Abstract
This study focused on the identification of QTL regions, candidate genes, and network related genes based on the first 3 lactations (LAC3) of milk, fat, and protein yields, and somatic cell score (SCS) in Portuguese Holstein cattle. Additionally, the results were compared with those from only first lactation (LAC1) data. The analyses were performed using the weighted single-step GWAS under an autoregressive test-day (TD) multiple lactations model. A total of 11,434,294 and 4,725,673 TD records from LAC3 and LAC1, respectively, including 38,323 autosomal SNPs and 1338 genotyped animals were used in GWAS analyses. A total of 51 (milk), 5 (fat), 24 (protein), and 4 (SCS) genes were associated to previously annotated relevant QTL regions for LAC3. The CACNA2D1 at BTA4 explained the highest proportion of genetic variance respectively for milk, fat, and protein yields. For SCS, the TRNAG-CCC at BTA14, MAPK10, and PTPN3 genes, both at BTA6 were considered important candidate genes. The accessed network refined the importance of the reported genes. CACNA2D1 regulates calcium density and activation/inactivation kinetics of calcium transport in the mammary gland; whereas TRNAG-CCC, MAPK10, and PTPN3 are directly involved with inflammatory processes widely derived from mastitis. In conclusion, potential candidate genes (TRNAG-CCC, MAPK10, and PTPN3) associated with somatic cell were highlighted, which further validation studies are needed to clarify its mechanism action in response to mastitis. Moreover, most of the candidate genes identified were present in both (LAC3 and LAC1) for milk, fat and protein yields, except for SCS, in which no candidate genes were shared between LAC3 and LAC1. The larger phenotypic information provided by LAC3 dataset was more effective to identify relevant genes, providing a better understanding of the genetic architecture of these traits over all lactations simultaneously.




Similar content being viewed by others
References
Aguilar I, Misztal I, Johnson DL, Legarra A, Tsuruta S, Lawlor TJ (2010) Hot topic: a unified approach to utilize phenotypic, full pedigree, and genomic information for genetic evaluation of Holstein final score1. J Dairy Sci 93:743–752. https://doi.org/10.3168/jds.2009-2730
Bindea G, Mlecnik B, Hackl H, Charoentong P, Tosolini M, Kirilovsky A, Fridman W, Pagès F, Trajanoski Z, Galon J (2009) ClueGO : a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25:1091–1093. https://doi.org/10.1093/bioinformatics/btp101
Bindea G, Galon J, Mlecnik B (2013) Systems biology CluePedia Cytoscape plugin : pathway insights using integrated experimental and in silico data. Bioinformatics 29:661–663. https://doi.org/10.1093/bioinformatics/btt019
Brown EL, Below JE, Fischer RSB, Essigmann HT, Hu H, Huff C, Robinson DA, Petty LE, Aguilar D, Bell GI, Hanis CL (2015) Genome-wide association study of Staphylococcus aureus carriage in a community-based sample of Mexican- Americans in Starr County , Texas. PLoS One 1–28. https://doi.org/10.1371/journal.pone.0142130
Carvalheira J, Pollak EJ, Quaas RL, Blake RW (2002) An autoregressive repeatability animal model for test-day records in multiple lactations. J Dairy Sci 85:2040–2045. https://doi.org/10.3168/jds.S0022-0302(02)74281-1
Carvalheira J, Salem MMI, Thompson G, Chen SY, Beja-Pereira A (2014) Genome-wide association study for milk and protein yields in Portuguese Holstein cattle. In: proceedings, 10th world congress of genetics applied to livestock production. pp 1–3
Chen C, Lanz RB, Walkey CJ, Chang W-H, Lu W, Johnson DL (2018) Maf1 and repression of RNA polymerase III-mediated transcription Crive Adipocyte differenctiation. Cell Rep 24:1852–1864. https://doi.org/10.1016/j.celrep.2018.07.046.Maf1
Christensen OF, Lund MS (2010) Genomic prediction when some animals are not genotyped. Genet Sel Evol 42:1–8
Do DN, Schenkel FS, Miglior F, Zhao X, Ibeagha-Awemu EM (2018) Genome wide association study identifies novel potential candidate genes for bovine milk cholesterol content. Scientific Reports 8:13239. https://doi.org/10.1038/s41598-018-31427-0
Frąszczak M, Szyda J (2016) Comparison of significant single nucleotide polymorphisms selections in GWAS for complex traits. J Appl Genet 57:207–213. https://doi.org/10.1007/s13353-015-0305-6
Frąszczak M, Suchocki T, Szyda J (2016) Utilization of information from gene networks towards a better understanding of functional similarities between complex traits : a dairy cattle model. J Appl Genet 57:129–133. https://doi.org/10.1007/s13353-015-0306-5
