Abstract
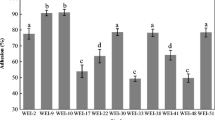
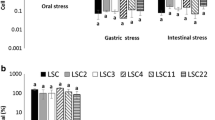
Lactic acid bacteria (LAB) are a well-used probiotics for health improvements in both humans and animals. Despite of several benefits, non-host-specific LAB showed poor probiotics effects due to difficulty in colonization and competition with normal flora. Therefore, the feasibility of porcine LAB isolates was evaluated as a probiotics. Ten of 49 Lactobacillus spp. isolates harbored 2∼10 kb plasmid DNA. Seven strains were selected based on the safety test, such as hemolytic activity, ammonia, indole, and phenylalanine production. After safety test, five strains were selected again by several tests, such as epithelial adherence, antimicrobial activity, tolerance against acid, bile, heat, and cold-drying, and production of acid and hydrogen peroxide. Then, enzyme profiles (ZYM test) and antibiotics resistance were analyzed for further characterization. Five Lactobacillus reuteri isolates from pig feces were selected by safety and functional tests. The plasmid DNA which was able to develop vector system was detected in the isolates. Together with these approaches, pig-specific Lactobacillus spp. originated from pigs were selected. These strains may be useful tools to develop oral delivery system.
Similar content being viewed by others
References
Adams, M.R. and P. Marteau. 1995. On the safety of lactic acid bacteria from food. Int. J. Food Microbiol. 27, 263–264.
Almstahl, A. and M. Wikstrom. 2005. Microflora in oral ecosystems in subjects with hyposalivation due to medicines or of unknown origin. Oral Health Prev. Dent. 3, 67–76.
Amoa-Awua, W.K., F.E. Appoh, and M. Jakobsen. 1996. Lactic acid fermentation of cassava dough into agbelima. Int. J. Food Microbiol. 31, 87–98.
Ashenafi, M. and M. Busse. 1991. Growth of Bacillus cereus in fermenting tempeh made from various beans and its inhibition by Lactobacillus plantarum. J. Appl. Bacteriol. 70, 329–333.
Axelsson, L.T., T.C. Chung, W.G. Dobrogosz, and S.E. Lindgren. 1989. Production of a broad spectrum antimicrobial substances by Lactobacillus reuteri. Microb. Ecol. Health Dis. 2, 131–136.
Aymerich, T., B. Martin, M. Garriga, M.C. Vidal-Carou, S. Bover-Cid, and M. Hugas. 2006. Safety properties and molecular strain typing of lactic acid bacteria from slightly fermented sausages. J. Appl. Microbiol. 100, 40–49.
Berg, R.D. 1998. Probiotics, prebiotics or ‘conbiotics’? Trends Microbiol. 6, 89–92.
Cappucino, J.G. and N. Sherman. 1987. Urease test, p. 157–160. Microbiology: a laboratory manual. Bejamings/Cummings Publishing Co. Inc., Menlo park, CA, USA.
Chauviere, G., M.H. Coconnier, S. Kerneis, J. Fourniat, and A.L. Servin. 1992. Adhesion of human Lactobacillus acidophilus strain LB to human enterocyte-like Caco-2 cells. J. Gen. Microbiol. 138(Pt 8), 1689–1696.
Choi, S.S., B.Y. Kang, M.J. Chung, S.D. Kim, S.H. Park, J.S. Kim, C.Y. Kang, and N.J. Ha. 2005. Safety assessment of potential lactic acid bacteria Bifidobacterium longum SPM1205 isolated from healthy Koreans. J. Microbiol. 43, 493–498.
Coconnier, M.H., V. Lievin, M.F. Bernet-Camard, S. Hudault, and A.L. Servin. 1997. Antibacterial effect of the adhering human Lactobacillus acidophilus strain LB. Antimicrob. Agents Chemother. 41, 1046–1052.
Codon, S. 1983. Aerobic metabolism of lactic acid bacteria. Ir. J. Food Sci. Technol. 7, 15–25.
Delgado, S., A. Suarez, and B. Mayo. 2006. Identification of dominant bacteria in feces and colonic mucosa from healthy Spanish adults by culturing and by 16S rDNA sequence analysis. Dig. Dis. Sci. 51, 744–751.
du Plessis, H.W., L.M. Dicks, I.S. Pretorius, M.G. Lambrechts, and M. du Toit. 2004. Identification of lactic acid bacteria isolated from South African brandy base wines. Int. J. Food Microbiol. 91, 19–29.
El-Ziney, M.G. and J.M. Debevere. 1998. The effect of Reuterin on Listeria monocytogenes and Escherichia coli O157:H7 in milk and cottage cheese. J. Food Prot. 61, 1275–1280.
