Abstract
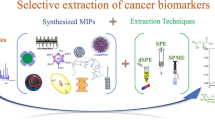
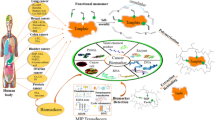
Breast cancer is threatening women all over the world. Invasive and noninvasive breast cancers have increased with time. According to 2021, 7.8 million women have been living with breast cancer, so that is making it the most prevalent cancer. Many women have no symptoms. Over time, in situ (stage 0) cancers can surround tissue, spread to lymph nodes, and continue to the other organs in the body (metastasis stage). Hence, the early diagnosis of breast cancer is so important. Breast cancer has been detected with mammograms, breast ultrasounds, and breast magnetic resonance imaging. For early detection, new sensitive, selective, invasive, reliable, and fast equipment or techniques are needed. At this point, molecularly imprinted polymers could be successful by electrochemical techniques. Molecularly imprinted polymers have selectivity features so that they can combine electrochemistry performance and give a successful output like the point of care equipment or lab-on-a-chip. In this chapter, recent developments in breast cancer detection by molecularly imprinted polymer-based electrochemical sensors are discussed. This chapter will be helpful to scientists, students, and industry in the development of new equipment, devices, and techniques for the detection of breast cancer markers.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Cancer. https://www.who.int/health-topics/cancer#tab=tab_1. Accessed 3 Apr 2022.
Asia S, Asia S, Hdi H. Source: Globocan. 2020;419:1–2.
Ruddy KJ, Winer EP. Male breast cancer: risk factors, biology, diagnosis, treatment, and survivorship. Ann Oncol. 2013;24:1434–43. https://doi.org/10.1093/ANNONC/MDT025.
Parada H, Sun X, Tse CK, et al. Lifestyle patterns and survival following breast cancer in the Carolina breast cancer study. Epidemiology. 2019;30:83–92. https://doi.org/10.1097/EDE.0000000000000933.
Board PS and PE. Breast cancer screening (PDQ®). PDQ Cancer Inf Summ. 2021:22–4. https://www.cancer.gov/types/breast/hp/breast-screening-pdq.
Gucalp A, Traina TA, Eisner JR, et al. Male breast cancer: a disease distinct from female breast cancer. Breast Cancer Res Treat. 2019;173:37–48. https://doi.org/10.1007/S10549-018-4921-9.
Clark BZ, Onisko A, Assylbekova B, et al. Breast cancer global tumor biomarkers: a quality assurance study of intratumoral heterogeneity. Mod Pathol. 2019;32:354–66. https://doi.org/10.1038/S41379-018-0153-0.
Narod SA. Personalised medicine and population health: breast and ovarian cancer. Hum Genet. 2018;137:769–78. https://doi.org/10.1007/S00439-018-1944-6.
Mahvi DA, Liu R, Grinstaff MW, et al. Local cancer recurrence: the realities, challenges, and opportunities for new therapies. CA Cancer J Clin. 2018;68:488–505. https://doi.org/10.3322/CAAC.21498.
McCormack V, McKenzie F, Foerster M, et al. Breast cancer survival and survival gap apportionment in sub-Saharan Africa (ABC-DO): a prospective cohort study. Lancet Glob Health. 2020;8:e1203–12. https://doi.org/10.1016/S2214-109X(20)30261-8.
Wang F, Fang Q, Ge Z, et al. Common BRCA1 and BRCA2 mutations in breast cancer families: a meta-analysis from a systematic review. Mol Biol Rep. 2012;39:2109–18. https://doi.org/10.1007/S11033-011-0958-0.
Piletska EV, Guerreiro AR, Whitcombe MJ, Piletsky SA. Influence of the polymerization conditions on the performance of molecularly imprinted polymers. Macromolecules. 2009;42:4921–8. https://doi.org/10.1021/MA900432Z/ASSET/IMAGES/MA900432Z.SOCIAL.JPEG_V03.
