Abstract
Purpose
Testing for BRCA1 and BRCA2 mutations in breast cancer patients is used to identify the risk of second primary cancers and the risk of cancer in the patients’ family. Women with triple-negative breast cancer (TNBC) are thought to be more likely to be BRCA1/2 mutation carriers, but most national guidelines for genetic testing, including those used in Germany and Austria, do not consider receptor triple negativity.
Methods
We determined the prevalence of BRCA1 and BRCA2 mutations within a cohort of 100 unselected TNBC cases, including patients from Germany and Austria to identify those BRCA-positive patients with a masked family history and who would have been missed due to respective current national guidelines. Double-stranded Sanger sequencing of all exons of BRCA1 and BRCA2, respectively, was performed.
Results
We identified a total of 13 deleterious mutations in BRCA1 and a total of four deleterious mutations in BRCA2. The total rate of deleterious BRCA1/2 mutation carriers was 21 % in our cohort. Six novel mutations, including two deleterious mutations, have been identified, which have not been described in public mutation databases so far. According to current German and Austrian national guidelines for genetic testing, 38.1 and 52.4 %, respectively, of BRCA1/2 mutation carriers would have been overlooked.
Conclusions
We conclude that the prevalence of BRCA1 and BRCA2 mutations is high in TNBC patients and that BRCA1/2 mutations are not restricted to young women or patients with a positive family history. Receptor triple negativity should therefore be considered in BRCA1/2 genetic testing guidelines.
Similar content being viewed by others
Introduction
Triple-negative breast cancer (TNBC), defined by the lack of expression of estrogen receptor (ER) and progesterone receptor (PR) as well as the absence of human epidermal growth factor receptor 2 (HER2) overexpression, accounts for approximately 15 % of all invasive breast cancer cases (Foulkes et al. 2010; Pal et al. 2011). TNBC is associated with aggressive behavior and a worse prognosis than other breast cancer subtypes (Irvin and Carey 2008; Pal et al. 2011). Given the lack of an established receptor target, targeted treatments are not available for TNBC, leaving chemotherapy as the mainstay of treatment, so far (Irvin and Carey 2008). That said, a number of targeted approaches to TNBC are undergoing clinical evaluation, including poly (ADP-ribose) polymerase (PARP) inhibitors, epidermal growth factor receptor inhibitors, and anti-angiogenic agents (Pal et al. 2011).
TNBC is a heterogeneous disease consisting of different subtypes each displaying a distinct biology (Mayer et al. 2014). In this regard, TNBC has been associated with breast cancer susceptibility gene 1 (BRCA1) and BRCA2 mutational status. Frequency of TNBC has been reported to be 57 % in BRCA1 mutation carriers and 23 % in BRCA2-positive patients (Atchley et al. 2008).
BRCA1 and BRCA2 are critically involved in pathways that regulate homologous-recombination-mediated DNA repair, cell cycle progression, and apoptosis (Foulkes and Shuen 2013) and are believed to be the two major contributors of hereditary breast cancer. It is estimated that women with deleterious BRCA1 or BRCA2 mutations have a lifetime risk of developing breast cancer of 50–85 % (Ford et al. 1994, 1998) and of developing ovarian cancer of 40–60 % (Antoniou et al. 2003; King et al. 2003). Genetic testing criteria may differ between countries based on mutation prevalence. According to current guidelines of some countries (e.g., Germany or the UK), genetic testing for BRCA1 and BRCA2 genes is recommended in women with breast cancer, who had at least a 10 % prior probability of carrying a BRCA1/2 mutation, primarily based on familial criteria (Gadzicki et al. 2011; Meindl 2002; Balmana et al. 2011). After a mutation has been identified within the index patient, predictive testing is then recommended for healthy relatives, who might benefit from preventive interventions.
