Abstract
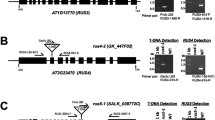
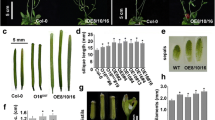
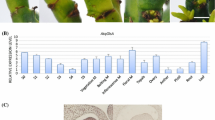
The detailed expression patterns of transcripts of two Arabidopsis arginase genes, ARGAH1 and ARGAH2, have not been previously described, and phylogenetic analysis suggests that they diverged independently of duplication events in other lineages. Therefore, we used β-glucuronidase reporter fusions and quantitative reverse-transcriptase PCR to analyze tissue-specific expression of ARGAH1 and ARGAH2 during Arabidopsis development, and in response to the availability of nutrients and exposure to methyl jasmonate (MeJA). We demonstrated tissue-specific transcript expression and enzyme activity in pollen for ARGAH1, but not ARGAH2. Conversely, we demonstrated MeJA-inducibility of ARGAH2, but not ARGAH1. In addition, we used microarrays to identify genes for which transcript abundance following MeJA treatment differed in wild type and ARGAH2 mutants. These ARGAH2 and MeJA responsive genes included a putative pathogenesis-related protein pathogenesis response-1 (At2g14610), and a gene of unknown function (At5g03090). Interestingly, these genes had opposite responses to the loss of ARGAH2, suggesting multiple downstream effects of arginase activity, following MeJA treatment. These results, and the variety and complexity of expression patterns of ARGAH1 and ARGAH2 transcript expression and their related reporter gene fusions that we observed point to multiple functions of arginase genes in Arabidopsis, some of which have resulted through a sub-functionalization not shared by all angiosperms.









Similar content being viewed by others
Abbreviations
- ARGAH :
-
Arginase (Arabidopsis)
- FDR:
-
False discovery rate
- GUS:
-
Beta-glucuronidase
- LeARG :
-
Arginase (tomato)
- MeJA:
-
Methyl jasmonate
- MS:
-
Murashige and Skoog
- NO:
-
Nitric oxide
- PR:
-
Pathogenesis related
- qRT-PCR:
-
Quantitative reverse-transcriptase PCR
- SA:
-
Salicyclic acid
- SAM:
-
Statistical analysis of microarrays
References
Alabadi D, Aguero MS, Perez-Amador MA et al (1996) Arginase, arginine decarboxylase, ornithine decarboxylase and polyamines in tomato ovaries—changes in unpollinated ovaries and parthenocarpic fruits induced by auxin or gibberellin. Plant Physiol 112:1237–1244
Bate N, Twell D (1998) Functional architecture of a late pollen promoter: pollen-specific transcription is developmentally regulated by multiple stage-specific and co-dependent activator elements. Plant Mol Biol 37:859–869
Boter M, Ruiz-Rivero O, Abdeen A et al (2004) Conserved myc transcription factors play a key role in jasmonate signaling both in tomato and arabidopsis. Gene Dev 18:1577–1591
Buchanan BB, Gruissem W, Jones RS (eds) (2000) Biochemistry and molecular biology of plants. American Society of Plant Biology, Rockville, MD
Chen H, McCaig BC, Melotto M et al (2004) Regulation of plant arginase by wounding, jasmonate, and the phytotoxin coronatine. J Biol Chem 279:45998–46007
Cooke JEK, Brown KA, Wu R et al (2003) Gene expression associated with n-induced shifts in resource allocation in poplar. Plant Cell Environ 26:757–770
Corpas FJ, Barroso JB, Carreras A et al (2006) Constitutive arginine-dependent nitric oxide synthase activity in different organs of pea seedlings during plant development. Planta 224:246–254
Devoto A, Turner JG (2005) Jasmonate-regulated arabidopsis stress signalling network. Physiol Plant 123:161–172
Devoto A, Ellis C, Magusin A et al (2005) Expression profiling reveals coi1 to be a key regulator of genes involved in wound- and methyl jasmonate-induced secondary metabolism, defence, and hormone interactions. Plant Mol Biol 58:497–513
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure from small quantities of fresh leaf tissues. Phytochem Bull 19:11–15
Filichkin SA, Leonard JM, Monteros A et al (2004) A novel endo-beta-mannanase associated with anther and gene in tomato leman5 is pollen development. Plant Physiol 134:1080–1087
Goldraij A, Polacco JC (1999) Arginase is inoperative in developing soybean embryos. Plant Physiol 119:297–303
Higo K, Ugawa Y, Iwamoto M et al (1999) Plant cis-acting regulatory DNA elements (place) database: 1999. Nucleic Acids Res 27:297–300
Hong-Qi Z, Croes AF, Linskens HF (1982) Protein synthesis in germinating pollen of petunia: role of proline. Planta 154:199–203
