Abstract
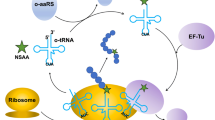
Natural proteins are normally made by 20 canonical amino acids. Genetic code expansion (GCE) enables incorporation of diverse chemically synthesized noncanonical amino acids (ncAAs) by orthogonal aminoacyl-tRNA synthetase (aaRS)/tRNA pairs using nonsense codons, which could significantly expand new functionalities of proteins in both scientific and biomedical applications. Here, by hijacking the cysteine biosynthetic enzymes, we describe a method combining amino acid biosynthesis and GCE to introduce around 50 structurally novel ncAAs into proteins by supplementation of commercially available aromatic thiol precursors, thus eliminating the need to chemically synthesize these ncAAs. A screening method is also provided for improving the incorporation efficiency of a particular ncAA. Furthermore, we demonstrate bioorthogonal groups, such as azide and ketone, that are compatible with our system and can be easily introduced into protein for subsequent site-specific labeling.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Macek B, Forchhammer K, Hardouin J, Weber-Ban E, Grangeasse C, Mijakovic I (2019) Protein post-translational modifications in bacteria. Nat Rev Microbiol 17(11):651–664. https://doi.org/10.1038/s41579-019-0243-0
Dumas A, Lercher L, Spicer CD, Davis BG (2015) Designing logical codon reassignment – expanding the chemistry in biology. Chem Sci 6(1):50–69. https://doi.org/10.1039/c4sc01534g
Wang L, Brock A, Herberich B, Schultz PG (2001) Expanding the genetic code of Escherichia coli. Science 292(5516):498–500. https://doi.org/10.1126/science.1060077
Wang J, Wang X, Fan X, Chen PR (2021) Unleashing the power of bond cleavage chemistry in living systems. ACS Cent Sci 7(6):929–943. https://doi.org/10.1021/acscentsci.1c00124
Wang X, Liu D, Shen L, Li F, Li Y, Yang L, Xu T, Tao H, Yao D, Wu L, Hirata K, Bohn LM, Makriyannis A, Liu X, Hua T, Liu Z-J, Wang J (2021) A genetically encoded F-19 NMR probe reveals the allosteric modulation mechanism of cannabinoid receptor 1. J Am Chem Soc 143(40):16320–16325. https://doi.org/10.1021/jacs.1c06847
Park SH, Ko W, Park SH, Lee HS, Shin I (2019) Evaluation of the interaction between Bax and Hsp70 in cells by using a FRET system consisting of a fluorescent amino acid and YFP as a FRET pair. Chembiochem 21(1–2):59–63. https://doi.org/10.1002/cbic.201900293
Zhang B, Sun J, Wang Y, Ji D, Yuan Y, Li S, Sun Y, Hou Y, Li P, Zhao L, Yu F, Ma W, Cheng B, Wu L, Hu J, Wang M, Song W, Li X, Li H, Fei Y, Chen H, Zhang L, Tsokos GC, Zhou D, Zhang X (2021) Site-specific PEGylation of interleukin-2 enhances immunosuppression via the sustained activation of regulatory T cells. Nat Biomed Eng 5(11):1288–1305. https://doi.org/10.1038/s41551-021-00797-8
Ptacin JL, Caffaro CE, Ma L, San Jose Gall KM, Aerni HR, Acuff NV, Herman RW, Pavlova Y, Pena MJ, Chen DB, Koriazova LK, Shawver LK, Joseph IB, Milla ME (2021) An engineered IL-2 reprogrammed for anti-tumor therapy using a semi-synthetic organism. Nat Commun 12(1). https://doi.org/10.1038/s41467-021-24987-9
Zhang J, Ji D, Shen W, Xiao Q, Gu Y, O’Shaughnessy J, Hu X (2022) Phase I trial of a novel anti-HER2 antibody–drug conjugate, ARX788, for the treatment of HER2-positive metastatic breast cancer. Clin Cancer Res:OF1–OF10. https://doi.org/10.1158/1078-0432.Ccr-22-0456
Ling X, Chang L, Chen H, Gao X, Yin J, Zuo Y, Huang Y, Zhang B, Hu J, Liu T (2021) Improving the efficiency of CRISPR-Cas12a-based genome editing with site-specific covalent Cas12a-crRNA conjugates. Mol Cell 81(22):4747–4756.e4747. https://doi.org/10.1016/j.molcel.2021.09.021
Ling X, Xie B, Gao X, Chang L, Zheng W, Chen H, Huang Y, Tan L, Li M, Liu T (2020) Improving the efficiency of precise genome editing with site-specific Cas9-oligonucleotide conjugates. Sci Adv 6(15). https://doi.org/10.1126/sciadv.aaz0051