Recent studies have shown that germline mutations of BRCA1 and to a lesser extent BRCA2 are overrepresented in patients with TNBC. In particular, high BRCA1 and BRCA2 mutation frequencies were reported in Ashkenazi Jewish women with TNBC revealing a BRCA1 mutation frequency of up to 29 % and a BRCA2 mutation frequency of 9 % (Comen et al. 2011). A recent meta-analysis including 2,533 high-risk breast cancer patients showed that women with TNBC are approximately five and a half times more likely to carry a BRCA1 mutation compared with non-TNBC phenotype and that approximately two in nine women with TNBC harbor a BRCA1 mutation (Tun et al. 2014). In studies including patient populations with TNBC regardless of a family history of cancer, reported BRCA1 mutation frequencies ranged between 6 and 15 % (Couch et al. 2014; Gonzalez-Angulo et al. 2011; Hartman et al. 2012; Rummel et al. 2013; Pern et al. 2012; Wong-Brown et al. 2015) and BRCA2 mutation frequencies ranged between 3 and 16 % (Couch et al. 2014; Gonzalez-Angulo et al. 2011; Hartman et al. 2012; Meyer et al. 2012; Pern et al. 2012; Wong-Brown et al. 2015).
Due to high BRCA1/2 mutation prevalence among TNBC patients, TNBC status was included in current National Comprehensive Cancer Network (NCCN) guidelines (version 1.2014) for BRCA1/2 testing (Daly et al. 2014). However, BRCA1/2 mutation frequency among patients with TNBC may differ between populations (Balmana et al. 2011) and receptor triple negativity is not included in genetic testing guidelines of several countries including Germany and Austria (Gadzicki et al. 2011; Singer et al. 2012). Therefore, we genotyped BRCA1 and BRCA2 genes in a cohort of 100 unselected TNBC patients from Southern Germany and Western Austria to identify those BRCA1-/2-positive patients with a masked family history of breast or ovarian cancer and who would have been missed due to respective current national guidelines.
Methods
Study population
From May 2011 through December 2013, we enrolled 100 consecutive Caucasian female patients diagnosed with TNBC, regardless of age and family history of cancer. Of these, 43 patients were recruited at the Academic Teaching Hospital Feldkirch (Feldkirch, Austria) and 57 patients were recruited at the Oncology Study Center Ravensburg (Ravensburg, Germany). All patients gave written informed consent for genetic testing and participation in the present study. The study protocol was approved by the Ethic Committee of the State of Vorarlberg (Austria).
ER, PR, and HER2 status were ascertained as follows: ER and PR status were determined by immunohistochemistry (IHC) and classified as negative if less than 1 % of cells showed stained nuclei in the tumor sample (Hammond et al. 2010). For HER2, IHC scores of 0 and +1 were considered as negative, and +2 scores were also classified as negative in case of a negative fluorescence in situ hybridization (FISH) result (FISH ratio of less than 1.8) (Wolff et al. 2007).
Tumors were classified according to the guidelines of the American Joint Committee on Cancer (AJCC) staging system (Edge et al. 2009). Family history of cancer was defined as diagnosed breast or ovarian cancer in first- and second-degree relatives.
BRCA1/2 mutation analysis
Genomic DNA was isolated from 5 ml EDTA blood using the QIAamp DNA Blood Midi Kit (Qiagen, Hilden, Germany). All exons and exon–intron boundaries of BRCA1 (exons 1–3, 5–24) and BRCA2 (exons 1–27) were amplified by polymerase chain reaction (PCR). Used PCR primers and PCR conditions are given in Online Resource 1. PCR products were purified using ExoSAP-IT® reagent (Affymetrix, Santa Clara, CA) and sequenced bidirectionally using BigDye® Cycle Sequencing Kit (Life Technologies, San Francisco, CA) on an ABI 96-capillary 3730xl DNA Analyzer (Life Technologies). Alignments to the reference sequences NG_005905.2 for BRCA1 and NG_012772.1 for BRCA2 were performed using SeqMan Pro software (DNASTAR, Inc., Madison, WI).