Poma A, Turner APF, Piletsky SA. Advances in the manufacture of MIP nanoparticles. Trends Biotechnol. 2010;28:629–37. https://doi.org/10.1016/J.TIBTECH.2010.08.006.
Morelli I, Chiono V, Vozzi G, et al. Molecularly imprinted submicron spheres for applications in a novel model biosensor-film. Sensors Actuators B Chem. 2010;150:394–401. https://doi.org/10.1016/J.SNB.2010.06.046.
Scorrano S, Mergola L, del Sole R, Vasapollo G. Synthesis of molecularly imprinted polymers for amino acid derivates by using different functional monomers. Int J Mol Sci. 2011;12:1735. https://doi.org/10.3390/IJMS12031735.
Puoci F, Cirillo G, Curcio M, et al. Molecularly imprinted solid-phase extraction for the selective HPLC determination of alpha-tocopherol in bay leaves. Anal Chim Acta. 2007;593:164–70. https://doi.org/10.1016/J.ACA.2007.04.053.
Baggiani C, Anfossi L, Giovannoli C. Solid-phase extraction of food contaminants using molecularly imprinted polymers. Anal Chim Acta. 2007;591:29–39. https://doi.org/10.1016/J.ACA.2007.01.056.
Ramström O, Mosbach K. Synthesis and catalysis by molecularly imprinted materials. Curr Opin Chem Biol. 1999;3:759–64. https://doi.org/10.1016/S1367-5931(99)00037-X.
Suwanwong Y, Boonpangrak S. Molecularly imprinted polymers for the extraction and determination of water-soluble vitamins: a review from 2001 to 2020. Eur Polym J. 2021;161:110835. https://doi.org/10.1016/J.EURPOLYMJ.2021.110835.
Vasapollo G, Del SR, Mergola L, et al. Molecularly imprinted polymers: present and future prospective. OPEN ACCESS Int J Mol Sci. 2011;12:12. https://doi.org/10.3390/ijms12095908.
Pichon V, Chapuis-Hugon F. Role of molecularly imprinted polymers for selective determination of environmental pollutants–a review. Anal Chim Acta. 2008;622:48–61. https://doi.org/10.1016/J.ACA.2008.05.057.
Andersson LI. Molecular imprinting: developments and applications in the analytical chemistry field. J Chromatogr B Biomed Sci Appl. 2000;745:3–13. https://doi.org/10.1016/S0378-4347(00)00135-3.
Wei S, Mizaikoff B. Recent advances on noncovalent molecular imprints for affinity separations. J Sep Sci. 2007;30:1794–805. https://doi.org/10.1002/JSSC.200700166.
Tamayo FG, Turiel E, Martín-Esteban A. Molecularly imprinted polymers for solid-phase extraction and solid-phase microextraction: recent developments and future trends. J Chromatogr A. 2007;1152:32–40. https://doi.org/10.1016/J.CHROMA.2006.08.095.
Haginaka J. Monodispersed, molecularly imprinted polymers as affinity-based chromatography media. J Chromatogr B Anal Technol Biomed Life Sci. 2008;866:3–13. https://doi.org/10.1016/J.JCHROMB.2007.07.019.
Lasáková M, Jandera P. Molecularly imprinted polymers and their application in solid-phase extraction. J Sep Sci. 2009;32:799–812. https://doi.org/10.1002/JSSC.200800506.
Campuzano S, Pedrero M, Pingarrón JM. Non-invasive breast cancer diagnosis through electrochemical biosensing at different molecular levels. Sensors. 2017;17:1993. https://doi.org/10.3390/S17091993.
Miao P, Wang B, Yu Z, et al. Ultrasensitive electrochemical detection of microRNA with star trigon structure and endonuclease mediated signal amplification. Biosens Bioelectron. 2015;63:365–70. https://doi.org/10.1016/J.BIOS.2014.07.075.
Cardoso AR, Moreira FTC, Fernandes R, Sales MGF. Novel and simple electrochemical biosensor monitoring attomolar levels of miRNA-155 in breast cancer. Biosens Bioelectron. 2016;80:621–30. https://doi.org/10.1016/J.BIOS.2016.02.035.