Gabashvili IS, Sokolowski BHA, Morton CC, Giersch ABS (2007) Ion channel gene expression in the inner ear. JARO - J Assoc Res Otolaryngol 8:305–328. https://doi.org/10.1007/s10162-007-0082-y
Grisart BG, Coppieters W, Farnir F, Karim L, Ford C, Berzi P, Cambisano N, Mni M, Reid S, Simon P, Spelman R, Georges M, Snell R (2002) Positional candidate cloning of a QTL in dairy cattle: Identification of a missense mutation in the bovine DGAT1 gene with major effect on milk yield and composition. Genome Res. 12:222–231. https://doi.org/10.1101/gr.224202
Hamashima K, Fujishima K, Masuda T, Sugahara J, Tomita M, Kanai A (2012) Nematode-specific tRNAs that decode an alternative genetic code for leucine. Nucleic Acids Res 40:3653–3662. https://doi.org/10.1093/nar/gkr1226
Iung LHS, Petrini J, Ramírez-Díaz J, Salvian M, Rovadoscki GA, Pilonetto F, Dauria BD, Machado PF, Coutinho LL, Wiggans GR, Mourão GB (2019) Genome-wide association study for milk production traits in a Brazilian Holstein population. J Dairy Sci 1:1–10. https://doi.org/10.3168/jds.2018-14811
Lu H, Bovenhuis H (2019) Genome-wide association studies for genetic effects that change during lactation in dairy cattle. J Dairy Sci 102. https://doi.org/10.3168/jds.2018-15994
Magotra A, Gupta ID, Verma A, Chaudhari M V, Arya A, Vineeth MR, Kumar R, Selvan AS (2017) Characterization and validation of point mutation in exon 19 of CACNA2D1 gene in Karan Fries (Bos taurus × Bos indicus) cattle. Indian J Anim Res 51:227–230. doi: https://doi.org/10.18805/ijar.5668
Magotra A, Gupta ID, Verma A, Alex R, Vineeth M, Ahmad T (2018) Candidate SNP of CACNA2D1 gene associated with clinical mastitis and production traits in Sahiwal (Bos taurus indicus) and Karan Fries (Bos taurus taurus × Bos taurus indicus). Anim Biotechnol 0:1–7
Misztal I, Legarra A, Aguilar I (2009) Computing procedures for genetic evaluation including phenotypic, full pedigree, and genomic information. J Dairy Sci 92:4648–4655. https://doi.org/10.3168/jds.2009-2064
Misztal I, Tsuruta S, Lourenco D, Aguilar I, Legarra A, Vitezica Z (2015) Manual for BLUPF90 family of programs
Nayeri S, Sargolzaei M, Abo-ismail MK, May N, Miller SP, Schenkel F, Moore SS, Stothard P (2016) Genome-wide association for milk production and female fertility traits in Canadian dairy Holstein cattle. BMC Genet:1–11. https://doi.org/10.1186/s12863-016-0386-1
Netzer N, Goodenbour JM, David A, Dittmar KA, Jones RB, Schneider JR, Boone D, Eves EM, Rosner MR, Gibbs JS, Embry A, Dolan B, Das S, Hickman HD, Berglund P, Bennink JR, Yewdell JW, Pan T (2009) Innate immune and chemically triggered oxidative stress modifies translational fidelity. Nature 462:522–526. https://doi.org/10.1038/nature08576
Oliveira HR, Cant JP, Brito LF, Feitosa FLB, Chud TCS, Fonseca PAS, Jamrozik J, Silva FF, Lourenco DAL, Schenkel FS (2019) Genome-wide association for milk production traits and somatic cell score in different lactation stages of Ayrshire , Holstein , and Jersey dairy cattle. J Dairy Sci 102:8159–8174. doi: https://doi.org/10.3168/jds.2019-16451
Pilecka I, Patrignani C, Pescini R, Curchod M, Perrin D, Xue Y, Yasenchak J, Clark A, Magnone MC, Zaratin P, Valenzuela D, Rommel C, Huijsduijnen HR (2007) Protein-tyrosine phosphatase H1 controls growth hormone receptor signaling and systemic growth * □. J Biol Chem 282:35405–35415. https://doi.org/10.1074/jbc.M705814200
Qanbari S, Pimentel ECG, Tetens J, Thaller G, Lichtner P, Sharifi AR, Simianer H (2010) The pattern of linkage disequilibrium in German Holstein cattle. Anim Genet:346–356. https://doi.org/10.1111/j.1365-2052.2009.02011.x
Ryman VE, Packiriswamy N, Sordillo LM (2015) Role of endothelial cells in bovine mammary gland health and disease. Anim Health Res Rev 16:135–149. https://doi.org/10.1017/S1466252315000158
Salem MMI, Thompson G, Chen S, Beja-Pereira A, Carvalheira J (2018) Linkage disequilibrium and haplotype block structure in Portuguese Holstein cattle. Czech J Anim Sci 63:61–69. Doi: https://doi.org/10.17221/56/2017-CJAS
Schaeffer LR, Jamrozik J, Kistemaker GJ, Van Doormaal BJ (2000) Experience with a test-day model. J Dairy Sci 83:1135–1144. https://doi.org/10.3168/jds.S0022-0302(00)74979-4