Detected sequence variants were checked for previously published reports in the dbSNP database (http://www.ncbi.nlm.nih.gov/snp), in the Breast Cancer Information Core (BIC) database (http://research.nhgri.nih.gov/bic/), in the Leiden Open Variation Database (LOVD, http://chromium.liacs.nl/LOVD2/home.php), and in the Universal Mutation Database (UMD, http://www.umd.be), as well as in PubMed (http://www.ncbi.nlm.nih.gov/pubmed).
The effect of missense mutations was evaluated with in silico bioinformatic tools “sorting intolerant from tolerant” (SIFT; http://sift.jcvi.org) (Kumar et al. 2009) and “polymorphism phenotyping v2” (PolyPhen-2; http://genetics.bwh.harvard.edu/pph2) (Adzhubei et al. 2010). SIFT predicts substitutions with scores less than 0.05 as deleterious, and PolyPhen-2 scores are evaluated as 0.000 (most probably benign) to 0.999 (most probably damaging). The effect of potential splice site variations was evaluated by “Human Splicing Finder” (http://www.umd.be/HSF/) (Desmet et al. 2009) and “Splice Site Prediction by Neural Network” (http://www.fruitfly.org/seq_tools/splice.html) (Reese et al. 1997) tools.
Current guidelines for genetic testing
Current NCCN guidelines for genetic testing are given in the “NCCN Guidelines for Genetic/Familial High-Risk Assessment: Breast and Ovarian” (version 1.2014) (Daly et al. 2014). In Austria, criteria for BRCA1/2 testing are given in the “Austrian guideline for prevention and early detection of breast and ovarian cancer in high-risk patients” (Singer et al. 2012), established by the “Austrian Society of Gynaecology and Obstetrics.” In Germany, BRCA1/2 testing criteria are cited in the “Interdisciplinary S3 guidelines for the Diagnosis, Treatment and Follow-up Care of Breast Cancer” (http://www.senologie.org/fileadmin/downloads/S3-Brustkrebs-v2012-OL-Kurzversion.pdf) (Gadzicki et al. 2011). Current guidelines for BRCA testing of the NCCN as well as in Germany and Austria are summarized in Online Resource 2. NCCN guidelines recommend genetic testing in women with TNBC diagnosed ≤60 years. In contrast, German and Austrian guidelines do not include receptor triple negativity.
Results
Mutational status
Of the 100 patients tested, 17 patients carried a deleterious BRCA1 mutation and four patients carried a deleterious BRCA2 mutation. Consequently, 21 % of our TNBC patients showed a positive BRCA mutational status. A total of 13 different deleterious BRCA1 mutations and four different deleterious BRCA2 mutations could be detected including nine frameshift, four missense, two nonsense, and two splice mutations as given in Table 1. According to previously published reports (Karchin et al. 2008) and in silico bioinformatic tools SIFT (score 0) and PolyPhen-2 (score 1), BRCA2 missense mutation c.9104A>C was classified as pathogenic although still assessed as an unclassified variant in BIC database and UMD. Four recurrent pathogenic BRCA1 mutations (c.3016_3019delCATT, c.4986+3G>C, c.5161C>T, and c.5265_5266insC) were found in our TNBC cohort, each affecting two unrelated cases.
Furthermore, two mutations with unclear clinical impact were detected: In silico analysis of BRCA1 variant c.1243G>A (p.Val415Ile) resulted in a proposed benign effect using SIFT and a possible damaging effect on protein function using PolyPhen-2 (score 0.902). Concordantly, the mutation is listed in UMD as an unclassified variant. Equally, missense BRCA1 mutation c.5151T>C (p.Trp1718Arg) showed controversial in silico prediction results with a benign effect using PolyPhen-2 and a deleterious effect using SIFT (score 0). The mutation is neither listed in public mutation databases nor quoted in PubMed, thus not allowing any comparisons with findings of others. Notably, the two carriers of mentioned unclassified variants showed a negative family history of breast or ovarian cancer pointing to a benign effect of both variants.
