Abstract
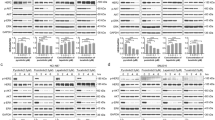
Triple-negative breast cancer (TNBC), which accounts for 10–20% of all breast cancers, has the worst prognosis. Although chemotherapy treatment is a standard for TNBC, it lacks a specific target. Therefore, new therapeutic strategies are required to be investigated. In this study, a combined doxorubicin (DOX) and small interfering RNA (siRNA) therapy is proposed as therapeutic strategy for targeting TGFβ1 gene. Hs578T cell line is used as in vitro model for TNBC, wherein TGFβ1siRNA therapy is employed to enhance therapeutic effects. Cell proliferation rate is measured using an MTT test, and morphological alterations are assed using microscopically approached, while gene expression is determined by qRT-PCR analysis. The combined treatment of TGFβ1siRNA and DOX reduced levels of cell proliferation and mitochondrial activity and promoted the alteration of cell morphology (dark-field microscopy). DOX treatment caused downregulation of six genes and upregulation of another six genes. The combined effects of DOX and TGFβ1siRNA resulted in upregulation of 13 genes and downregulation of four genes. Silencing of TGFβ1 resulted in activation of cell death mechanisms in Hs578T cells, to potentiate the effects of DOX, but not in an additive manner, due to the activation of genes involved in resistance to therapy (ABCB1 and IL-6).






Similar content being viewed by others
Abbreviations
- 2-HER2:
-
Human epidermal growth factor
- ABCB1:
-
ATP-binding cassette sub-family B member 1
- cDNA:
-
Complementary DNA
- DFS:
-
Disease-free survival
- DNA:
-
Deoxyribonucleic acid
- DOX:
-
Doxorubicin
- ER:
-
Estrogen receptor
- FC:
-
Fold change
- IC50:
-
Half maximal inhibitory concentration
- IL-6/IL-8:
-
Interleukin 6/interleukin 8
- MGMT:
-
Methylation of O6-methylguanine-DssNA methyltransferase
- Ml:
-
Milliliter
- μM:
-
Micromolar
- nM:
-
Nanomolar
- OS:
-
Overall survival
- PR:
-
Progesterone receptor
- PTEN:
-
Phosphatase and tensin homolog
- RNAi:
-
RNA interference
- SD:
-
Standard deviation
- siRNA:
-
Small interfering RNA
- TCGA:
-
The cancer genome atlas
- TGFβ1:
-
Transforming growth factor beta 1
- TMRE:
-
Tetra methylrhodamine ethyl ester
- TNBC:
-
Triple-negative breast cancer
References
Wahba HA, El-Hadaad HA (2015) Current approaches in treatment of triple-negative breast cancer. Cancer Biol Med 12:106–116. https://doi.org/10.7497/j.issn.2095-3941.2015.0030
Braicu C, Berindan-Neagoe I, Pileczki V, Cojocneanu-Petric R, Pop LA, Puscas E, Irimie A, Buiga R (2014) Breast tumor bank: an important resource for developing translational cancer research in Romania. Cancer Biomark 14:119–127. https://doi.org/10.3233/cbm-130309
Blows FM, Driver KE, Schmidt MK, Broeks A, van Leeuwen FE, Wesseling J, Cheang MC, Gelmon K, Nielsen TO, Blomqvist C, Heikkila P, Heikkinen T, Nevanlinna H, Akslen LA, Begin LR, Foulkes WD, Couch FJ, Wang X, Cafourek V, Olson JE, Baglietto L, Giles GG, Severi G, McLean CA, Southey MC, Rakha E, Green AR, Ellis IO, Sherman ME, Lissowska J, Anderson WF, Cox A, Cross SS, Reed MW, Provenzano E, Dawson SJ, Dunning AM, Humphreys M, Easton DF, Garcia-Closas M, Caldas C, Pharoah PD, Huntsman D (2010) Subtyping of breast cancer by immunohistochemistry to investigate a relationship between subtype and short and long term survival: a collaborative analysis of data for 10,159 cases from 12 studies. PLoS Med 7:e1000279. https://doi.org/10.1371/journal.pmed.1000279
