Abstract
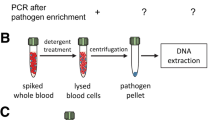
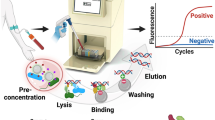
Automation in DNA isolation is a necessity for routine practice employing molecular diagnosis of infectious agents. To this end, the development of automated systems for the molecular diagnosis of microorganisms directly in blood samples is at its beginning. Important characteristics of systems demanded for routine use include high recovery of microbial DNA, DNA-free containment for the reduction of DNA contamination from exogenous sources, DNA-free reagents and consumables, ideally a walkaway system, and economical pricing of the equipment and consumables. Such full automation of DNA extraction evaluated and in use for sepsis diagnostics is yet not available. Here, we present protocols for the semiautomated isolation of microbial DNA from blood culture and low- and high-volume blood samples. The protocols include a manual pretreatment step followed by automated extraction and purification of microbial DNA.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Podnecky NL, Elrod MG, Newton BR et al (2013) Comparison of DNA extraction kits for detection of Burkholderia pseudomallei in spiked human whole blood using real-time PCR. PLoS One 8:58032. doi:10.1371/journal.pone.0058032
Hansen WLJ, Bruggeman CA, Wolffs PFG (2009) Evaluation of new preanalysis sample treatment tools and DNA isolation protocols to improve bacterial pathogen detection in whole blood. J Clin Microbiol 47:2629–2631
Loonen AJM, Jansz AR, Kreeftenberg H et al (2011) Acceleration of the direct identification of Staphylococcus aureus versus coagulase-negative staphylococci from blood culture material: a comparison of six bacterial DNA extraction methods. Eur J Clin Microbiol Infect Dis 30:337–342
Wiesinger-Mayr H, Jordana-Lluch E, Martró E et al (2011) Establishment of a semi-automated pathogen DNA isolation from whole blood and comparison with commercially available kits. J Microbiol Methods 85:206–213
Laakso S, Mäki M (2013) Assessment of a semi-automated protocol for multiplex analysis of sepsis-causing bacteria with spiked whole blood samples. Microbiologyopen 2:284–292
Hansen WLJ, Bruggeman CA, Wolffs PFG (2013) Pre-analytical sample treatment and DNA extraction protocols for the detection of bacterial pathogens from whole blood. Methods Mol Biol 943:81–90
Loonen AJM, Bos MP, van Meerbergen B et al (2013) Comparison of pathogen DNA isolation methods from large volumes of whole blood to improve molecular diagnosis of bloodstream infections. PLoS One 8:e72349. doi:10.1371/journal.pone.0072349
Kellogg JA, Manzella JP, Bankert DA (2000) Frequency of low-level bacteremia in children from birth to fifteen years of age. J Clin Microbiol 38:2181–2185
Yagupsky P, Nolte FS (1990) Quantitative aspects of septicemia. Clin Microbiol Rev 3: 269–279
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2015 Springer Science+Business Media New York
About this protocol
Cite this protocol
Lorenz, M.G., Disqué, C., Mühl, H. (2015). Bacterial and Fungal DNA Extraction from Blood Samples: Automated Protocols. In: Mancini, N. (eds) Sepsis. Methods in Molecular Biology, vol 1237. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-1776-1_12
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1776-1_12
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-1775-4
Online ISBN: 978-1-4939-1776-1
eBook Packages: Springer Protocols