In total, six variants have been identified, which have not been described in public BRCA mutation databases (dbSNP, BIC, LOVD, and UMD) or in PubMed so far: BRCA1: c.-82G>A c.212+157delA, c.3915delC, c.5151T>C, and c.5230_5237delAGAAACCA; BRCA2: c.8010G>A. Novel mutations are given in Online Resource 3, and respective screenshots of sequence chromatograms are presented in the Online Resource 4. BRCA1 nonsense mutation c.3915delC introduces a premature stop codon at amino acid position 1306 (p.Leu1306Ter), and BRCA1 frameshift mutation c.5230_5237delAGAAACCA introduces a premature stop codon at amino acid position 1826 (p.Arg1744Profs1826Ter). Therefore, both mutations were classified as pathogenic. Mutation c.5151T>C (p.Trp1718Arg) was classified as an unclassified variant (see above). Using mentioned in silico tools, remaining novel mutations did not show any deleterious effect on protein function (data not shown) and therefore were classified as benign.
Patients’ characteristics
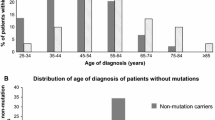
Clinical characteristics of patients with likely causal BRCA1 and BRCA2 mutations compared to non-BRCA carriers (including carriers of unclassified variants) are given in Table 2. Carriers of BRCA1 or BRCA2 mutations were younger at TNBC diagnosis compared to non-carriers, but association between BRCA1/2 mutation status and age at diagnosis did not reach statistical significance in our study. Other clinical parameters including stage and grade were not significantly associated with BRCA1/2 mutation status. As to be expected, BRCA1/2 mutation status was significantly linked with a family history of breast or ovarian cancer.
BRCA1/2 mutation carriers meeting current guidelines
Applying current guidelines for genetic testing revealed that among the 21 TNBC patients carrying a causal BRCA1 or BRCA2 mutation, 19 cases (90.5 %) met testing criteria of the current NCCN guidelines, 13 cases (61.9 %) met criteria of the current German guidelines, and 10 cases (47.6 %) met criteria of the current Austrian guidelines (Table 3).
Discussion
In the present study, 21 % of our TNBC patients unselected for age and family history of breast or ovarian cancer carried a deleterious germline BRCA1 or BRCA2 mutation. The observed total BRCA1/2 mutation frequency in unselected TNBC cases is comparable with previous findings (Gonzalez-Angulo et al. 2011; Meyer et al. 2012; Pern et al. 2012), although some studies reported a lower BRCA1/2 mutation prevalence (Couch et al. 2014; Hartman et al. 2012; Wong-Brown et al. 2015). Particularly, in the so far largest series of TNBC cases unselected for family history of cancer, comprising 1,824 subjects, BRCA1 and BRCA2 mutations were found in 11.2 % of patients (Couch et al. 2014). Differences in ancestral backgrounds, patient selection criteria, and genotyping methods may have contributed to deviating BRCA1/2 mutation frequencies between studies. However, in most of the studies the prevalence of BRCA1/2 mutations in unselected TNBC patients was equal or clearly exceeded 10 % (Couch et al. 2014; Gonzalez-Angulo et al. 2011; Hartman et al. 2012; Pern et al. 2012; Wong-Brown et al. 2015), used as an a priori probability threshold of carrying a BRCA1/2 mutation in current guidelines for genetic testing (Gadzicki et al. 2011; Meindl 2002; Balmana et al. 2011).