Braicu C, Chiorean R, Irimie A, Chira S, Tomuleasa C, Neagoe E, Paradiso A, Achimas-Cadariu P, Lazar V, Berindan-Neagoe I (2016) Novel insight into triple-negative breast cancers, the emerging role of angiogenesis, and antiangiogenic therapy. Expert Rev Mol Med 18:e18. https://doi.org/10.1017/erm.2016.17
Chiorean R, Braicu C, Berindan-Neagoe I (2013) Another review on triple negative breast cancer. Are we on the right way towards the exit from the labyrinth? Breast 22:1026–1033. https://doi.org/10.1016/j.breast.2013.08.007
Barnard ME, Boeke CE, Tamimi RM (2015) Established breast cancer risk factors and risk of intrinsic tumor subtypes. Biochim Biophys Acta 1856:73–85. https://doi.org/10.1016/j.bbcan.2015.06.002
Huang J, Luo Q, Xiao Y, Li H, Kong L, Ren G (2017) The implication from RAS/RAF/ERK signaling pathway increased activation in epirubicin treated triple negative breast cancer. Oncotarget 8:108249–108260. https://doi.org/10.18632/oncotarget.22604
Braicu C, Raduly L, Morar-Bolba G, Cojocneanu R, Jurj A, Pop LA, Pileczki V, Ciocan C, Moldovan A, Irimie A, Eniu A, Achimas-Cadariu P, Paradiso A, Berindan-Neagoe I (2018) Aberrant miRNAs expressed in HER-2 negative breast cancers patient. J Exp Clin Cancer Res 37:257. https://doi.org/10.1186/s13046-018-0920-2
Hernandez-Aya LF, Chavez-Macgregor M, Lei X, Meric-Bernstam F, Buchholz TA, Hsu L, Sahin AA, Do K-A, Valero V, Hortobagyi GN, Gonzalez-Angulo AM (2011) Nodal status and clinical outcomes in a large cohort of patients with triple-negative breast cancer. J Clin Oncol 29:2628–2634. https://doi.org/10.1200/JCO.2010.32.1877
Hwa HL, Kuo WH, Chang LY, Wang MY, Tung TH, Chang KJ, Hsieh FJ (2008) Prediction of breast cancer and lymph node metastatic status with tumour markers using logistic regression models. J Eval Clin Pract 14:275–280. https://doi.org/10.1111/j.1365-2753.2007.00849.x
Narrandes S, Huang S, Murphy L, Xu W (2018) The exploration of contrasting pathways in triple negative breast cancer (TNBC). BMC Cancer 18:22. https://doi.org/10.1186/s12885-017-3939-4
Mehanna J, Haddad FG, Eid R, Lambertini M, Kourie HR (2019) Triple-negative breast cancer: current perspective on the evolving therapeutic landscape. Int J Women's Health 11:431–437. https://doi.org/10.2147/IJWH.S178349
Cojocneanu Petric R, Braicu C, Raduly L, Zanoaga O, Dragos N, Monroig P, Dumitrascu D, Berindan-Neagoe I (2015) Phytochemicals modulate carcinogenic signaling pathways in breast and hormone-related cancers. Onco Targets Ther 8:2053–2066. https://doi.org/10.2147/OTT.S83597
Xu F, Wang F, Yang T, Sheng Y, Zhong T, Chen Y (2014) Differential drug resistance acquisition to doxorubicin and paclitaxel in breast cancer cells. Cancer Cell Int 14:142–142. https://doi.org/10.1186/s12935-014-0142-4
Krausz AE, Adler BL, Makdisi J, Schairer D, Rosen J, Landriscina A, Navati M, Alfieri A, Friedman JM, Nosanchuk JD, Rodriguez-Gabin A, Ye KQ, McDaid HM, Friedman AJ (2018) Nanoparticle-encapsulated doxorubicin demonstrates superior tumor cell kill in triple negative breast cancer subtypes intrinsically resistant to doxorubicin. Precis Nanomed 1:173–182. https://doi.org/10.33218/prnano1(3).181029.1
Zhao L, Qi Y, Xu L, Tao X, Han X, Yin L, Peng J (2018) MicroRNA-140-5p aggravates doxorubicin-induced cardiotoxicity by promoting myocardial oxidative stress via targeting Nrf2 and Sirt2. Redox Biol 15:284–296. https://doi.org/10.1016/j.redox.2017.12.013