Tang H, Dai Z, Qin X, Cai W, Hu L, Huang Y, Cao W, Yang F, Wang C, Liu T (2018) Proteomic identification of protein tyrosine phosphatase and substrate interactions in living mammalian cells by genetic encoding of irreversible enzyme inhibitors. J Am Chem Soc 140(41):13253–13259. https://doi.org/10.1021/jacs.8b06922
Gaston MA, Zhang LW, Green-Church KB, Krzycki JA (2011) The complete biosynthesis of the genetically encoded amino acid pyrrolysine from lysine. Nature 471(7340):647–650. https://doi.org/10.1038/nature09918
Mehl RA, Anderson JC, Santoro SW, Wang L, Martin AB, King DS, Horn DM, Schultz PG (2003) Generation of a bacterium with a 21 amino acid genetic code. J Am Chem Soc 125(4):935–939. https://doi.org/10.1021/ja0284153
Chen Y, Loredo A, Gordon A, Tang J, Yu C, Ordonez J, Xiao H (2018) A noncanonical amino acid-based relay system for site-specific protein labeling. Chem Commun 54(52):7187–7190. https://doi.org/10.1039/c8cc03819h
Chen Y, Tang J, Wang L, Tian Z, Cardenas A, Fang X, Chatterjee A, Xiao H (2020) Creation of bacterial cells with 5-hydroxytryptophan as a 21st amino acid building block. Chem 6(10):2717–2727. https://doi.org/10.1016/j.chempr.2020.07.013
Jung J-E, Lee SY, Park H, Cha H, Ko W, Sachin K, Kim DW, Chi DY, Lee HS (2014) Genetic incorporation of unnatural amino acids biosynthesized from α-keto acids by an aminotransferase. Chem Sci 5(5). https://doi.org/10.1039/c3sc51617b
Kim S, Sung BH, Kim SC, Lee HS (2018) Genetic incorporation ofl-dihydroxyphenylalanine (DOPA) biosynthesized by a tyrosine phenol-lyase. Chem Commun 54(24):3002–3005. https://doi.org/10.1039/c8cc00281a
Exner MP, Kuenzl T, To TMT, Ouyang Z, Schwagerus S, Hoesl MG, Hackenberger CPR, Lensen MC, Panke S, Budisa N (2017) Design of S-allylcysteine in situ production and incorporation based on a novel pyrrolysyl-tRNA synthetase variant. Chembiochem 18(1):85–90. https://doi.org/10.1002/cbic.201600537
Zhang MS, Brunner SF, Huguenin-Dezot N, Liang AD, Schmied WH, Rogerson DT, Chin JW (2017) Biosynthesis and genetic encoding of phosphothreonine through parallel selection and deep sequencing. Nat Methods 14(7):729–736. https://doi.org/10.1038/nmeth.4302
Maier THP (2003) Semisynthetic production of unnatural L-α-amino acids by metabolic engineering of the cysteine-biosynthetic pathway. Nat Biotechnol 21(4):422–427. https://doi.org/10.1038/nbt807
Wang Y, Chen X, Cai W, Tan L, Yu Y, Han B, Li Y, Xie Y, Su Y, Luo X, Liu T (2021) Expanding the structural diversity of protein building blocks with noncanonical amino acids biosynthesized from aromatic thiols. Angew Chem Int Ed 60(18):10040–10048. https://doi.org/10.1002/anie.202014540
Liu Y, Kim J, Seo H, Park S, Chae J (2015) Copper(II)-catalyzed single-step synthesis of aryl thiols from aryl halides and 1,2-ethanedithiol. Adv Synth Catal 357(10):2205–2212. https://doi.org/10.1002/adsc.201400941
Brai A, Martelli F, Riva V, Garbelli A, Fazi R, Zamperini C, Pollutri A, Falsitta L, Ronzini S, Maccari L, Maga G, Giannecchini S, Botta M (2019) DDX3X helicase inhibitors as a new strategy to fight the West Nile virus infection. J Med Chem 62(5):2333–2347. https://doi.org/10.1021/acs.jmedchem.8b01403
Wirtz M, Hell R (2003) Production of cysteine for bacterial and plant biotechnology: application of cysteine feedback-insensitive isoforms of serine acetyltransferase. Amino Acids 24(1):195–203. https://doi.org/10.1007/s00726-002-0313-9
Liu CC, Schultz PG (2010) Adding new chemistries to the genetic code. Annu Rev Biochem 79(1):413–444. https://doi.org/10.1146/annurev.biochem.052308.105824
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Wang, Y., Cai, W., Han, B., Liu, T. (2023). Protein Expression with Biosynthesized Noncanonical Amino Acids. In: Tsai, YH., Elsässer, S.J. (eds) Genetically Incorporated Non-Canonical Amino Acids. Methods in Molecular Biology, vol 2676. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3251-2_6
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3251-2_6
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3250-5
Online ISBN: 978-1-0716-3251-2
eBook Packages: Springer Protocols