Identification of a deleterious BRCA1/2 mutation in a patient with TNBC significantly impacts clinical management. BRCA1-/2-positive patients are more likely to develop bilateral or ovarian cancer. Thus, identification of BRCA1/2 mutations is critical in optimizing surgical management and risk reduction strategies (Rummel et al. 2013). In addition, in breast cancer patients, response rates to therapies including DNA-damaging chemotherapeutics, such as platinum, or novel DNA repair-targeted agents, such as PARP inhibitors, appear to be influenced by BRCA1 or BRCA2 mutations (Telli 2014; Fong et al. 2009). Consequently, BRCA1/2 mutation status has emerged as an important potential predictive biomarker for these therapies. Furthermore, children of a parent with a BRCA1 or BRCA2 mutation have a 50 % risk of having inherited the mutation. Women carrying a BRCA1/2 mutation have strongly elevated risks of breast cancer and ovarian cancer, with a lifetime risk of breast cancer as high as 85 % (Ford et al. 1994, 1998) and a up to 60 % chance of developing ovarian cancer (Antoniou et al. 2003; King et al. 2003). In particular with mutations in the BRCA2 gene, risk is increased in women and men for pancreatic cancer and in men for prostate cancer (Mersch et al. 2015). It is, therefore, important that adult relatives are informed about cancer risks and should be made aware of the options for genetic counselling, testing, and management, as necessary.
Current guidelines for genetic testing in a breast cancer patient are primarily based on a family history of cancer or the age at onset. However, not all of our BRCA1-/2-positive patients showed an appropriate family history of cancer or early age of diagnosis missing common criteria for genetic testing. As a matter of fact, according to current German and Austrian national guidelines for genetic testing, which do not include TNBC, 38.1 and 52.4 %, respectively, of BRCA1/2 carriers would have been missed in our cohort of TNBC patients. These results suggest that beside a positive family history of cancer or early age of diagnosis of breast cancer, receptor triple negativity should be considered in the guidelines for genetic analysis of BRCA1 and BRCA2. Noteworthily, according to less stringent NCCN guidelines (Daly et al. 2014) recommending genetic testing in TNBC patients before or at age 60, still 9.5 % of BRCA1/2 carriers would have been overlooked. Therefore, perhaps all patients with TNBC irrespective of age or family history should be offered genetic testing as also proposed by other authors (Couch et al. 2014; Rummel et al. 2013).
Our study indentified six variants, including two deleterious mutations, which were not listed in public BRCA mutation databases (dbSNP, BIC, LOVD, and UMD) or were described in PubMed so far. Particularly, only 10 out of 17 deleterious BRCA1/2 mutations were given in the frequently used BIC database. Therefore, public databases still appear incomplete. This gap may be filled by the implementation of large future studies facilitated by the use of new sequencing technologies decreasing extensively time and cost of mutation screening. In fact, very recently, mutations in BRCA1 and BRCA2 as well as in other breast cancer susceptibility genes have been screened in large series of TNBC patients using next-generation sequencing technologies (Couch et al. 2014; Wong-Brown et al. 2015).
There are potential limitations of our study which should be considered. Sample size of the included study population is moderate. Therefore, we cannot exclude that some associations not found to be significant in our study may have reached statistical significance with a larger population. Notably, however, studies on BRCA1/2 genes in unselected TNBC patients from Germany or Austria are scarce and our study discovered several mutations, including two deleterious mutations, undescribed so far. In addition, BRCA1/2 mutation detection was performed by direct double-stranded Sanger DNA sequencing, which still represents the gold standard in mutation analysis (Balmana et al. 2011). That said, analysis of large exonic deletions or duplications in BRCA1 or BRCA2 was not performed in our study. However, large rearrangements are rare among TNBC patients with frequencies between 0.7 and 2.0 % (Couch et al. 2014; Hartman et al. 2012). Therefore, one to two carriers of a rearrangement mutation could have been missed in our cohort of 100 TNBC cases and BRCA1/2 mutation rate may be even higher as observed.
We conclude that the prevalence of BRCA1 and BRCA2 mutations is high in TNBC patients and that BRCA1/2 mutations are not restricted to young women or patients with a positive family history of breast or ovarian cancer. TNBC patients with BRCA1/2 mutations require different strategies for patient management and counselling and respond differently to targeted therapies and chemotherapies. Therefore, TNBC patients may benefit from BRCA genetic testing to indentify those with a masked family history and who do not meet current guidelines for genetic testing. Receptor triple negativity should therefore be considered in BRCA1/2 genetic testing guidelines.