Honys D, Twell D (2003) Comparative analysis of the arabidopsis pollen transcriptome. Plant Physiol 132:640–652
Jenkinson CP, Grody WW, Cederbaum SD (1996) Comparative properties of arginases. Comp Biochem Physiol 114B:107–132
Jiang YQ, Deyholos MK (2006) Comprehensive transcriptional profiling of NaCl-stressed Arabidopsis roots reveals novel classes of responsive genes. BMC Plant Biol 6:25
King JE, Gifford DJ (1997) Amino acid utilization in seeds of loblolly pine during germination and early seedling growth. Plant Physiol 113:1125–1135
Klessig DF, Malamy J (1994) The salicylic-acid signal in plants. Plant Mol Biol 26:1439–1458
Krumpelman PM, Freyermuth SK, Cannon JF et al (1995) Nucleotide-sequence of arabidopsis-thaliana arginase expressed in yeast. Plant Physiol 107:1479–1480
Lebel E, Heifetz P, Thorne L et al (1998) Functional analysis of regulatory sequences controlling PR-1 gene expression in arabidopsis. Plant J16:223–233
Lescot M, DehaisP, Thijs G et al (2002) Plantcare, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30:325–327
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative pcr and the 2(t)(-delta delta c) method. Methods 25:402–408
Micallef BJ, Shelp BJ (1989a) Arginine metabolism in developing soybean cotyledons. 3. Utilization. Plant Physiol 91:170–174
Micallef BJ, Shelp BJ (1989b) Arginine metabolism in developing soybean cotyledons. 1. Relationship to nitrogen nutrition. Plant Physiol 90:624–630
Palmieri L, Todd CD, Arrigoni R et al (2006) Arabidopsis mitochondria have two basic amino acid transporters with partially overlapping specificities and differential expression in seedling development. Bioenergetics 1757:1277–1283
Perez-Amador MA, Lidder P, Johnson MA et al (2001) New molecular phenotypes in the dst mutants of arabidopsis revealed by DNA microarray analysis. Plant Cell 13:2703–2717
Prestridge DS (1991) Signal scan—a computer-program that scans DNA-sequences for eukaryotic transcriptional elements. Comput Appl Biosci 7:203–206
Rogers HJ, Bate N, Combe J et al (2001) Functional analysis of cis-regulatory elements within the promoter of the tobacco late pollen gene g10. Plant Mol Biol 45:577–585
Rouster J, van Mechelen J, Cameron-Mills V (1998) The untranslated leader sequence of the barley lipoxygenase 1 (lox1) gene confers embryo-specific expression. Plant J 15:435–440
Saeed AI, Sharov V, White J et al (2003) TM4: a free, open-source system for microarray data management and analysis. Biotechniques 34:374–384
Schwacke R, Grallath S, Breitkreuz KE et al (1999) Leprot1, a transporter for proline, glycine betaine, and gamma-amino butyric acid in tomato pollen. Plant Cell 11:377–391
Todd CD, Cooke JEK, Mullen RT et al (2001) Regulation of loblolly pine (Pinus taeda L.) arginase in developing seedling tissue during germination and post-germinative growth. Plant Mol Biol 45:555–565
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 98:5116–5121
Twell D, Yamaguchi J, Wing RA et al (1991) Promoter analysis of genes that are coordinately expressed during pollen development reveals pollen-specific enhancer sequences and shared regulatory elements. Gene Dev 5:496–507
Ward ER, Uknes SJ, Williams SC et al (1991) Coordinate gene activity in response to agents that induce systemic acquired-resistance. Plant Cell 3:1085–1094
Weigel D, Glazebrook J (2002) How to transform Arabidopsis. In: Arabidopsis: A laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY
Zonia LE, Stebbins NE, Polacco JC (1995) Essential role of urease in germination of nitrogen-limited Arabidopsis thaliana seeds. Plant Physiol 107:1097–1103
Acknowledgments
Microarrays were printed at the University of Alberta Department of Biological Sciences by Dr. Anthony Cornish. We thank Mohsen Mohammadi for assistance with phylogenetic software. This research was funded by separate NSERC Discovery Grants to MKD and the late David Gifford.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brownfield, D.L., Todd, C.D. & Deyholos, M.K. Analysis of Arabidopsis arginase gene transcription patterns indicates specific biological functions for recently diverged paralogs. Plant Mol Biol 67, 429–440 (2008). https://doi.org/10.1007/s11103-008-9336-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-008-9336-2