Moscovici M, Bhimji A, Kelley SO. Rapid and specific electrochemical detection of prostate cancer cells using an aperture sensor array. Lab Chip. 2013;13:940–6. https://doi.org/10.1039/C2LC41049D.
Zhu Y, Chandra P, Shim Y-B. Ultrasensitive and selective electrochemical diagnosis of breast cancer based on a hydrazine–Au nanoparticle–aptamer bioconjugate. Anal Chem. 2013;85:1058–64. https://doi.org/10.1021/ac302923k.
Azimzadeh M, Rahaie M, Nasirizadeh N, et al. An electrochemical nanobiosensor for plasma miRNA-155, based on graphene oxide and gold nanorod, for early detection of breast cancer. Biosens Bioelectron. 2016;77:99–106. https://doi.org/10.1016/J.BIOS.2015.09.020.
Chen M, Wang Y, Su H, et al. Three-dimensional electrochemical DNA biosensor based on 3D graphene-Ag nanoparticles for sensitive detection of CYFRA21-1 in non-small cell lung cancer. Sensors Actuators B Chem. 2018;255:2910–8. https://doi.org/10.1016/j.snb.2017.09.111.
Wang K, He MQ, Zhai FH, et al. A novel electrochemical biosensor based on polyadenine modified aptamer for label-free and ultrasensitive detection of human breast cancer cells. Talanta. 2017;166:87–92. https://doi.org/10.1016/J.TALANTA.2017.01.052.
Fuller MS, Lee CI, Elmore JG. Breast cancer screening: an evidence-based update. Med Clin North Am. 2015;99:451–68. https://doi.org/10.1016/J.MCNA.2015.01.002.
Nazário ACP, Facina G, Filassi JR. Breast cancer: news in diagnosis and treatment. Rev Assoc Med Bras. 2015;61:543–52. https://doi.org/10.1590/1806-9282.61.06.543.
Pace LE, Keating NL. A systematic assessment of benefits and risks to guide breast cancer screening decisions. JAMA. 2014;311:1327–35. https://doi.org/10.1001/JAMA.2014.1398.
Friedewald SM, Rafferty EA, Rose SL, et al. Breast cancer screening using tomosynthesis in combination with digital mammography. JAMA. 2014;311:2499–507. https://doi.org/10.1001/JAMA.2014.6095.
Svahn TM, Houssami N, Sechopoulos I, Mattsson S. Review of radiation dose estimates in digital breast tomosynthesis relative to those in two-view full-field digital mammography. Breast. 2015;24:93–9. https://doi.org/10.1016/J.BREAST.2014.12.002.
Brem RF, Lenihan MJ, Lieberman J, Torrente J. Screening breast ultrasound: past, present, and future. AJR Am J Roentgenol. 2015;204:234–40. https://doi.org/10.2214/AJR.13.12072.
Lehman CD, Gatsonis C, Kuhl CK, et al. MRI evaluation of the contralateral breast in women with recently diagnosed breast cancer. N Engl J Med. 2007;356:1295–303. https://doi.org/10.1056/NEJMOA065447.
Houssami N, Turner R, Macaskill P, et al. An individual person data meta-analysis of preoperative magnetic resonance imaging and breast cancer recurrence. J Clin Oncol. 2014;32:392–401. https://doi.org/10.1200/JCO.2013.52.7515.
Berg WA, Blume JD, Cormack JB, et al. Combined screening with ultrasound and mammography vs mammography alone in women at elevated risk of breast cancer. JAMA. 2008;299:2151–63. https://doi.org/10.1001/JAMA.299.18.2151.
Liberman L. Centennial dissertation. Percutaneous imaging-guided core breast biopsy: state of the art at the millennium. AJR Am J Roentgenol. 2000;174:1191–9. https://doi.org/10.2214/AJR.174.5.1741191.
Villegas-Carlos F, Andino-Araque V, Valverde-Quintana M, et al. Predictive factors of invasion in ductal carcinoma in situ diagnosed by core-needle biopsy. Cir Cir. 2022;90:41–9. https://doi.org/10.24875/CIRU.21000136.