References
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249
Antoniou A, Pharoah PD, Narod S, Risch HA, Eyfjord JE, Hopper JL, Loman N, Olsson H, Johannsson O, Borg A, Pasini B, Radice P, Manoukian S, Eccles DM, Tang N, Olah E, Anton-Culver H, Warner E, Lubinski J, Gronwald J, Gorski B, Tulinius H, Thorlacius S, Eerola H, Nevanlinna H, Syrjakoski K, Kallioniemi OP, Thompson D, Evans C, Peto J, Lalloo F, Evans DG, Easton DF (2003) Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case Series unselected for family history: a combined analysis of 22 studies. Am J Hum Genet 72:1117–1130
Atchley DP, Albarracin CT, Lopez A, Valero V, Amos CI, Gonzalez-Angulo AM, Hortobagyi GN, Arun BK (2008) Clinical and pathologic characteristics of patients with BRCA-positive and BRCA-negative breast cancer. J Clin Oncol 26:4282–4288
Balmana J, Diez O, Rubio IT, Cardoso F (2011) BRCA in breast cancer: ESMO clinical practice guidelines. Ann Oncol 22(Suppl 6):vi31–vi34
Comen E, Davids M, Kirchhoff T, Hudis C, Offit K, Robson M (2011) Relative contributions of BRCA1 and BRCA2 mutations to “triple-negative” breast cancer in Ashkenazi Women. Breast Cancer Res Treat 129:185–190
Couch FJ, Hart SN, Sharma P, Toland AE, Wang X, Miron P, Olson JE, Godwin AK, Pankratz VS, Olswold C, Slettedahl S, Hallberg E, Guidugli L, Davila JI, Beckmann MW, Janni W, Rack B, Ekici AB, Slamon DJ, Konstantopoulou I, Fostira F, Vratimos A, Fountzilas G, Pelttari LM, Tapper WJ, Durcan L, Cross SS, Pilarski R, Shapiro CL, Klemp J, Yao S, Garber J, Cox A, Brauch H, Ambrosone C, Nevanlinna H, Yannoukakos D, Slager SL, Vachon CM, Eccles DM, Fasching PA (2014) Inherited mutations in 17 breast cancer susceptibility genes among a large triple-negative breast cancer cohort unselected for family history of breast cancer. J Clin Oncol 33:304–311
Daly MB, Pilarski R, Axilbund JE, Buys SS, Crawford B, Friedman S, Garber JE, Horton C, Kaklamani V, Klein C, Kohlmann W, Kurian A, Litton J, Madlensky L, Marcom PK, Merajver SD, Offit K, Pal T, Pasche B, Reiser G, Shannon KM, Swisher E, Voian NC, Weitzel JN, Whelan A, Wiesner GL, Dwyer MA, Kumar R (2014) Genetic/familial high-risk assessment: breast and ovarian, version 1. 2014. J Natl Compr Cancer Netw 12:1326–1338
Desmet FO, Hamroun D, Lalande M, Collod-Beroud G, Claustres M, Beroud C (2009) Human splicing finder: an online bioinformatics tool to predict splicing signals. Nucleic Acids Res 37:e67
Edge SB, Byrd DR, Compton CC (2009) AJCC cancer staging manual, 7th edn. Springer, Chicago
Fong PC, Boss DS, Yap TA, Tutt A, Wu P, Mergui-Roelvink M, Mortimer P, Swaisland H, Lau A, O’Connor MJ, Ashworth A, Carmichael J, Kaye SB, Schellens JH, de Bono JS (2009) Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N Engl J Med 361:123–134
Ford D, Easton DF, Bishop DT, Narod SA, Goldgar DE (1994) Risks of cancer in BRCA1-mutation carriers. Breast cancer linkage consortium. Lancet 343:692–695
Ford D, Easton DF, Stratton M, Narod S, Goldgar D, Devilee P, Bishop DT, Weber B, Lenoir G, Chang-Claude J, Sobol H, Teare MD, Struewing J, Arason A, Scherneck S, Peto J, Rebbeck TR, Tonin P, Neuhausen S, Barkardottir R, Eyfjord J, Lynch H, Ponder BA, Gayther SA, Zelada-Hedman M (1998) Genetic heterogeneity and penetrance analysis of the BRCA1 and BRCA2 genes in breast cancer families. The breast cancer linkage consortium. Am J Hum Genet 62:676–689