Silva AA, Silva DA, Silva FF, Costa CN, Lopes PS, Caetano AR, Thompson G, Carvalheira J (2019a) Autoregressive single-step test-day model for genomic evaluations of Portuguese Holstein cattle. J Dairy Sci:1–10. https://doi.org/10.3168/jds.2018-15191
Silva AA, Silva FF, Silva DA, Silva HT, Costa CN, Lopes PS, Veroneze R, Thompson G, Carvalheira J (2019b) Genotype imputation strategies for Portuguese Holstein cattle using different SNP panels. Czech J Anim Sci 64:377–386. doi: doi.org/10.17221/120/2019-CJAS
Szyda J, Suchocki T, Qanbari S, Liu Z, Simianer H (2017) Assessing the degree of stratification between closely related Holstein-Friesian populations. J Appl Genet 58:521–526. https://doi.org/10.1007/s13353-017-0409-2
VanHouten JN, Neville MC, Wysolmerski JJ (2007) The calcium-sensing receptor regulates plasma membrane calcium adenosine triphosphatase isoform 2 activity in mammary epithelial cells: a mechanism for calcium-regulated calcium transport into milk. Endocrinology 148:5943–5954. https://doi.org/10.1210/en.2007-0850
VanRaden PM (2008) Efficient methods to compute genomic predictions. J Dairy Sci 91:4414–4423. https://doi.org/10.3168/jds.2007-0980
Viitala S, Szyda J, Blott S, Schulman N, Lidauer M, Mäki-Tanila A, Georges M, Vilkki J (2006) The role of the bovine growth hormone receptor and prolactin receptor genes in milk, fat and protein production in Finnish Ayrshire dairy cattle. Genetics 173:2151–64. https://doi.org/10.1534/genetics.105.046730
Wang H, Misztal I, Aguilar I, Legarra A, Muir WM (2012) Genome-wide association mapping including phenotypes from relatives without genotypes. Genet Res (Camb) 94:73–83. https://doi.org/10.1017/S0016672312000274
Wang H, Misztal I, Aguilar I, Legarra A, Fernando RL, Vitezica Z, Okimoto R, Wing T, Hawken R, Muir WM (2014) Genome-wide association mapping including phenotypes from relatives without genotypes in a single-step (ssGWAS) for 6-week body weight in broiler chickens. Front Genet 5
Wu X, Lund MS, Sahana G, Guldbrandtsen B, Sun D, Zhang Q, Su G (2015) Association analysis for udder health based on SNP-panel and sequence data in Danish Holsteins. Genet Sel Evol 47:1–14. https://doi.org/10.1186/s12711-015-0129-1
Yue SJ, Zhao YQ, Gu XR, Yin B, Jiang YL, Wang ZH, Shi KR (2017) A genome-wide association study suggests new candidate genes for milk production traits in Chinese Holstein cattle. Anim Genet 48:677–681. https://doi.org/10.1111/age.12593
Zhou C, Li C, Cai W, Liu S, Yin H, Shi S, Zhang Q, Zhang S (2019) Genome-wide association study for milk protein composition traits in a Chinese Holstein population using a single-step approach. Front Genet 10:1–17. https://doi.org/10.3389/fgene.2019.00072
Acknowledgments
The authors acknowledge Portuguese Dairy Cattle Breeders Association (ANABLE) and Embrapa Dairy Cattle for providing the used data. We also thank Dr. Hinayah R. Oliveira (Department of Animal Sciences, Purdue University) for her valuable and constructive suggestions.
Funding
This study was partially financed by CAPES/FCT (99999.008462/2014-03) and CNPq/INCT-CA.
Author information
Authors and Affiliations
Contributions
AA Silva designed the study, performed the statistical analysis, interpreted the results, and wrote the manuscript. DA Silva, HT Silva, CN Costa, PS Lopes, R Veroneze, and G Thompson interpreted the results and revised the manuscript. J Carvalheira and FF Silva supervised all stages of this work. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed. The approval’s register of the Ethics Committee on the Use of Animal at the Universidade Federal de Viçosa is 001/2017.
Additional information
Communicated by: Maciej Szydlowski
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Fig. S1
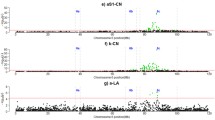
GWAS results of milk production traits for the first lactation (LAC1). a milk yield, and b fat yield. Each dot represents one SNP window of 100 kb. On the y-axis is the chromosomes (Chr) (PDF 192 kb)
Fig. S2
GWAS results of milk production traits for the first lactation (LAC1). a protein yield, and b somatic cell score (SCS). Each dot represents one SNP window of 100 kb. On the y-axis is the chromosomes (Chr) (PDF 250 kb)
Rights and permissions
About this article
Cite this article
Silva, A.A., Silva, D.A., Silva, F.F. et al. GWAS and gene networks for milk-related traits from test-day multiple lactations in Portuguese Holstein cattle. J Appl Genetics 61, 465–476 (2020). https://doi.org/10.1007/s13353-020-00567-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-020-00567-3