Abstract
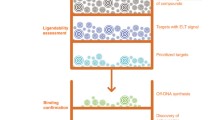
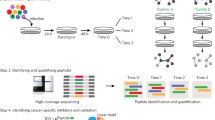
Functional genomic screens can identify several proteins as potential targets for drug development in cancer. Typically, these drug targets are validated with pharmacological inhibition using small molecules. Given that chemical inhibitors do not exist for a many of these proteins, several promising candidates often remain unexplored. In this chapter, we describe methods for generating protein-based inhibitors of intracellular targets using phage display. This is a scalable and inexpensive approach that can be applied to several protein targets identified in genetic screens. We describe methods for expression of target proteins, construction of phage-display libraries and selection of binding proteins. These synthetic binding proteins can block natural protein interactions within the cancer cell and act as inhibitors. Protein inhibitors have utility in validation of drug targets and can also guide small-molecule drug development.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
O’Loughlin TA, Gilbert LA (2019) Functional genomics for cancer research: applications in vivo and in vitro. Annual Rev Cancer Biol 3(1):345–363. https://doi.org/10.1146/annurev-cancerbio-030518-055742
Haley B, Roudnicky F (2020) Functional genomics for cancer drug target discovery. Cancer Cell 38(1):31–43. https://doi.org/10.1016/j.ccell.2020.04.006
Parameswaran S, Kundapur D, Vizeacoumar FS et al (2019) A road map to personalizing targeted cancer therapies using synthetic lethality. Trends Cancer 5(1):11–29. https://doi.org/10.1016/j.trecan.2018.11.001
Sato M (2020) Phenotypic screening using large-scale genomic libraries to identify drug targets for the treatment of cancer. Oncol Lett 19(6):3617–3626. https://doi.org/10.3892/ol.2020.11512
Weiss WA, Taylor SS, Shokat KM (2007) Recognizing and exploiting differences between RNAi and small-molecule inhibitors. Nat Chem Biol 3(12):739–744. https://doi.org/10.1038/nchembio1207-739
Edwards AM, Isserlin R, Bader GD et al (2011) Too many roads not taken. Nature 470(7333):163–165. https://doi.org/10.1038/470163a
Antolin AA, Tym JE, Komianou A et al (2018) Objective, quantitative, data-driven assessment of chemical probes. Cell Chem Biol 25(2):194–205. e195. https://doi.org/10.1016/j.chembiol.2017.11.004
Kehoe JW, Kay BK (2005) Filamentous phage display in the new millennium. Chem Rev 105(11):4056–4072. https://doi.org/10.1021/cr000261r
Smith GP (1985) Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 228(4705):1315–1317
Sachdev SS, Frederic AF (2006) Making antibodies in bacteria. In: Making and using antibodies. CRC Press, Boca Raton, Florida, pp 157–180. https://doi.org/10.1201/9781420005196.ch810.1201/9781420005196.ch8
Grover P, Shi H, Baumgartner M et al (2015) Fluorescence polarization screening assays for small molecule allosteric modulators of ABL kinase function. PLoS One 10(7):e0133590. https://doi.org/10.1371/journal.pone.0133590
Lea WA, Simeonov A (2011) Fluorescence polarization assays in small molecule screening. Expert Opin Drug Discov 6(1):17–32. https://doi.org/10.1517/17460441.2011.537322
Batyuk A, Wu Y, Honegger A et al (2016) DARPin-based crystallization chaperones exploit molecular geometry as a screening dimension in protein crystallography. J Mol Biol 428(8):1574–1588. https://doi.org/10.1016/j.jmb.2016.03.002
Koide S (2009) Engineering of recombinant crystallization chaperones. Curr Opin Struct Biol 19(4):449–457. https://doi.org/10.1016/j.sbi.2009.04.008
Arkin MR, Tang Y, Wells JA (2014) Small-molecule inhibitors of protein-protein interactions: progressing toward the reality. Chem Biol 21(9):1102–1114. https://doi.org/10.1016/j.chembiol.2014.09.001
Ilic S, Akabayov SR, Arthanari H et al (2016) Identification of DNA primase inhibitors via a combined fragment-based and virtual screening. Sci Rep 6:36322. https://doi.org/10.1038/srep36322
Reis JM, Xu X, McDonald S et al (2018) Discovering selective binders for Photoswitchable proteins using phage display. ACS Synth Biol 7(10):2355–2364. https://doi.org/10.1021/acssynbio.8b00123
Nord K, Gunneriusson E, Ringdahl J et al (1997) Binding proteins selected from combinatorial libraries of an alpha-helical bacterial receptor domain. Nat Biotechnol 15(8):772–777. https://doi.org/10.1038/nbt0897-772
Leung I, Jarvik N, Sidhu SS (2017) A highly diverse and functional naive ubiquitin variant library for generation of intracellular affinity reagents. J Mol Biol 429(1):115–127. https://doi.org/10.1016/j.jmb.2016.11.016
Schlatter D, Brack S, Banner DW et al (2012) Generation, characterization and structural data of chymase binding proteins based on the human Fyn kinase SH3 domain. MAbs 4(4):497–508. https://doi.org/10.4161/mabs.20452
Uppalapati M, Lee DJ, Mandal K et al (2016) A potent D-protein antagonist of VEGF-A is nonimmunogenic, metabolically stable, and longer-circulating in vivo. ACS Chem Biol 11(4):1058–1065. https://doi.org/10.1021/acschembio.5b01006
Uppalapati M, Sidhu SS, Kerman A (2014) Scaffolded peptidic libraries and methods of making and screening the same, International Patent WO 2014/140882 A2, 14 Mar 2014
Gilbreth RN, Esaki K, Koide A et al (2008) A dominant conformational role for amino acid diversity in minimalist protein-protein interfaces. J Mol Biol 381(2):407–418. https://doi.org/10.1016/j.jmb.2008.06.014
Kunkel TA, Roberts JD, Zakour RA (1987) Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol 154:367–382. https://doi.org/10.1016/0076-6879(87)54085-x
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
McDonald, S., Annan Sudarsan, A., Babeker, H., Budharaju, K., Uppalapati, M. (2021). Generation of Protein Inhibitors for Validation of Cancer Drug Targets Identified in Functional Genomic Screens. In: Vizeacoumar, F.J., Freywald, A. (eds) Mapping Genetic Interactions. Methods in Molecular Biology, vol 2381. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-1740-3_17
Download citation
DOI: https://doi.org/10.1007/978-1-0716-1740-3_17
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-1739-7
Online ISBN: 978-1-0716-1740-3
eBook Packages: Springer Protocols