze single-core performance for the system size of 0.16M molecules. Since at least two threads per core are needed to fill the instruction pipeline [13], hyperthreading is expected to play a crucial role on the Xeon Phi. To enable a fair comparison, we also evaluate the hyperthreading performance on IVB.
In the hyperthreading analysis, we pin threads/processes to the same core and disregard any “classical” overhead of parallelization: threads running on the same core can be cheaply synchronized via the L1 cache, and, as they share physical resources, any load-imbalance among them automatically implies more physical resources for the thread with the higher computational load.
We also investigate the performance penalty of the “patched” access to the SoAs by considering a variant of the c08 mode, which accesses data in “colored” mode even when only a single thread is running (“col 1 thread” as opposed to “noncol 1 thread”).
Hyperthreading on Xeon Phi. The penalty of the colored access is deduced from measurements “noncol 1 thread” and “col 1 thread”, cf. Fig. 3. It is highest for 1CLJ rc3 and amounts up to 16 %, since this interaction is the least compute intensive and, hence, most memory intensive. As all other scenarios are more compute intensive, the impact is lower (3–5 %).
Next, we compare the measurements “1 thread” and “2 threads” in Fig. 3(b). The observed speed-ups range between 1.5 and 1.75, with the best speed-up obtained in the 1CLJ rc3 case and smaller speed-ups in the other scenarios. This behavior is due to bad pipelining properties of the LJ kernel and is in agreement with previous work [3]. Using 3 and 4 threads, further speed-ups are obtained. This is due to overlapping gather-scatter operations with FLOPs, and probably also the fact that the Xeon Phi cores feature an in-order execution mode. Consistent performance improvements are obtained and the 4-thread variant always delivers the highest FLOP-rate. This suggests that an optimal Xeon Phi implementation will employ the full 240 threads per MIC card. Oversubscription with 5 threads was also investigated, but led to degraded performance.
Hyperthreading on IvyBridge. Fig. 4 shows the respective analysis on IVB for c08 and win. Comparing “noncol 1 thread” and “col 1 thread” variants, we observe the same trend as on MIC, only this time it is weaker –7 % for 1CLJ rc3, 0–3% for the remaining interactions. This shows, that the colored scheme is also a good solution for hyperthreading on the IVB architecture. Next, we compare “col 1 thread” and “win 1 process”, and “col 2 threads” and “win 2 processes” in Fig. 4(a). A visible difference is observed for 1CLJ (35 % for 1CLJ rc3, 14 % for 1CLJ rc5) likely due to differing memory management. The 4CLJ scenarios are sufficiently computationally intensive to hide the differences in memory management (1.6 % and 0.4 % for 4CLJ with rc3 and rc5). Speed-ups due to hyperthreading in Fig. 4 arise as expected [4]. In our experiments, 40 % for 1CLJ rc3 and ca. 20 % in the other scenarios are obtained.
(a) Single core performance as a function of the number of interaction sites. On MIC the “ c08 col 4 threads” variant was used, while on IVB “ win 2 processes”. “Site” denotes the version before the loop-interchange, “CoM” - afterwards. (b) Final GFLOP/sec for 240/32 threads/processes, 1.32M molecules. Hatched bar indicates total speed-up of the c08 mode against reference version [7].
Absolute Performance. The dependence of the absolute performance from the number of interaction sites is shown in Fig. 6(a). On both MIC and IVB, 1CLJ is an exception from the trends. This is due to the fact, that the 1CLJ case spends the least amount of time inside the LJ Kernel, and, hence, runs closer to the higher FLOP-rate of the cutoff-evaluation, see Fig. 5. On MIC, the difference between 1CLJ and the other cases is also emphasized by the special treatment of the single-centered molecules in the if-statement of Listing 1.1.
On IVB, the performance is saturated at around 4–5 CLJ, as the performance of the LJ-Kernel is attained. This is around 14 % of the theoretical single-core peak performance, which is in agreement with previous results. Overall the values lie in the range of 9–14%.
The MIC implementation has become scalable in the number of interaction sites. The performance continually rises, reaching 9.3 % of single-core peak for 16CLJ, as the floating point performance becomes more and more saturated. Overall, the values range between 4 % and 8 %. In contrast, the version without the loop-interchange stagnates at around 5 % of peak-efficiency, which is likely the limit of the gather-scatter performance. In the 2–10 CLJ range, the gain continually increases from 1.1 to 1.7 with both implementations exhibiting a boost at 8CLJ and 4CLJ, as in those cases the gather- and scatter operations collect only contiguous data from one and two cache-lines, respectively. The gains at 4CLJ are 1.24 and 1.32 for the rc3 and rc5 cases, respectively. The gains for the no3 mode (not shown) were observed to be roughly half of those presented for c08, as the no3 mode does not need to scatter forces to the second SoA. Another factor, affecting the Xeon Phi performance, is the lack of addition and multiplication operations, which can be fused - only a total of 7 operations out of 43 can be fused: 2 in the cutoff-evaluation and 5 in the LJ-Kernel. This means that most of the time, the FPU is executing non-fused (i.e., either add or multiply) operations, which halves floating point performance. This effect is not as limiting on the IVB, due to the employed hyperthreading.