Foulkes WD, Shuen AY (2013) In brief: BRCA1 and BRCA2. J Pathol 230:347–349
Foulkes WD, Smith IE, Reis-Filho JS (2010) Triple-negative breast cancer. N Engl J Med 363:1938–1948
Gadzicki D, Evans DG, Harris H, Julian-Reynier C, Nippert I, Schmidtke J, Tibben A, van Asperen CJ, Schlegelberger B (2011) Genetic testing for familial/hereditary breast cancer—comparison of guidelines and recommendations from the UK, France, the Netherlands and Germany. J Community Genet 2:53–69
Gonzalez-Angulo AM, Timms KM, Liu S, Chen H, Litton JK, Potter J, Lanchbury JS, Stemke-Hale K, Hennessy BT, Arun BK, Hortobagyi GN, Do KA, Mills GB, Meric-Bernstam F (2011) Incidence and outcome of BRCA mutations in unselected patients with triple receptor-negative breast cancer. Clin Cancer Res 17:1082–1089
Hammond ME, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S, Fitzgibbons PL, Francis G, Goldstein NS, Hayes M, Hicks DG, Lester S, Love R, Mangu PB, McShane L, Miller K, Osborne CK, Paik S, Perlmutter J, Rhodes A, Sasano H, Schwartz JN, Sweep FC, Taube S, Torlakovic EE, Valenstein P, Viale G, Visscher D, Wheeler T, Williams RB, Wittliff JL, Wolff AC (2010) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol 28:2784–2795
Hartman AR, Kaldate RR, Sailer LM, Painter L, Grier CE, Endsley RR, Griffin M, Hamilton SA, Frye CA, Silberman MA, Wenstrup RJ, Sandbach JF (2012) Prevalence of BRCA mutations in an unselected population of triple-negative breast cancer. Cancer 118:2787–2795
Irvin WJ Jr, Carey LA (2008) What is triple-negative breast cancer? Eur J Cancer 44:2799–2805
Karchin R, Agarwal M, Sali A, Couch F, Beattie MS (2008) Classifying variants of undetermined significance in BRCA2 with protein likelihood ratios. Cancer Inform 6:203–216
King MC, Marks JH, Mandell JB (2003) Breast and ovarian cancer risks due to inherited mutations in BRCA1 and BRCA2. Science 302:643–646
Knies K, Schuster B, Ameziane N, Rooimans M, Bettecken T, de Winter J, Schindler D (2012) Genotyping of fanconi anemia patients by whole exome sequencing: advantages and challenges. PLoS One 7:e52648
Kumar P, Henikoff S, Ng PC (2009) Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 4:1073–1081
Mayer IA, Abramson VG, Lehmann BD, Pietenpol JA (2014) New strategies for triple-negative breast cancer–deciphering the heterogeneity. Clin Cancer Res 20:782–790
Meindl A (2002) Comprehensive analysis of 989 patients with breast or ovarian cancer provides BRCA1 and BRCA2 mutation profiles and frequencies for the German population. Int J Cancer 97:472–480
Mersch J, Jackson MA, Park M, Nebgen D, Peterson SK, Singletary C, Arun BK, Litton JK (2015) Cancers associated with BRCA1 and BRCA2 mutations other than breast and ovarian. Cancer 121:269–275
Meyer P, Landgraf K, Hogel B, Eiermann W, Ataseven B (2012) BRCA2 mutations and triple-negative breast cancer. PLoS One 7:e38361
Pal SK, Childs BH, Pegram M (2011) Triple negative breast cancer: unmet medical needs. Breast Cancer Res Treat 125:627–636