Bennett NC, Farah CS. Next-generation sequencing in clinical oncology: next steps towards clinical validation. Cancers (Basel). 2014;6:2296–312. https://doi.org/10.3390/CANCERS6042296.
Perou CM, Sørile T, Eisen MB, et al. Molecular portraits of human breast tumors. Nature. 2000;406:747–52. https://doi.org/10.1038/35021093.
Ross JS, Linette GP, Stec J, et al. Breast cancer biomarkers and molecular medicine. Expert Rev Mol Diagn. 2003;3:573–85. https://doi.org/10.1586/14737159.3.5.573.
Dowsett M, Dunbier AK. Emerging biomarkers and new understanding of traditional markers in personalized therapy for breast cancer. Clin Cancer Res. 2008;14:8019–26. https://doi.org/10.1158/1078-0432.CCR-08-0974.
Lim E, Metzger-Filho O, Winer EP. The natural history of hormone receptor-positive breast cancer. Oncology (Williston Park). 2012;26:688–95.
Pan H, Gray R, Braybrooke J, et al. 20-year risks of breast-cancer recurrence after stopping endocrine therapy at 5 years. N Engl J Med. 2017;377:1836–46. https://doi.org/10.1056/NEJMOA1701830.
Patani N, Martin LA, Dowsett M. Biomarkers for the clinical management of breast cancer: international perspective. Int J Cancer. 2013;133:1–13. https://doi.org/10.1002/IJC.27997.
Mohammed H, Russell IA, Stark R, et al. Progesterone receptor modulates ERα action in breast cancer. Nature. 2015;523:313–7. https://doi.org/10.1038/NATURE14583.
Rubin I, Yarden Y. The basic biology of HER2. Ann Oncol Off J Eur Soc Med Oncol. 2001;12 Suppl 1:S3–8. https://doi.org/10.1093/ANNONC/12.SUPPL_1.S3.
Ross JS, Slodkowska EA, Symmans WF, et al. The HER-2 receptor and breast cancer: ten years of targeted anti-HER-2 therapy and personalized medicine. Oncologist. 2009;14:320–68. https://doi.org/10.1634/THEONCOLOGIST.2008-0230.
Luporsi E, André F, Spyratos F, et al. Ki-67: level of evidence and methodological considerations for its role in the clinical management of breast cancer: analytical and critical review. Breast Cancer Res Treat. 2012;132:895–915. https://doi.org/10.1007/S10549-011-1837-Z.
Ding Y, Ding K, Qian H, et al. Impact on survival of estrogen receptor, progesterone receptor, and Ki-67 expression discordance pre- and post-neoadjuvant chemotherapy in breast cancer. PLoS One. 2020;15:e0231895. https://doi.org/10.1371/JOURNAL.PONE.0231895.
Huang SH, O’Sullivan B. Overview of the 8th edition TNM classification for head and neck cancer. Curr Treat Options in Oncol. 2017;18:1–13. https://doi.org/10.1007/S11864-017-0484-Y.
Fischer JP, Wes AM, Tuggle CT, et al. Mastectomy with or without immediate implant reconstruction has similar 30-day perioperative outcomes. J Plast Reconstr Aesthet Surg. 2014;67:1515–22. https://doi.org/10.1016/J.BJPS.2014.07.021.
Veronesi U, Saccozzi R, Del Vecchio M, et al. Comparing radical mastectomy with quadrantectomy, axillary dissection, and radiotherapy in patients with small cancers of the breast. N Engl J Med. 1981;305:6–11. https://doi.org/10.1056/NEJM198107023050102.
Giuliano AE, Kirgan DM, Guenther JM, Morton DL. Lymphatic mapping and sentinel lymphadenectomy for breast cancer. Ann Surg. 1994;220:391–401. https://doi.org/10.1097/00000658-199409000-00015.