Lovitt CJ, Shelper TB, Avery VM (2018) Doxorubicin resistance in breast cancer cells is mediated by extracellular matrix proteins. BMC Cancer 18:41. https://doi.org/10.1186/s12885-017-3953-6
Jabłońska-Trypuć A, Krętowski R, Kalinowska M, Świderski G, Cechowska-Pasko M, Lewandowski W (2018) Possible mechanisms of the prevention of doxorubicin toxicity by cichoric acid—antioxidant nutrient. Nutrients 10:44
Ascolani G, Lio P (2014) Modeling TGF-beta in early stages of cancer tissue dynamics. PLoS One 9:e88533. https://doi.org/10.1371/journal.pone.0088533
Worthington JJ, Klementowicz JE, Travis MA (2011) TGFbeta: a sleeping giant awoken by integrins. Trends Biochem Sci 36:47–54. https://doi.org/10.1016/j.tibs.2010.08.002
Gulei D, Mehterov N, Ling H, Stanta G, Braicu C, Berindan-Neagoe I (2017) The “good-cop bad-cop” TGF-beta role in breast cancer modulated by non-coding RNAs. Biochim Biophys Acta 1861:1661–1675. https://doi.org/10.1016/j.bbagen.2017.04.007
Ganapathy V, Ge R, Grazioli A, Xie W, Banach-Petrosky W, Kang Y, Lonning S, McPherson J, Yingling JM, Biswas S, Mundy GR, Reiss M (2010) Targeting the transforming growth factor-beta pathway inhibits human basal-like breast cancer metastasis. Mol Cancer 9:122. https://doi.org/10.1186/1476-4598-9-122
Wang XG, Meng Q, Qi FM, Yang QF (2014) Blocking TGF-β inhibits breast cancer cell invasiveness via ERK/S100A4 signal. Eur Rev Med Pharmacol Sci 18:3844–3853
Gulei D, Mehterov N, Ling H, Stanta G, Braicu C, Berindan-Neagoe I (2017) The “good-cop bad-cop” TGF-beta role in breast cancer modulated by non-coding RNAs. Biochim Biophys Acta Gen Subj 1861:1661–1675. https://doi.org/10.1016/j.bbagen.2017.04.007
Lin X, Li L, Wang R, Wilcox D, Zhao X, Song J, Huang X, Hansen TM, Dande P, Wada C, Hubbard RD, Kohlbrenner WM, Fesik SW, Shen Y (2011) A robust in vivo positive-readout system for monitoring siRNA delivery to xenograft tumors. RNA 17:603–612. https://doi.org/10.1261/rna.2546011
Irimie AI, Braicu C, Cojocneanu-Petric R, Berindan-Neagoe I, Campian RS (2015) Novel technologies for oral squamous carcinoma biomarkers in diagnostics and prognostics. Acta Odontol Scand 73:161–168. https://doi.org/10.3109/00016357.2014.986754
Irimie AI, Braicu C, Pileczki V, Petrushev B, Soritau O, Campian RS, Berindan-Neagoe I (2016) Knocking down of p53 triggers apoptosis and autophagy, concomitantly with inhibition of migration on SSC-4 oral squamous carcinoma cells. Mol Cell Biochem 419:75–82. https://doi.org/10.1007/s11010-016-2751-9
Pileczki V, Braicu C, Gherman CD, Berindan-Neagoe I (2012) TNF-alpha gene knockout in triple negative breast cancer cell line induces apoptosis. Int J Mol Sci 14:411–420. https://doi.org/10.3390/ijms14010411
Buduru S, Zimta AA, Ciocan C, Braicu C, Dudea D, Irimie AI, Berindan-Neagoe I (2018) RNA interference: new mechanistic and biochemical insights with application in oral cancer therapy. Int J Nanomed 13:3397–3409. https://doi.org/10.2147/ijn.S167383
Pileczki V, Pop L, Braicu C, Budisan L, Bolba Morar G, Del C Monroig-Bosque P, Sandulescu RV, Berindan-Neagoe I (2016) Double gene siRNA knockdown of mutant p53 and TNF induces apoptosis in triple-negative breast cancer cells. Onco Targets Ther 9:6921–6933. https://doi.org/10.2147/ott.S110719
Braicu C, Pileczki V, Irimie A, Berindan-Neagoe I (2013) p53siRNA therapy reduces cell proliferation, migration and induces apoptosis in triple negative breast cancer cells. Mol Cell Biochem 381:61–68. https://doi.org/10.1007/s11010-013-1688-5