Flint, J.F. and E.R. Angert. 2005. Development of a strain-specific assay for detection of viable Lactobacillus spp. HOFG1 after application to cattle feed. J. Microbiol. Methods 61, 235–243.
Florez, A.B., S. Delgado, and B. Mayo. 2005. Antimicrobial susceptibility of lactic acid bacteria isolated from a cheese environment. Can. J. Microbiol. 51, 51–58.
Gevers, D., M. Danielsen, G. Huys, and J. Swings. 2003a. Molecular characterization of tet(M) genes in Lactobacillus isolates from different types of fermented dry sausage. Appl. Environ. Microbiol. 69, 1270–1275.
Gevers, D., G. Huys, and J. Swings. 2003b. In vitro conjugal transfer of tetracycline resistance from Lactobacillus isolates to other Gram-positive bacteria. FEMS Microbiol. Lett. 225, 125–130.
Haddadin, M.S.Y., S.S. Awaisheh, and R.K. Robinson. 2004. The production of yogurt with probiotic bacteria isolated from infants in Jordan. Pakistan J. Nutr. 3, 290–293.
Hudault, S., V. Lievin, M.F. Bernet-Camard, and A.L. Servin. 1997. Antagonistic activity exerted in vitro and in vivo by Lactobacillus casei (strain GG) against Salmonella typhimurium C5 infection. Appl. Environ. Microbiol. 63, 513–518.
Isenberg, H.D. 1992. Indole test/Urease. Clinical Microbiology Procedures Handbook. American Society of Microbiology, Washington DC, USA. 1, 1.19.13–11.19.15 / 12.16.18.
Ishibashi, N. and S. Yamazaki. 2001. Probiotics and safety. Am. J. Clin. Nutr. 73, 465S–470S.
Jacobsen, C.N., V. Rosenfeldt Nielsen, A.E. Hayford, P.L. Moller, K.F. Michaelsen, A. Paerregaard, B. Sandstrom, M. Tvede, and M. Jakobsen. 1999. Screening of probiotic activities of forty-seven strains of Lactobacillus spp. by in vitro techniques and evaluation of the colonization ability of five selected strains in humans. Appl. Environ. Microbiol. 65, 4949–4956.
Juven, B.J., F. Schved, and P. Lindner. 1992. Antagonistic compounds produced by a chicken intestinal strains of Lactobacillus acidophilius. J. Food Prot. 55, 157–161.
Kastner, S., V. Perreten, H. Bleuler, G. Hugenschmidt, C. Lacroix, and L. Meile. 2006. Antibiotic susceptibility patterns and resistance genes of starter cultures and probiotic bacteria used in food. Syst. Appl. Microbiol. 29, 145–155.
Kim, K.S., J.O. Morrison, and A.S. Bayer. 1982. Deficient autolytic enzyme activity in antibiotic-tolerant lactobacilli. Infect. Immun. 36, 582–585.
Klein, G., C. Hallmann, I.A. Casas, J. Abad, J. Louwers, and G. Reuter. 2000. Exclusion of vanA, vanB and vanC type glycopeptide resistance in strains of Lactobacillus reuteri and Lactobacillus rhamnosus used as probiotics by polymerase chain reaction and hybridization methods. J. Appl. Microbiol. 89, 815–824.
Lee, D.Y., S.G. Kang, N. Rayamajhi, M.L. Kang, and H.S. Yoo. 2008. Analysis of nucleotide sequence of a novel plasmid, pILR091, from Lactobacillus reuteri L09 isolated from pig. Korean J. Vet. Res. 48, 441–449.
Makelainen, H., R. Tahvonen, S. Salminen, and A.C. Ouwehand. 2003. In vivo safety assessment of two Bifidobacterium longum strains. Microbiol. Immunol. 47, 911–914.
Marteau, P.R. 2002. Probiotics in clinical conditions. Clin. Rev. Aller. Immunol. 22, 255–273.
Mastromarino, A., B.S. Reddy, and E.L. Wynder. 1976. Metabolic epidemiology of colon cancer: enzymic activity of fecal flora. Am. J. Clin. Nutr. 29, 1455–1460.
Mayo, B., C. Hardisson, and A.F. Brana. 1989. Selected characteristics of several strains of Lactobacillus plantarum. Microbiologia 5, 105–112.
Menendez, S., J.A. Centeno, R. Godinez, and J.L. Rodriguez-Otero. 2000. Effects of Lactobacillus strains on the ripening and organoleptic characteristics of Arzua-Ulloa cheese. Int. J. Food Microbiol. 59, 37–46.
Mercenier, A., H. Muller-Alouf, and C. Grangette. 2000. Lactic acid bacteria as live vaccines. Curr. Iss. Mol. Biol. 2, 17–25.