Wang X, Xu L, Yin Z, et al. Locoregional recurrence-associated factors and risk-adapted postmastectomy radiotherapy for breast cancer staged in cT1-2N0-1 after neoadjuvant chemotherapy. Cancer Manag Res. 2018;10:4105–12. https://doi.org/10.2147/CMAR.S173628.
Tang L, Matsushita H, Jingu K. Controversial issues in radiotherapy after breast-conserving surgery for early breast cancer in older patients: a systematic review. J Radiat Res. 2018;59:789–93. https://doi.org/10.1093/JRR/RRY071.
Turaka A, Freedman GM, Li T, et al. Young age is not associated with increased local recurrence for DCIS treated by breast-conserving surgery and radiation. J Surg Oncol. 2009;100:25–31. https://doi.org/10.1002/JSO.21284.
Ng SP, David S, Alamgeer M, Ganju V. Impact of pretreatment combined (18)F-fluorodeoxyglucose positron emission tomography/computed tomography staging on radiation therapy treatment decisions in locally advanced breast cancer. Int J Radiat Oncol Biol Phys. 2015;93:111–7. https://doi.org/10.1016/J.IJROBP.2015.05.012.
Haviland JS, Owen JR, Dewar JA, et al. The UK Standardisation of Breast Radiotherapy (START) trials of radiotherapy hypofractionation for treatment of early breast cancer: 10-year follow-up results of two randomized controlled trials. Lancet Oncol. 2013;14:1086–94. https://doi.org/10.1016/S1470-2045(13)70386-3.
Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004;351:2817–26. https://doi.org/10.1056/NEJMOA041588.
Wu YT, Xu Z, Zhang K, et al. Efficacy and cardiac safety of the concurrent use of trastuzumab and anthracycline-based neoadjuvant chemotherapy for HER2-positive breast cancer: a systematic review and meta-analysis. Ther Clin Risk Manag. 2018;14:1789–97. https://doi.org/10.2147/TCRM.S176214.
Tolaney SM, Barry WT, Dang CT, et al. Adjuvant paclitaxel and trastuzumab for node-negative, HER2-positive breast cancer. N Engl J Med. 2015;372:134–41. https://doi.org/10.1056/NEJMOA1406281.
Mavroudis D, Saloustros E, Malamos N, et al. Six versus 12 months of adjuvant trastuzumab in combination with dose-dense chemotherapy for women with HER2-positive breast cancer: a multicenter randomized study by the Hellenic Oncology Research Group (HORG). Ann Oncol Off J Eur Soc Med Oncol. 2015;26:1333–40. https://doi.org/10.1093/ANNONC/MDV213.
Pagani O, Regan MM, Walley BA, et al. Adjuvant exemestane with ovarian suppression in premenopausal breast cancer. N Engl J Med. 2014;371:107–18. https://doi.org/10.1056/NEJMOA1404037.
Davies C, Pan H, Godwin J, et al. Long-term effects of continuing adjuvant tamoxifen to 10 years versus stopping at 5 years after diagnosis of estrogen receptor-positive breast cancer: ATLAS, a randomized trial. Lancet (London, England). 2013;381:805–16. https://doi.org/10.1016/S0140-6736(12)61963-1.
Leal F, Liutti VT, Antunes dos Santos VC, et al. Neoadjuvant endocrine therapy for resectable breast cancer: a systematic review and meta-analysis. Breast. 2015;24:406–12. https://doi.org/10.1016/J.BREAST.2015.03.004.
Finn RS, Crown JP, Lang I, et al. The cyclin-dependent kinase 4/6 inhibitor palbociclib in combination with letrozole versus letrozole alone as first-line treatment of estrogen receptor-positive, HER2-negative, advanced breast cancer (PALOMA-1/TRIO-18): a randomized phase 2 study. Lancet Oncol. 2015;16:25–35. https://doi.org/10.1016/S1470-2045(14)71159-3.
Turner NC, Ro J, André F, et al. Palbociclib in hormone-receptor-positive advanced breast cancer. N Engl J Med. 2015;373:209–19. https://doi.org/10.1056/NEJMOA1505270.