Berindan-Neagoe I, Braicu C, Irimie A (2012) Combining the chemotherapeutic effects of epigallocatechin 3-gallate with siRNA-mediated p53 knock-down results in synergic pro-apoptotic effects. Int J Nanomed 7:6035–6047. https://doi.org/10.2147/ijn.S36523
Gorini S, De Angelis A, Berrino L, Malara N, Rosano G, Ferraro E (2018) Chemotherapeutic drugs and mitochondrial dysfunction: focus on doxorubicin, trastuzumab, and sunitinib. Oxidative Med Cell Longev 2018:7582730–7582730. https://doi.org/10.1155/2018/7582730
Ding MJ, Su KE, Cui GZ, Yang WH, Chen L, Yang M, Liu YQ, Dai DL (2016) Association between transforming growth factor-beta1 expression and the clinical features of triple negative breast cancer. Oncol Lett 11:4040–4044. https://doi.org/10.3892/ol.2016.4497
Menendez D, Shatz M, Resnick MA (2013) Interactions between the tumor suppressor p53 and immune responses. Curr Opin Oncol 25:85–92. https://doi.org/10.1097/CCO.0b013e32835b6386
Liu Z, Jiang Z, Gao Y, Wang L, Chen C, Wang X (2019) TP53 mutations promote immunogenic activity in breast cancer. J of Oncol 2019:5952836–5952836. https://doi.org/10.1155/2019/5952836
Braicu C, Buse M, Busuioc C, Drula R, Gulei D, Raduly L, Rusu A, Irimie A, Atanasov AG, Slaby O, Ionescu C, Berindan-Neagoe I (2019) A comprehensive review on MAPK: a promising therapeutic target in cancer. Cancers (Basel). https://doi.org/10.3390/cancers11101618
Palmieri D, Duchnowska R, Woditschka S, Hua E, Qian Y, Biernat W, Sosinska-Mielcarek K, Gril B, Stark AM, Hewitt SM, Liewehr DJ, Steinberg SM, Jassem J, Steeg PS (2014) Profound prevention of experimental brain metastases of breast cancer by temozolomide in an MGMT-dependent manner. Clin Cancer Res 20:2727–2739. https://doi.org/10.1158/1078-0432.ccr-13-2588
Neto JC, Ikoma MM, Carvalho KC, Vassallo J, De Brot M, Gobbi H, Soares FA, Rocha RM (2012) MGMT and PTEN as potential prognostic markers in breast cancer. Exp Mol Pathol 92:20–26. https://doi.org/10.1016/j.yexmp.2011.09.019
Delou JMdA, Vignal GM, Índio-do-Brasil V, Accioly MTdS, da Silva TSL, Piranda DN, Sobral-Leite M, de Carvalho MA, Capella MAM, Vianna-Jorge R (2017) Loss of constitutive ABCB1 expression in breast cancer associated with worse prognosis. Breast Cancer (Dove Medical Press) 9:415–428. https://doi.org/10.2147/BCTT.S131284
Hientz K, Mohr A, Bhakta-Guha D, Efferth T (2017) The role of p53 in cancer drug resistance and targeted chemotherapy. Oncotarget 8:8921–8946. https://doi.org/10.18632/oncotarget.13475
von Manstein V, Yang CM, Richter D, Delis N, Vafaizadeh V, Groner B (2013) Resistance of cancer cells to targeted therapies through the activation of compensating signaling loops. Curr Signal Transduct Ther 8:193–202. https://doi.org/10.2174/1574362409666140206221931
Hartman ZC, Poage GM, den Hollander P, Tsimelzon A, Hill J, Panupinthu N, Zhang Y, Mazumdar A, Hilsenbeck SG, Mills GB, Brown PH (2013) Growth of triple-negative breast cancer cells relies upon coordinate autocrine expression of the proinflammatory cytokines IL-6 and IL-8. Cancer Res 73:3470–3480. https://doi.org/10.1158/0008-5472.Can-12-4524-t
Poage GM, Hartman ZC, Brown PH (2013) Revealing targeted therapeutic opportunities in triple-negative breast cancers: a new strategy. Cell Cycle 12:2705–2706. https://doi.org/10.4161/cc.25871