Miele, A., M. Bandera, and B.P. Goldstein. 1995. Use of primers selective for vancomycin resistance genes to determine van genotype in enterococci and to study gene organization in VanA isolates. Antimicrob. Agents Chemother. 39, 1772–1778.
Nigatu, A. 2000. Evaluation of numerical analyses of RAPD and API 50 CH patterns to differentiate Lactobacillus plantarum, Lact. fermentum, Lact. rhamnosus, Lact. sake, Lact. parabuchneri, Lact. gallinarum, Lact. casei, Weissella minor and related taxa isolated from kocho and tef. J. Appl. Microbiol. 89, 969–978.
Organization., F.a.A.O.o.t.U.N.a.W. 2002. Guidelines for the evalu ation of probiotics in food. Joint FAO and WHO Working Group Report London, Ontario, Canada April 30 and May 1.
O’sullivan, D.J. and T.R. Klaenhammer. 1993. Rapid mini-prep isolation of high-quality plasmid DNA from Lactococcus and Lactobacillus spp. Appl. Environ. Microbiol. 59, 2730–2733.
Pryde, S.E., A.J. Richardson, C.S. Stewart, and H.J. Flint. 1999. Molecular analysis of the microbial diversity present in the colonic wall, colonic lumen, and cecal lumen of a pig. Appl. Environ. Microbiol. 65, 5372–5377.
Putman, M., H.W. van Veen, J.E. Degener, and W.N. Konings. 2001. The lactococcal secondary multidrug transporter LmrP confers resistance to lincosamides, macrolides, streptogramins and tetracyclines. Microbiology 147, 2873–2880.
Rafii, F., R. Wynne, T.M. Heinze, and D.D. Paine. 2003. Mechanism of metronidazole-resistance by isolates of nitroreductase-producing Enterococcus gallinarum and Enterococcus casseliflavus from the human intestinal tract. FEMS Microbiol. Lett. 225, 195–200.
Rautio, M., H. Jousimies-Somer, H. Kauma, I. Pietarinen, M. Saxelin, S. Tynkkynen, and M. Koskela. 1999. Liver abscess due to a Lactobacillus rhamnosus strain indistinguishable from L.trhamnosus strain GG. Clin. Infect. Dis. 28, 1159–1160.
Reid, G. 2002. Safety of lactobacillus strains as probiotic agents. Clin. Infect. Dis. 35, 349–350.
Reid, G., J. Jass, M.T. Sebulsky, and J.K. McCormick. 2003. Potential uses of probiotics in clinical practice. Clin. Microbiol. Rev. 16, 658–672.
Ruiz-Barba, J.L., J.C. Piard, and R. Jimenez-Diaz. 1991. Plasmid profiles and curing of plasmids in Lactobacillus plantarum strains isolated from green olive fermentations. J. Appl. Bacteriol. 71, 417–421.
Sandine, W.E. 1979. Role of Lactobacillus in the intestinal tract. J. Food Prot. 42, 259–262.
Sarem, F., L.O. Sarem-Damerdji, and J.P. Nicolas. 1996. Comparison of the adherence of three Lactobacillus strains to Caco-2 and Int-407 human intestinal cell lines. Lett. Appl. Microbiol. 22, 439–442.
Tuomola, E.M. and S.J. Salminen. 1998. Adhesion of some probiotic and dairy Lactobacillus strains to Caco-2 cell cultures. Int. J. Food Microbiol. 41, 45–51.
Tynkkynen, S., R. Satokari, M. Saarela, T. Mattila-Sandholm, and M. Saxelin. 1999. Comparison of ribotyping, randomly amplified polymorphic DNA analysis, and pulsed-field gel electrophoresis in typing of Lactobacillus rhamnosus and L. casei strains. Appl. Environ. Microbiol. 65, 3908–3914.
Tynkkynen, S., K.V. Singh, and P. Varmanen. 1998. Vancomycin resistance factor of Lactobacillus rhamnosus GG in relation to enterococcal vancomycin resistance (van) genes. Int. J. Food Microbiol. 41, 195–204.
Yeung, P.S., C.L. Kitts, R. Cano, P.S. Tong, and M.E. Sanders. 2004. Application of genotypic and phenotypic analyses to commercial probiotic strain identity and relatedness. J. Appl. Microbiol. 97, 1095–1104.
Yeung, P.S., M.E. Sanders, C.L. Kitts, R. Cano, and P.S. Tong. 2002. Species-specific identification of commercial probiotic strains. J. Dairy Sci. 85, 1039–1051.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lee, D.Y., Seo, YS., Rayamajhi, N. et al. Isolation, characterization, and evaluation of wild isolates of Lactobacillus reuteri from pig feces. J Microbiol. 47, 663–672 (2009). https://doi.org/10.1007/s12275-009-0124-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-009-0124-8