Abstract
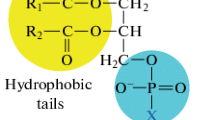
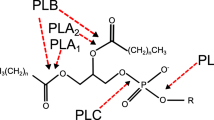
Phospholipase C (PLC) catalyzes the hydrolysis of phospholipids to produce phosphate monoesters and diacylglycerol. It has many applications in the enzymatic degumming of plant oils. PLC Bc , a bacterial PLC from Bacillus cereus, is an optimal choice for this activity in terms of its wide substrate spectrum, high activity, and approved safety. Unfortunately, its large-scale production and reliable high-throughput screening of PLC Bc remain challenging. Herein, we summarize the research progress regarding PLC Bc with emphasis on the screening methods, expression systems, catalytic mechanisms and inhibitor of PLC Bc . This review hopefully will inspire new achievements in related areas, to promote the sustainable development of PLC Bc and its application.




Similar content being viewed by others
References
Antikainen NM, Hergenrother PJ, Harris MM, Corbett W, Martin SF (2003a) Altering substrate specificity of phosphatidylcholine-preferring phospholipase C of Bacillus cereus by random mutagenesis of the headgroup binding site. Biochemistry 42:1603–1610
Antikainen NM, Monzingo AF, Franklin CL, Robertus JD, Martin SF (2003b) Using X-ray crystallography of the Asp55Asn mutant of the phosphatidylcholine-preferring phospholipase C from Bacillus cereus to support the mechanistic role of Asp55 as the general base. Arch Biochem Biophys 417:81–86
Barton NR (2008) A new process for degumming: the use of phospholipase C to improve yields during refining of high phosphorus vegetable oils. In: 99th AOCS annual meeting and expo, Seattle: abstracts, p 120
Benfield AP, Goodey NM, Phillips LT, Martin SF (2007) Structural studies examining the substrate specificity profiles of PC-PLC Bc protein variants. Arch Biochem Biophys 460:41–47
Bulnes PG, Roura AG, Canals D, Delgado A, Casas J, Llebaria A (2010) 2-Aminohydroxamic acid derivatives as inhibitors of Bacillus cereus phosphatidylcholine preferred phospholipase C PC-PLC Bc . Bioorg Med Chem 18:8549–8555
Cheong DE, Choi JH, Song JJ, Kim GJ (2013) Construction of non-invasively constitutive expression vectors using a metagenome-derived promoter for soluble expression of proteins. Bioprocess Biosyst Eng 36:667–676
Clausen K (2001) Enzymatic oil-degumming by a novel microbial phospholipase. Eur J Lipid Sci Technol 103:333–340
Dijkstra AJ (2010) Enzymatic degumming. Eur J Lipid Sci Technol 112:1178–1189
Durban M, Bornscheuer U (2003) An assay system for the detection of phospholipase C activity. Eur J Lipid Sci Technol 105:633–637
Durban MA, Bornscheuer UT (2007) An improved assay for the determination of phospholipase C activity. Eur J Lipid Sci Technol 109:469–473
Durban MA, Silbersack J, Schweder T, Schauer F, Bornscheuer UT (2007) High level expression of a recombinant phospholipase C from Bacillus cereus in Bacillus subtilis. Appl Microbiol Biotechnol 74:634–639
Flieger A, Gong SM, Faigle M, Neumeister B (2000) Critical evaluation of p-nitrophenylphosphorylcholine (p-NPPC) as artificial substrate for the detection of phospholipase C. Enzyme Microb Technol 26:451–458
Franklin CL, Li H, Martin SF (2003) Design, synthesis, and evaluation of water-soluble phospholipid analogues as inhibitors of phospholipase C from Bacillus cereus. J Org Chem 68:7298–7307
Graille J, Pina M, Montet D (1988) Biotechnologie des lipides: quelques applications possibles. Riv Ital Sostanze Gr 65:423–428
Gramatikova S, Hazlewood G, Lam D, Barton NR (2007) US Patent 7,226,771
Gramatikova S, Hazlewood G, Lam D, Barton NR (2008) US Patent Application Publication 2008/0317731
Grit M, Underberg WJ, Crommelin DJ (1993) Hydrolysis of saturated soybean phosphatidylcholine in aqueous liposome dispersions. J Pharm Sci 82:362–366
Hansen S, Kristian L, Hough H, Hough E (1992) Crystal-structures of phosphate, iodide and iodate-inhibited phospholipase C from Bacillus cereus and structural investigations of the binding of reaction products and a substrate-analog. J Mol Biol 225:543–549
Hansen S, Hough E, Svensson LA, Wong YL, Martin SF (1993) Crystal structure of phospholipase C from Bacillus cereus complexed with a substrate analog. J Mol Biol 234:179–187
Hergenrother PJ, Martin SF (2001) Phosphatidylcholine-preferring phospholipase C from B. cereus. Function, structure, and mechanism. Bioorg Chem Biol Signal Transduct 211:131–167
Hergenrother PJ, Spaller MR, Haas MK, Martin SF (1995) Chromogenic assay for phospholipase C from Bacillus cereus. Anal Biochem 229:313–316
Hough E et al (1989) High resolution (1.5 Å) crystal structure of phospholipase C from Bacillus cereus. Nature 338:357–360
Huang S, Liang M, Xu Y, Aamir R, Chun L (2014) Characteristics and vegetable oils degumming of recombinant phospholipase B. Chem Eng J 237:23–28
Ivanova N et al (2003) Genome sequence of Bacillus cereus and comparative analysis with Bacillus anthracis. Nature 423:87–91