Assuming “perfect upscaling” by a factor of 60 (MIC) and 16 (IVB), we see that MIC has the potential to outperform IVB by a factor of up to 1.4–1.5.
4.3 Shared-Memory Parallelization
Figure 7 presents the performed strong scaling experiments. The non-colored versions of the c08 and c18 variants were used for the timings on 1 thread.
Comparing the c08, c18 and no3 schemes using 1 thread on MIC shows that restoring N3 gives a speed-up of 1.3 to 1.55. The scheme c08 performs around 5 %\(-\)10 % faster than c18, except in the 1.32M rc3 4CLJ case, where both schemes show similar performance. At 60 cores we observe a parallel efficiency of around 85 % for all rc3 scenarios except 0.16M rc3 1CLJ (75 %). For the rc5 scenarios, the values are between 90 and 94 % for all cases except 0.16M rc5 1CLJ (82 %). This demonstrates that our implementation and approach are very well suited for manycore architectures such as MIC. The speed-ups in the hyperthreading range are almost 2 for the 1.32M and 0.16M rc3 configurations which is in agreement with the results from Sect. 4.2. The 0.16M rc5 case, however, does not exhibit a sufficient level of parallelism due to the small number of linked-cells in the strong scaling limit.
Considering IVB performance we also see that the adapted “Flops are free” approach is indeed reasonable and the MPI implementation is hard to beat, despite its redundant calculations, and in agreement with [2]. Analyzing in more detail, we see that the source of the good win performance does not lie in scalability, but in superior sequential performance. The obtained speed-ups, relative to the base version of the same parallelization mode, are in all cases higher for the c08 variant, ranging between 16.9 and 18.4 versus values between 13.5 and 17.5 for win. Comparing the start of the curves in detail, the win can, however, be up to 10–20% faster than the colored variant, likely due to the better cache performance. Parallel efficiency on 16 cores is between 81 % in the 1CLJ 160k rc3 case and \(\ge \)89 % in the other cases for the c08 variant.
In the 1.32M scenarios (and even in some 0.16M scenarios), we attain speed-ups of comparable magnitude to the ones predicted in Sect. 4.2. Comparing IVB win and MIC c08 at full thread utilization, MIC is faster than IVB by factors 1.09 (1CLJ rc3), 1.23 (1CLJ rc5), 1.27 (4CLJ rc3), and 1.22 (4CLJ rc5). This is close to our simple upper bound (1.4–1.5) assuming perfect upscaling, cf. Sect. 4.2. These speed-ups are not consistently attained in the 0.16M case as this configuration exhibits around 8 times fewer linked-cells and, respectively, parallelism.
Figure 6(b) shows the final attained FLOP-rates and total speed-up compared to the reference ls1-mardyn version used in [7]. The 4CLJ scenarios benefit from both the gather-scatter and the N3 optimization, which multiplied together, give the observed speed-ups of 1.7–1.8. The 1CLJ scenarios experience only the N3 speed-up, explaining the factor of 1.4 for 1CLJ rc5. The 1CLJ rc3 scenario, being least computationally intensive, experienced a (considerably) larger speed-up than 1.4 due to the addressed bottleneck of the SoA reallocation, as mentioned in Sect. 3.
5 Summary
We presented an improvement to the gather-scatter-based molecular dynamics implementation and a shared-memory parallelization approach operating on the linked-cell structure of the molecular dynamics simulation.
As seen in Sect. 4.2, the number of expensive gather- and scatter-operations could be reduced by simple swapping of loops. This allowed to speed-up simulations of multi-centered molecules. The presented c08 scheme efficiently parallelizes the linked-cell algorithm in shared-memory environments with minimal synchronization steps and satisfying the N3 optimization. In particular, the Xeon Phi architecture could be exploited and delivered slightly more performance than the Ivy Bridge host. The accelerator card thus builds a valuable hardware extension for molecular dynamics. Hybrid MPI-OpenMP simulations using this scheme and extending the analysis to the whole simulation, as opposed to the presented calculation, are part of future work.
