Abstract
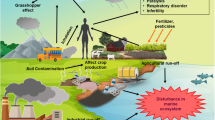
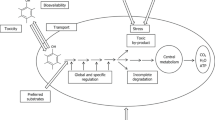
The biotransformation of synthetic chemicals that enter the environment is dependent on the capacity of microbial enzymes to recognize xenobiotic substrates and catalyze reactions with stable structural elements, such as carbon-halogen bonds. The range of compounds that can be enzymatically degraded strongly influences the environmental fate of many potentially harmful compounds. Environmental recalcitrance thus is, in part, a problem of enzyme activity and specificity. This can be well illustrated with the halogenated aliphatics, which are frequently encountered as environmental pollutants due to losses and emissions during their use in industry and agriculture. Halogenated aliphatic compounds are used as blowing agents (methylchloride), cooling liquids (ethylchloride), soil fumigants (1,3-dichloropropylene, methylbromide), insecticides (hexachlorocyclohexane), intermediates in chemical synthesis (1,2-dichloroethane, vinyl chloride, chloroacetates) and as solvents (trichloroethanes, tri- and tetrachloroethene). Several related structures (e.g., chlorinated alkanes, ethers and alcohols) occur as wastes or contaminants of products.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Curragh, H., Flynn, O., Larkin, M. J., Stafford, T. M., Hamilton, J. T. G., and Harper, D. B., 1994. Haloalkane degradation and assimilation by Rhodococcus rhodochrous NCIMB 13064, Microbiology 140/ 1433–1442.
Derewenda, Z. S., and Sharpe, A. M., 1993. News from the interface: The molecular structure of triacylglyceride lipases, Trends Biochem. Sci. 18:20–25.
Fox, B. G., Borneman, J. G., Wackett, L. P., and Lipscomb, J. D., 1990. Haloalkane oxidation by the soluble methane monooxygenase from Methylosinus trichosporium OB3b: mechanistic and environmental implications, Biochemistry 29:6419–6427.
Janssen, D. B., Pries, F., and van der Ploeg, J. R., 1994. Genetics and biochemistry of dehalogenating enzymes, Ann. Rev. Microbiol. 48:163–191.
Kasai, N., Tsujimura, K., Unoura, K., and Suzuki, T., 1990. Degradation of 2,3-dichloro-l-propanol by a Pseudomonas sp., Agric. Biol. Chem. 54:3185–3190.
Kawasaki, H., Tsuda, K., Matsushita, I., and Tonomura K., 1992. Lack of homology between two haloacetate de-halogenase genes encoded on a plasmid from Moraxella sp. strain B., J. Gen. Microbiol. 138:1317–1323.
Keuning, S., Janssen, D. B., and Witholt, B., 1985. Purification and characterization of hydrolytic haloalkane dehalogenase from Xanthobacter autotrophicus GJ10, J. Bacteriol. 163: 635–6
Kennes, C., Pries, F., Krooshof, G. H., Bokma, E., Kingma, J., and Janssen, D. B., 1995. Replacement of tryptophan residues in haloalkane dehalogenase reduces halide binding and catalytic activity, Eur. J. Biochem. 228:403–07.
Nagata, Y, Nariya, T., Ohtomo, R., Fukuda, M., Yano, K., and Takagi, M., 1993. Cloning and sequencing of a dehalogenase gene encoding an enzyme with hydrolase activity involved in the degradation of γ-hexachloro-cyclohexane in Pseudomonas paucimobilis, J. Bacteriol. 175: 6403–64
Nakamura, T., Nagasawa, T., Yu, F., Watanabe, I., and Yamada, H., 1992. Resolution and some properties of enzymes involved in enantioselective transformation of l,3-dichloro-2-propanol to (R)-3-chloro-l,2-propanediol by Corynebacterium sp. strain N-1074, J. Bacteriol. 174:7613–7619.
Ollis, D. L., Cheah, E., Cygler, M, Dijkstra, B. W., Frolow, F., Franken, S. M., Haral, M., Remington, S. J., Silman, I., Schrag, J., Sussman, J. L., Verschueren, K. H. G., and Goldman, A., 1992. The a/β-hydrolase fold, Protein Eng. 5:197–211.
Omori, T., Kimura, T., Kodama, T., 1991. Cloning of haloalkane dehalogenase genes of Rhodococcus sp. m 15-3 and expression and sequencing of the genes. pp498–503. In: On-Site Bioreclamation. Processes for Xenobiotic and Hydrocarbon Treatment. R.E. Hinchee & R.F. Olfenbuffel, Eds. Butterworth-Heinemann, Boston.
Pries, F., Kingma, J., Pentenga, M., van Pouderoyen, G., Jeronimus-Stratingh, C. M., Bruins, A. P., and Janssen, D. B., 1994a. Site-directed mutagenesis and oxygen isotope incorporation studies of the nucleophilic aspartate of haloalkane dehalogenase, Biochemistry 33:1242–1247.