Pern F, Bogdanova N, Schurmann P, Lin M, Ay A, Langer F, Hillemanns P, Christiansen H, Park-Simon TW, Dork T (2012) Mutation analysis of BRCA1, BRCA2, PALB2 and BRD7 in a hospital-based series of German patients with triple-negative breast cancer. PLoS One 7:e47993
Reese MG, Eeckman FH, Kulp D, Haussler D (1997) Improved splice site detection in Genie. J Comput Biol 4:311–323
Rummel S, Varner E, Shriver CD, Ellsworth RE (2013) Evaluation of BRCA1 mutations in an unselected patient population with triple-negative breast cancer. Breast Cancer Res Treat 137:119–125
Singer CF, Tea MK, Pristauz G, Hubalek M, Rappaport C, Riedl C, Helbich T (2012) Guideline for the prevention and early detection of breast and ovarian cancer in high risk patients, particularly in women from HBOC (hereditary breast and ovarian cancer) families. Wien Klin Wochenschr 124:334–339
Stegel V, Krajc M, Zgajnar J, Teugels E, De Greve J, Hocevar M, Novakovic S (2011) The occurrence of germline BRCA1 and BRCA2 sequence alterations in Slovenian population. BMC Med Genet 12:9
Telli M (2014) Optimizing chemotherapy in triple-negative breast cancer: the role of platinum. Am Soc Clin Oncol Educ Book. e37–e42
Tun NM, Villani G, Ong K, Yoe L, Bo ZM (2014) Risk of having BRCA1 mutation in high-risk women with triple-negative breast cancer: a meta-analysis. Clin Genet 85:43–48
Wolff AC, Hammond ME, Schwartz JN, Hagerty KL, Allred DC, Cote RJ, Dowsett M, Fitzgibbons PL, Hanna WM, Langer A, McShane LM, Paik S, Pegram MD, Perez EA, Press MF, Rhodes A, Sturgeon C, Taube SE, Tubbs R, Vance GH, van de Vijver M, Wheeler TM, Hayes DF (2007) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 25:118–145
Wong-Brown MW, Meldrum CJ, Carpenter JE, Clarke CL, Narod SA, Jakubowska A, Rudnicka H, Lubinski J, Scott RJ (2015) Prevalence of BRCA1 and BRCA2 germline mutations in patients with triple-negative breast cancer. Breast Cancer Res Treat 150:71–80
Acknowledgments
We are grateful to Mag. Gabriela Dür and the Vorarlberger Landesregierung (Bregenz, Austria), to Franz Rauch and the Vorarlberger Industriellenvereinigung (Bregenz, Austria), to Dr. Peter Woess and the Vorarlberger Aerztekammer (Dornbirn, Austria), as well as to Dr. Peter Fraunberger and the Institute for Clinical Chemistry at the Academic Teaching Hospital Feldkirch (Feldkirch, Austria) for continuously supporting our research institute. The present study was partly financed by the European Union’s European Regional Development Fund through the INTERREG IV Programme “Alpenrhein-Bodensee-Hochrhein,” Project Number: 179.
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical standard
All procedures performed in this study involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Axel Muendlein and Bettina H. Rohde are first authors and have contributed equally to this article.
Thomas Decker and Alois H. Lang are senior authors and have contributed equally to this article.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Muendlein, A., Rohde, B.H., Gasser, K. et al. Evaluation of BRCA1/2 mutational status among German and Austrian women with triple-negative breast cancer. J Cancer Res Clin Oncol 141, 2005–2012 (2015). https://doi.org/10.1007/s00432-015-1986-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-015-1986-2