Optimization Notice: Software and workloads used in performance tests may have been optimized for performance only on Intel microprocessors. Performance tests, such as SYSmark and MobileMark, are measured using specific computer systems, components, software, operations and functions. Any change to any of those factors may cause the results to vary. You should consult other information and performance tests to assist you in fully evaluating your contemplated purchases, including the performance of that product when combined with other products. For more information go to http://www.intel.com/performance.
Intel, Xeon, and Intel Xeon Phi are trademarks of Intel Corporation in the U.S. and/or other countries.
References
Brown, W.M., Wang, P., Plimpton, S.J., Tharrington, A.N.: Implementing molecular dynamics on hybrid high performance computers short range forces. Comput. Phys. Commun. 182(4), 898–911 (2011)
Buchholz, M.: Framework zur Parallelisierung von Molekulardynamiksimulationen in verfahrenstechnischen Anwendungen. Dissertation, Institut für Informatik, Technische Universität München, München, August 2010
Eckhardt, W.: Efficient HPC implementations for large-scale molecular simulation in process engineering. Dissertation, Institut für Informatik, Technische Universität München, München, June 2014
Eckhardt, W., Heinecke, A., Bader, R., Brehm, M., Hammer, N., Huber, H., Kleinhenz, H.-G., Vrabec, J., Hasse, H., Horsch, M., Bernreuther, M., Glass, C.W., Niethammer, C., Bode, A., Bungartz, H.-J.: 591 TFLOPS multi-trillion particles simulation on SuperMUC. In: Kunkel, J.M., Ludwig, T., Meuer, H.W. (eds.) ISC 2013. LNCS, vol. 7905, pp. 1–12. Springer, Heidelberg (2013)
Griebel, M., Knapek, S., Zumbusch, G.: Numerical Simulation in Molecular Dynamics: Numerics, Algorithms, Parallelization, Applications, vol. 5, Springer, Heidelberg (2007)
Heinecke, A.: Boosting scientific computing applications through leveraging data parallel architectures. Dissertation, Institut für Informatik, Technische Universität München, München, January 2014
Heinecke, A., Eckhardt, W., Horsch, M., Bungartz, H.J.: Supercomputing for molecular-dynamics simulations: handling multi-trillion particles in nanofluidics. Springer Briefs in Computer Science, Springer, June 2015
Horsch, M., Niethammer, C., Vrabec, J., Hasse, H.: Computational molecular engineering as an emerging technology in process engineering. it-Information Technology Methoden und innovative Anwendungen der Informatik und Informationstechnik, 55(3), 97–101 (2013)
Kunaseth, M., Richards, D.F., et al.: Scalable data-privatization threading for hybrid MPI/OpenMP parallelization of molecular dynamics. In: International Conference on Parallel and Distributed Processing Techniques and Applications (2011)
Neumann, P., Bungartz, H.J., Mehl, M., Neckel, T., Weinzierl, T.: A coupled approach for fluid dynamic problems using the pde framework peano. Commun. Comput. Phys. 12, 65–84 (2012)
Niethammer, C., Becker, S., Bernreuther, M., Buchholz, M., Eckhardt, W., Heinecke, A., Werth, S., Bungartz, H.J., Glass, C.W., Hasse, H., Vrabec, J., Horsch, M.: ls1 mardyn: the massively parallel molecular dynamics code for large systems. J. Chem. Theor. Comput. 10(10), 4455–4464 (2014)
Pennycook, S., Hughes, C., Smelyanskiy, M., Jarvis, S.: Exploring SIMD for molecular dynamics, using intel xeon; processors and intel xeon phi coprocessors. In: 2013 IEEE 27th International Symposium on Parallel Distributed Processing (IPDPS), pp. 1085–1097, May 2013
Ranman, R.: Intel Xeon Phi Coprocessor Architecture and Tools. Apress open, New York (2013)
Acknowledgements
This work was partially supported by the Intel Parallel Computing Center ExScaMIC, and the Bundesministerium für Bildung in Forschung (Germany) in the scope of the project “SkaSim - Skalierbare HPC-Software für molekulare Simulationen in der chemischen Industrie”, Förderkennzeichen 01IH13005F. We further thank Wolfgang Hölzl for productive discussions.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer International Publishing Switzerland
About this paper
Cite this paper
Tchipev, N. et al. (2015). Optimized Force Calculation in Molecular Dynamics Simulations for the Intel Xeon Phi. In: Hunold, S., et al. Euro-Par 2015: Parallel Processing Workshops. Euro-Par 2015. Lecture Notes in Computer Science(), vol 9523. Springer, Cham. https://doi.org/10.1007/978-3-319-27308-2_62
Download citation
DOI: https://doi.org/10.1007/978-3-319-27308-2_62
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-27307-5
Online ISBN: 978-3-319-27308-2
eBook Packages: Computer ScienceComputer Science (R0)