Pries, F., van den Wijngaard, A. J., Bos, R., Pentenga, M., and Janssen, D. B., 1994b. The role of spontaneous cap domain mutations in haloalkane dehalogenase specificity and evolution, J. Biol. Chem. 269:17490–17494.
Pries, F., Kingma, J., Krooshof, G. H., Jeronimus-Stratingh, C. M., Bruins, A. P., and Janssen, D. B., 1995. Histidine 289 is essential for hydrolysis of the alkyl-enzyme intermediate of haloalkane dehalogenase, J. Biol Chem. 270:10405–10411.
Poulos, T. L., Finzel, B. C., and Howard, A., 1987, High resolution crystal structure of P450cam, J. Mol Biol 195:687–700.
Rosenzweig, A. C., Frederick, C. A., Lippard, S. J., and Nordlund, P., 1993. Crystal structure of a bacterial non-haem iron hydroxylase that catalyses the biological oxidation of methane, 366:537–543.
Sallis, P. J., Armfield, S. J., Bull, A. T., and Hardman, D. J., 1990. Isolation and characterization of a haloalkane halidohydrolase from Rhodococcus erythropolis Y2, J. Gen. Microbiol. 136:115–120.
Scholtz, R., Leisinger, T., Suter, F., and Cook, A. M., 1987. Characterization of 1-chlorohexane halidehydrolase, a dehalogenase of wide substrate range from an Arthrobacter sp., J. Bacteriol. 169: 5016–502
Stucki, G., and Thüer, M., 1995. Experiences of a large-scale application of 1,2-dichloroethane degrading microorganisms for groundwater treatment, Environ. Sci. Technol. 29:2339–2345.
Tardiff,C, Greer, C. W., Labbe, D., and Lau, P. C. K., 1991, Involvement of a large plasmid in the degradation of 1,2-dichloroethane by Xanthobacter autotrophicus GJ10, Appl Environ. Microbiol. 57:1853–1857.
van den Wijngaard, A. J., van der Kamp, K., van der Ploeg, J., Kazemier, B., Pries, F., and Janssen, D. B. 1992. Degradation of 1,2-dichloroethane by facultative methylotrophic bacteria, Appl Env. Microbiol. 58:976–983.
Verhagen, C., Smit, E., Janssen, D. B., and van Elsas, J. D., 1995. Bacterial dichloropropene degradation in soil, screening of soils and involvement of plasmids carrying the dhlA gene, Soil Biol Biochem. 27: 1547–155
Verschueren, K. H. G., Franken, S. M., Rozeboom, H. J., Kalk, K. H., and Dijkstra, B. W., 1993a. Refined X-ray structure of haloalkane dehalogenase at pH6.2 and pH8.2 and implications for the reaction mechanism, J. Mol. Biol. 232:856–872.
Verschueren, K. H. G., Seljée, F., Rozeboom, H. J., Kalk, K. H., and Dijkstra, B. W., 1993b. Crystallographic analysis of the catalytic mechanism of haloalkane dehalogenase, Nature 363:693–698.
Verschueren, K. H. G., Kingma, J., Rozeboom, H. J., Kalk, K. H., Janssen, D. B., and Dijkstra, B. W., 1993c. Crystallographic and fluorescence studies of the interaction of haloalkane dehalogenase with halide ions. Studies with halide compounds reveal a halide binding site in the active site, Biochemistry 32:9031–9037.
Wackett, L. P., Sadowsky, M. J., Newman, L. M., Hur, H-G., and Li, S., 1994. Metabolism of polyhalogenated compounds by a genetically engineered bacterium, Nature 368:627–629.
Yokota, T., Fuse, H., Omori, T., and Minoda, Y, 1986. Microbial dehalogenation of haloalkanes mediated by oxygenase or halidohydrolase, Agric. Biol Chem. 50:453–460.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 1997 Springer Science+Business Media New York
About this chapter
Cite this chapter
Schanstra, J.P., Poelarends, G.J., Bosma, T., Janssen, D.B. (1997). Engineering Enzymes and Microorganisms for the Transformation of Synthetic Compounds. In: Sayler, G.S., Sanseverino, J., Davis, K.L. (eds) Biotechnology in the Sustainable Environment. Environmental Science Research, vol 54. Springer, Boston, MA. https://doi.org/10.1007/978-1-4615-5395-3_5
Download citation
DOI: https://doi.org/10.1007/978-1-4615-5395-3_5
Publisher Name: Springer, Boston, MA
Print ISBN: 978-1-4613-7463-3
Online ISBN: 978-1-4615-5395-3
eBook Packages: Springer Book Archive