Wang K, Zhu X, Zhang K, Yin Y, Chen Y, Zhang T (2018) Interleukin-6 contributes to chemoresistance in MDA-MB-231 cells via targeting HIF-1alpha. J Biochem Mol Toxicol 32:e22039. https://doi.org/10.1002/jbt.22039
Amara D, Wolf DM, Van’t Veer L, Esserman L, Campbell M, Yau C (2017) Co-expression modules identified from published immune signatures reveal five distinct immune subtypes in breast cancer. Breast Cancer Res Treat 161:41–50. https://doi.org/10.1007/s10549-016-4041-3
Tulsyan S, Mittal RD, Mittal B (2016) The effect of ABCB1 polymorphisms on the outcome of breast cancer treatment. Pharmacogenomics and personalized medicine 9:47–58. https://doi.org/10.2147/PGPM.S86672
Oba T, Izumi H, Ito K-I (2016) ABCB1 and ABCC11 confer resistance to eribulin in breast cancer cell lines. Oncotarget 7:70011–70027. https://doi.org/10.18632/oncotarget.11727
Etemadmoghadam D, Weir BA, Au-Yeung G, Alsop K, Mitchell G, George J, Australian Ovarian Cancer Study G, Davis S, D'Andrea AD, Simpson K, Hahn WC, Bowtell DDL (2013) Synthetic lethality between CCNE1 amplification and loss of BRCA1. Proc Natl Acad Sci U S A 110:19489–19494. https://doi.org/10.1073/pnas.1314302110
Wei C-Y, Tan Q-X, Zhu X, Qin Q-H, Zhu F-B, Mo Q-G, Yang W-P (2015) Expression of CDKN1A/p21 and TGFBR2 in breast cancer and their prognostic significance. Int J Clin Exp Pathol 8:14619–14629
Guiu S, Charon-Barra C, Vernerey D, Fumoleau P, Campone M, Spielmann M, Roché H, Mesleard C, Arnould L, Lemonnier J, Lacroix-Triki M (2015) Coexpression of androgen receptor and FOXA1 in nonmetastatic triple-negative breast cancer: ancillary study from PACS08 trial. Future Oncol 11:2283–2297. https://doi.org/10.2217/fon.15.102
Nadler Y, González AM, Camp RL, Rimm DL, Kluger HM, Kluger Y (2010) Growth factor receptor-bound protein-7 (Grb7) as a prognostic marker and therapeutic target in breast cancer. Ann Oncol 21:466–473. https://doi.org/10.1093/annonc/mdp346
de Silva HC, Lin MZ, Phillips L, Martin JL, Baxter RC (2019) IGFBP-3 interacts with NONO and SFPQ in PARP-dependent DNA damage repair in triple-negative breast cancer. Cell Mol Life Sci 76:2015–2030. https://doi.org/10.1007/s00018-019-03033-4
Malone MK, Smrekar K, Park S, Blakely B, Walter A, Nasta N, Park J, Considine M, Danilova LV, Pandey NB, Fertig EJ, Popel AS, Jin K (2020) Cytokines secreted by stromal cells in TNBC microenvironment as potential targets for cancer therapy. Cancer Biol Ther. https://doi.org/10.1080/15384047.2020.1739484
Jing X, Liang H, Hao C, Yang X, Cui X (2019) Overexpression of MUC1 predicts poor prognosis in patients with breast cancer. Oncol Rep 41:801–810. https://doi.org/10.3892/or.2018.6887
Hata T, Rajabi H, Takahashi H, Yasumizu Y, Li W, Jin C, Long MD, Hu Q, Liu S, Fushimi A, Yamashita N, Kui L, Hong D, Yamamoto M, Miyo M, Hiraki M, Maeda T, Suzuki Y, Samur MK, Kufe D (2019) MUC1-C activates the NuRD complex to drive dedifferentiation of triple-negative breast cancer cells. Cancer Res 79:5711–5722. https://doi.org/10.1158/0008-5472.Can-19-1034
Rohrberg J, Van de Mark D, Amouzgar M, Lee JV, Taileb M, Corella A, Kilinc S, Williams J, Jokisch ML, Camarda R, Balakrishnan S, Shankar R, Zhou A, Chang AN, Chen B, Rugo HS, Dumont S, Goga A (2020) MYC dysregulates mitosis, revealing cancer vulnerabilities. Cell Rep 30:3368–3382.e7. https://doi.org/10.1016/j.celrep.2020.02.041
