Abstract
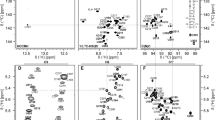
The dysregulation of translation contributes to many pathogenic conditions in humans. Discovering new translational mechanisms is important to understanding the diversity of this process and its potential mechanisms. Such mechanisms can be initially observed in viruses. With this in mind, we studied the viral protein genome-linked VPg factor from the largest genus of plant viruses. Studies in plants show that VPg binds to the eukaryotic translation initiation factor eIF4E for translation of viral RNAs. VPg contains no known eIF4E binding motifs and no sequence homology to any known proteins. Thus, as a first step in understanding the structural basis of this interaction, we carried out NMR assignments of the VPg from the potato virus Y potyvirus protein.


Similar content being viewed by others
References
Bastet A, Robaglia C, Gallois J (2017) eIF4E resistance: natural variation should guide gene editing. Trends Plant Sci 22:411–419
Carroll M, Borden KL (2013) The oncogene eIF4E: using biochemical insights to target cancer. J Interferon Cytokine Res 33:227–238
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
German-Retana S et al (2008) Mutational analysis of plant cap-binding protein eIF4E reveals key amino acids involved in biochemical functions and potyvirus infection. J Virol 82:7601–7612
Goddard TD, Kneller DG (2003) Sparky 3. University of California, San Francisco
Hyberts SG, Takeuchi K, Wagner G (2010) Poisson-gap sampling and forward maximum entropy reconstruction for enhancing the resolution and sensitivity of protein NMR data. J Am Chem Soc 132:2145–2147
Kentsis A, Dwyer EC, Perez JM, Sharma M, Chen A, Pan ZQ, Borden KL (2001) The RING domains of the promyelocytic leukemia protein PML and the arenaviral protein Z repress translation by directly inhibiting translation initiation factor eIF4E. J Mol Biol 312:609–623
Osborne MJ, Borden KL (2015) The eukaryotic translation initiation factor eIF4E in the nucleus: taking the road less traveled. Immunol Rev 263:210–223
Peter D et al (2017) GIGYF1/2 proteins use auxiliary sequences to selectively bind to 4EHP and repress target mRNA expression. Genes Dev 31:1147–1161
Robaglia C, Caranta C (2006) Translation initiation factors: a weak link in plant virus infection. Trends Plant Sci 11:40–45
Shen Y, Bax A (2013) Protein backbone and sidechain torsion angles predicted from NMR chemical shifts using artificial neural networks. J Biomol NMR 56:227–241
Tan NG, Ardley HC, Scott GB, Rose SA, Markham AF, Robinson PA (2003) Human homologue of ariadne promotes the ubiquitylation of translation initiation factor 4E homologous protein, 4EHP. FEBS Lett 554:501–504
Topisirovic I, Borden KL (2005) Homeodomain proteins and eukaryotic translation initiation factor 4E (eIF4E): an unexpected relationship. Histol Histopathol 20:1275–1284
Volpon L, Osborne MJ, Capul AA, de la Torre JC, Borden KL (2010) Structural characterization of the Z RING-eIF4E complex reveals a distinct mode of control for eIF4E. Proc Natl Acad Sci USA 107:5441–5446
Vranken WF et al (2005) The CCPN data model for NMR spectroscopy: development of a software pipeline. Proteins 59:687–696
Wang A, Krishnaswamy S (2012) Eukaryotic translation intiation factor 4E-mediated recessive resistance to plant viruses and its utility in crop improvement. Mol Plant Pathol 13:795–803
Acknowledgements
800 MHz NMR data were acquired at QANUC and 600 MHz NMR data at IRIC which were both supported in part by CFI. This work was supported by the following grants to KLBB: RO1 NIH 98571; RO1 NIH80728 and KLBB holds the Canada Research Chair in Molecular Biology of the Cell Nucleus.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Coutinho de Oliveira, L., Volpon, L., Osborne, M.J. et al. Chemical shift assignment of the viral protein genome-linked (VPg) from potato virus Y. Biomol NMR Assign 13, 9–13 (2019). https://doi.org/10.1007/s12104-018-9842-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-018-9842-3