Swain SM, Baselga J, Kim S-B, et al. Pertuzumab, trastuzumab, and docetaxel in HER2-positive metastatic breast cancer. N Engl J Med. 2015;372:724–34. https://doi.org/10.1056/NEJMOA1413513.
Kilpatrick ES, Lind MJ. Appropriate requesting of serum tumor markers. BMJ. 2009;339:859. https://doi.org/10.1136/BMJ.B3111.
Shah R, Rosso K, David Nathanson S. Pathogenesis, prevention, diagnosis and treatment of breast cancer. World J Clin Oncol. 2014;5:283–98. https://doi.org/10.5306/WJCO.V5.I3.283.
Banegas MP, Bird Y, Moraros J, et al. Breast cancer knowledge, attitudes, and early detection practices in United States-Mexico border Latinas. J Women’s Health (Larchmt). 2012;21:101–7. https://doi.org/10.1089/JWH.2010.2638.
Donepudi MS, Kondapalli K, Amos SJ, Venkanteshan P. Breast cancer statistics and markers. J Cancer Res Ther. 2014;10:506–11. https://doi.org/10.4103/0973-1482.137927.
Duffy MJ, Evoy D, McDermott EW. CA 15-3: uses and limitation as a biomarker for breast cancer. Clin Chim Acta. 2010;411:1869–74. https://doi.org/10.1016/J.CCA.2010.08.039.
Kurian S, Khan M, Grant M. CA 27-29 in patients with breast cancer with pulmonary fibrosis. Clin Breast Cancer. 2008;8:538–40. https://doi.org/10.3816/CBC.2008.N.067.
Zheng G, Yu H, Hemminki A, et al. Familial associations of female breast cancer with other cancers. Int J Cancer. 2017;141:2253–9. https://doi.org/10.1002/IJC.30927.
Couto E, Hemminki K. Estimates of heritable and environmental components of familial breast cancer using family history information. Br J Cancer. 2007;96:1740–2. https://doi.org/10.1038/SJ.BJC.6603753.
Claus EB, Schildkraut JM, Thompson WD, Risch NJ. The genetic attributable risk of breast and ovarian cancer. Cancer. 1996;77:2318–24. https://doi.org/10.1002/(sici)1097-0142(19960601)77:11<2318::aid-cncr21>3.0.co;2-z.
Apostolou P, Fostira F. Hereditary breast cancer: the era of new susceptibility genes. Biomed Res Int. 2013;2013:747318. https://doi.org/10.1155/2013/747318.
Nielsen FC, Van Overeem HT, Sørensen CS. Hereditary breast and ovarian cancer: new genes in confined pathways. Nat Rev Cancer. 2016;16:599–612. https://doi.org/10.1038/NRC.2016.72.
Mavaddat N, Barrowdale D, Andrulis IL, et al. Pathology of breast and ovarian cancers among BRCA1 and BRCA2 mutation carriers: results from the Consortium of Investigators of Modifiers of BRCA1/2 (CIMBA). Cancer Epidemiol Biomark Prev. 2012;21:134–47. https://doi.org/10.1158/1055-9965.EPI-11-0775.
Vocka M, Zimovjanova M, Bielcikova Z, et al. Estrogen receptor status oppositely modifies breast cancer prognosis in BRCA1/BRCA2 mutation carriers versus non-carriers. Cancers (Basel). 2019;11. https://doi.org/10.3390/CANCERS11060738.
Yadav S, Couch FJ. Germline genetic testing for breast cancer risk: the past, present, and future. Am Soc Clin Oncol Educ Book Am Soc Clin Oncol Annu Meet. 2019;39:61–74. https://doi.org/10.1200/EDBK_238987.
Gilkes DM, Semenza GL. Role of hypoxia-inducible factors in breast cancer metastasis. Future Oncol. 2013;9:1623–36. https://doi.org/10.2217/FON.13.92.
Salceda S, Caro J. Hypoxia-inducible factor 1alpha (HIF-1alpha) protein is rapidly degraded by the ubiquitin-proteasome system under normoxic conditions. Its stabilization by hypoxia depends on redox-induced changes. J Biol Chem. 1997;272:22642–7. https://doi.org/10.1074/JBC.272.36.22642.