Jiang J, Thyagarajan-Sahu A, Loganathan J, Eliaz I, Terry C, Sandusky GE, Sliva D (2012) BreastDefend™ prevents breast-to-lung cancer metastases in an orthotopic animal model of triple-negative human breast cancer. Oncol Rep 28:1139–1145. https://doi.org/10.3892/or.2012.1936
Jin T, Suk Kim H, Ki Choi S, Hye Hwang E, Woo J, Suk Ryu H, Kim K, Moon A, Kyung Moon W (2017) microRNA-200c/141 upregulates SerpinB2 to promote breast cancer cell metastasis and reduce patient survival. Oncotarget 8:32769–32782. https://doi.org/10.18632/oncotarget.15680
Schäfer SA, Hülsewig C, Barth P, von Wahlde MK, Tio J, Kolberg HC, Bernemann C, Blohmer JU, Kiesel L, Kolberg-Liedtke C (2019) Correlation between SFRP1 expression and clinicopathological parameters in patients with triple-negative breast cancer. Future Oncol 15:1921–1938. https://doi.org/10.2217/fon-2018-0564
Kavanagh E, Joseph B (2011) The hallmarks of CDKN1C (p57, KIP2) in cancer. Biochem Biophys Acta 1816:50–56. https://doi.org/10.1016/j.bbcan.2011.03.002
Domagala P, Hybiak J, Rys J, Byrski T, Cybulski C, Lubinski J (2016) Pathological complete response after cisplatin neoadjuvant therapy is associated with the downregulation of DNA repair genes in BRCA1-associated triple-negative breast cancers. Oncotarget 7:68662–68673. https://doi.org/10.18632/oncotarget.11900
Barroso-Sousa R, Keenan TE, Pernas S, Exman P, Jain E, Garrido-Castro AC, Hughes ME, Bychkovsky B, Umeton R, Files JL, Lindeman NI, MacConaill LE, Hodi FS, Krop I, Dillon DA, Winer EP, Wagle N, Lin NU, Mittendorf EA, Van Allen EM, Tolaney SM (2020) Tumor mutational burden and PTEN alterations as molecular correlates of response to PD-1/L1 blockade in metastatic triple-negative breast cancer. Clin Cancer Res. https://doi.org/10.1158/1078-0432.Ccr-19-3507
Keniry M, Parsons R (2008) The role of PTEN signaling perturbations in cancer and in targeted therapy. Oncogene 27:5477–5485. https://doi.org/10.1038/onc.2008.248
Funding
This paper was published under the frame of European Social Found, Human Capital Operational Programme 2014–2020, Project No. POCU/380/6/13/125171, and PNCDI III 2015–2020 “Increasing the performance of scientific research and technology transfer in translational medicine through the formation of a new generation of young researchers”—ECHITAS, no. 29PFE/ 18.10. 2018. Cristina Alexandra Ciocan-Cȃrtiţă received a grant for doctoral research Grant Number 1300/11/13.01.2017.
Author information
Authors and Affiliations
Contributions
Conceptualization was done by CACC, IBN, CB; contributions to methodology were done by AJ, CACC, LR, RC, AM, VP, LB; software was provided by RC, LAP, CB; validation was done by AJ, CB; formal analysis was carried out LAP, CB; investigation was done by IBN.; resources were gathered by IBN, CB; data curation was performed by RC; writing—original draft preparation—was done by AJ, CACC; writing—review and editing—was done by CB, IBN, SSK; visualization was carried out by CB, RC; supervision was done by IBN; editing was done by SSK; project administration was done by IBN; funding was acquired by IBN, CACC, CB.
Corresponding authors
Ethics declarations
Conflict of interests
The authors declare that they have no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ciocan-Cȃrtiţă, C.A., Jurj, A., Raduly, L. et al. New perspectives in triple-negative breast cancer therapy based on treatments with TGFβ1 siRNA and doxorubicin. Mol Cell Biochem 475, 285–299 (2020). https://doi.org/10.1007/s11010-020-03881-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-020-03881-w