Johansen T, Holm T, Guddal PH, Sletten K, Haugli FB, Little C (1988) Cloning and sequencing of the gene encoding the phosphatidylcholine preferring phospholipase C of Bacillus cereus. Gene 65:293–304
Kozawa O, Suzuki A, Kaida T, Tokuda H, Uematsu T (1997) Tumor necrosis factor-alpha autoregulates interleukin-6 synthesis via activation of protein kinase C—function of sphingosine 1-phosphate and phosphatidylcholine-specific phospholipase C. J Biol Chem 272:25099–25104
Kuppe A, Evans LM, Mcmillen DA, Griffith OH (1989) Phosphatidylinositol specific phospholipase C of Bacillus cereus—cloning, sequencing, and relationship to other phospholipases. J Bacteriol 171:6077–6083
Lee J, Kim Y, Min D (2010) Laser desorption/ionization mass spectrometric assay for phospholipase activity based on graphene oxide/carbon nanotube double-layer films. J Am Chem Soc Commun 132:14714–14717
Li YH, Maher P, Schubert D (1998) Phosphatidylcholine-specific phospholipase C regulates glutamate-induced nerve cell death. Proc Natl Acad Sci USA 95:7748–7753
Liao RZ, Yu JG, Himo F (2010) Reaction mechanism of the trinuclear zinc enzyme phospholipase C: a density functional theory study. J Phys Chem B 114:2533–2540
Liu Y, Ogawa K, Schanze KS (2008) Conjugated polyelectrolyte based real-time fluorescence assay for phospholipase C. Anal Chem 80:150–158
Liu M, Gu J, Xie W, Yu H (2013) Directed co-evolution of an endoglucanase and a β-glucosidase in Escherichia coli by a novel high-throughput screening method. Chem Commun 49:7219–7221
Liu M, Xie W, Xu H, Gu J, Lv X, Yu H, Ye L (2014) Directed evolution of an exoglucanase facilitated by a co-expressed β-glucosidase and construction of a whole engineered cellulase system in Escherichia coli. Biotechnol Lett 36:1801–1807
Luberto C, Hannun YA (1998) Sphingomyelin synthase, a potential regulator of intracellular levels of ceramide and diacylglycerol during SV40 transformation—does sphingomyelin synthase account for the putative, phosphatidylcholine-specific phospholipase C? J Biol Chem 273:14550–14559
Martin SF, Spaller MR, Hergenrother PJ (1996) Expression and site-directed mutagenesis of the phosphatidylcholine-preferring phospholipase C of Bacillus cereus: probing the role of the active site Glu146. Biochemistry 35:12970–12977
Martin SF, Follows BC, Hergenrother PJ, Franklin CL (2000a) A novel class of zinc-binding inhibitors for the phosphatidylcholine-preferring phospholipase C from Bacillus cereus. J Org Chem 65:4509–4514
Martin SF, Follows BC, Hergenrother PJ, Trotter BK (2000b) The choline binding site of phospholipase C (Bacillus cereus): insights into substrate specificity. Biochemistry 39:3410–3415
McGaughey CA, Chu HP (1948) The egg-yolk reaction of aerobic sporing Bacilli. J Gen Microbiol 2:334–340
Roberts MF, Wu YQ, Zhou C, Geng D, Tan C (1996) Mechanism and structure based inhibitors of phospholipase C enzymes. Adv Enzyme Regul 36:57–71
Roura AG, Casas J, Llebaria A (2002) Synthesis and phospholipase C inhibitory activity of D609 diastereomers. Lipids 37:401–406
Roura AG, Navarro I, Delgado A, Llebaria A, Casas J (2004) Disclosing new inhibitors by finding similarities in three-dimensional active-site architectures of polynuclear zinc phospholipases and aminopeptidases. Angew Chem Int Ed 43:862–865
Seo K, Rhee J (2004) High-level expression of recombinant phospholipase C form Bacillus cereus in Pichia pastoris and its characterization. Biotechnol Lett 26:1475–1479
Sø JB, Turner M (2007). US Patent Application Publication 2007/0298157
Sundell S, Hansen S, Hough E (1994) A proposal for the catalytic mechanism in phospholipase-C based on interaction energy and distance geometry calculations. Protein Eng 7:571–577
Tan CA, Hehir MJ, Roberts MF (1997) Cloning, overexpression, refolding, and purification of the nonspecific phospholipase C from Bacillus cereus. Protein Expr Purif 10:365–372
Thrige DD, Buur JRB, Jorgensen FS (1997) Substrate binding and catalytic mechanism in phospholipase C from Bacillus cereus: a molecular mechanics and molecular dynamics study. Biopolymers 42:319–336
Wargovich MJ et al (2000) Efficacy of potential chemopreventive agents on rat colon aberrant crypt formation and progression. Carcinogenesis 21:1149–1155
Zhang L et al (2010) D609 inhibits progression of preexisting atheroma and promotes lesion stability in apolipoprotein E−/− mice a role of phosphatidylcholine-specific phospholipase in atherosclerosis. Arterioscler Thromb Vasc Biol 30:U411–U490
Zufarov O, Schmidt Š, Sekretár S (2008) Degumming of rapeseed and sunflower oils. ACS 1:321–328
Acknowledgments
This work was supported by a Grant from Wilmar (Shanghai) Biotechnology Research and Development Center Co., Ltd.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lyu, Y., Ye, L., Xu, J. et al. Recent research progress with phospholipase C from Bacillus cereus . Biotechnol Lett 38, 23–31 (2016). https://doi.org/10.1007/s10529-015-1962-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10529-015-1962-6