Masoumi Moghaddam S, Amini A, Morris DL, Pourgholami MH. Significance of vascular endothelial growth factor in growth and peritoneal dissemination of ovarian cancer. Cancer Metastasis Rev. 2012;31:143–62. https://doi.org/10.1007/S10555-011-9337-5.
Zhang L, Kirchhoff T, Yee CJ, Offit K. A rapid and reliable test for BRCA1 and BRCA2 founder mutation analysis in paraffin tissue using pyrosequencing. J Mol Diagn. 2009;11:176–81. https://doi.org/10.2353/JMOLDX.2009.080137.
Meisel JL, Hyman DM, Garg K, et al. The performance of BRCA1 immunohistochemistry for detecting germline, somatic, and epigenetic BRCA1 loss in high-grade serous ovarian cancer. Ann Oncol Off J Eur Soc Med Oncol. 2014;25:2372–8. https://doi.org/10.1093/ANNONC/MDU461.
Garg K, Levine DA, Olvera N, et al. BRCA1 immunohistochemistry in a molecularly characterized cohort of ovarian high-grade serous carcinomas. Am J Surg Pathol. 2013;37:138–46. https://doi.org/10.1097/PAS.0B013E31826CABBD.
Smith KL, Isaacs C. BRCA mutation testing in determining breast cancer therapy. Cancer J. 2011;17:492–9. https://doi.org/10.1097/PPO.0B013E318238F579.
You M, Yang S, Tang W, et al. Molecularly imprinted polymers-based electrochemical DNA biosensor for the determination of BRCA-1 amplified by SiO 2@Ag. Biosens Bioelectron. 2018;112:72–8. https://doi.org/10.1016/J.BIOS.2018.04.038.
Ali IU, Campbell G, Lidereau R, Callahan R. Amplification of c-erbB-2 and aggressive human breast tumors? Science. 1988;240:1795–6. https://doi.org/10.1126/SCIENCE.3289120.
Garnock-Jones KP, Keating GM, Scott LJ. Trastuzumab: a review of its use as adjuvant treatment in human epidermal growth factor receptor 2 (HER2)-positive early breast cancer. Drugs. 2010;70:215–39. https://doi.org/10.2165/11203700-000000000-00000.
Wolff AC, Hammond MEH, Hicks DG, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. Arch Pathol Lab Med. 2014;138:241–56. https://doi.org/10.5858/ARPA.2013-0953-SA.
Wolff AC, Hammond MEH, Schwartz JN, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. Arch Pathol Lab Med. 2007;131:18–43. https://doi.org/10.5858/2007-131-18-ASOCCO.
Pacheco JG, Rebelo P, Freitas M, et al. Breast cancer biomarker (HER2-ECD) detection using a molecularly imprinted electrochemical sensor. Sensors Actuators B Chem. 2018;273:1008–14. https://doi.org/10.1016/J.SNB.2018.06.113.
Amayo AA, Kuria JG. Clinical application of tumor markers: a review. East Afr Med J. 2009;86. https://doi.org/10.4314/EAMJ.V86I12.62909.
Pacheco JG, Silva MSV, Freitas M, et al. Molecularly imprinted electrochemical sensor for the point-of-care detection of a breast cancer biomarker (CA 15-3). Sensors Actuators B Chem. 2018;256:905–12. https://doi.org/10.1016/J.SNB.2017.10.027.
Ribeiro JA, Pereira CM, Silva AF, Sales MGF. Disposable electrochemical detection of breast cancer tumour marker CA 15-3 using poly(Toluidine Blue) as imprinted polymer receptor. Biosens Bioelectron. 2018;109:246–54. https://doi.org/10.1016/J.BIOS.2018.03.011.
Gomes RS, Moreira FTC, Fernandes R, Goreti Sales MF. Sensing CA 15-3 in point-of-care by electropolymerizing O-phenylenediamine (oPDA) on Au-screen printed electrodes. PLoS One. 2018;13:e0196656. https://doi.org/10.1371/JOURNAL.PONE.0196656.
Santos ART, Moreira FTC, Helguero LA, Sales MGF. Antibody biomimetic material made of pyrrole for CA 15-3 and its application as sensing material in ion-selective electrodes for potentiometric detection. Biosensors. 2018;8:8. https://doi.org/10.3390/BIOS8010008.
Hammarström S. The carcinoembryonic antigen (CEA) family: structures, suggested functions, and expression in normal and malignant tissues. Semin Cancer Biol. 1999;9:67–81. https://doi.org/10.1006/SCBI.1998.0119.
Grunnet M, Sorensen JB. Carcinoembryonic antigen (CEA) as a tumor marker in lung cancer. Lung Cancer. 2012;76:138–43. https://doi.org/10.1016/J.LUNGCAN.2011.11.012.
Wu S g, He Z y, Zhou J, et al. Serum levels of CEA and CA15-3 in different molecular subtypes and prognostic value in Chinese breast cancer. Breast. 2014;23:88–93. https://doi.org/10.1016/J.BREAST.2013.11.003.
Park BW, Oh JW, Kim JH, et al. Preoperative CA 15-3 and CEA serum levels as predictor for breast cancer outcomes. Ann Oncol Off J Eur Soc Med Oncol. 2008;19:675–81. https://doi.org/10.1093/ANNONC/MDM538.
Shao Y, Sun X, He Y, et al. Elevated levels of serum tumor markers CEA and CA15-3 are prognostic parameters for different molecular subtypes of breast cancer. PLoS One. 2015;10:e0133830. https://doi.org/10.1371/JOURNAL.PONE.0133830.
Bahrami-Ahmadi A, Makarian F, Mortazavizadeh MR, et al. Symptomatic metastasis prediction with serial measurements of CA 15.3 in primary breast cancer patients. J Res Med Sci. 2012;17:850.
Asad-Ur-Rahman F, Saif MW. Elevated level of serum carcinoembryonic antigen (CEA) and search for a malignancy: a case report. Cureus. 2016;8:e648. https://doi.org/10.7759/CUREUS.648.
Lee JS, Park S, Park JM, et al. Elevated levels of preoperative CA 15-3 and CEA serum levels have independently poor prognostic significance in breast cancer. Ann Oncol Off J Eur Soc Med Oncol. 2013;24:1225–31. https://doi.org/10.1093/ANNONC/MDS604.
Puglisi F, Fontanella C, Numico G, et al. Follow-up of patients with early breast cancer: is it time to rewrite the story? Crit Rev Oncol Hematol. 2014;91:130–41. https://doi.org/10.1016/J.CRITREVONC.2014.03.001.
Li X, Dai D, Chen B, et al. Clinicopathological and prognostic significance of cancer antigen 15-3 and carcinoembryonic antigen in breast cancer: a meta-analysis including 12,993 patients. Dis Markers. 2018;2018:1–15. https://doi.org/10.1155/2018/9863092.
Hing JX, Mok CW, Tan PT, et al. Clinical utility of tumor marker velocity of cancer antigen 15-3 (CA 15-3) and carcinoembryonic antigen (CEA) in breast cancer surveillance. Breast. 2020;52:95–101. https://doi.org/10.1016/J.BREAST.2020.05.005.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Bakirhan, N.K., Yucel, C. (2022). Recent Progress in Detection of Breast Cancer Biomarkers by Clinical and Imprinting Polymer-Based Sensors. In: Chaughule, R.S., Patkar, D.P., Ramanujan, R.V. (eds) Nanomaterials for Cancer Detection Using Imaging Techniques and Their Clinical Applications. Springer, Cham. https://doi.org/10.1007/978-3-031-09636-5_11
Download citation
DOI: https://doi.org/10.1007/978-3-031-09636-5_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-09635-8
Online ISBN: 978-3-031-09636-5
eBook Packages: Physics and AstronomyPhysics and Astronomy (R0)