Abstract
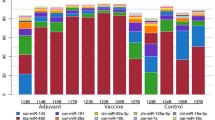
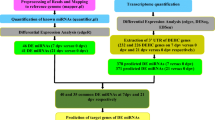
Previous studies have demonstrated the key regulatory roles played by microRNAs (miRNAs) in influenza virus-host interactions. To gain more insight into the contribution of miRNAs to the host immune response, spleen tissues from mice infected with A/Swine/GD/2/12 (H1N1) virus were harvested 5 days postinfection, and miRNA deep sequencing was performed. The results showed that 50 miRNAs were modulated. Interestingly, pathway analysis of miRNAs and targets showed that upregulated miR-124-3p interacts with innate immune-related pathways such as the Toll-like receptor pathway, RIG-I-like receptor signaling pathway, NOD-like receptor signaling pathway and JAK-STAT signaling pathway, and this might play a major role in the anti-inflammatory response. Further understanding of the roles played by these miRNAs in influenza virus infection will provide new insights into host-pathogen interactions.


Similar content being viewed by others
References
Li Y, Chan EY, Li J, Ni C, Peng X, Rosenzweig E, Tumpey TM, Katze MG (2010) MiRNA expression and virulence in pandemic influenza virus-infected mice. J Virol 84:3023–3032
Smith GJ, Vijaykrishna D, Bahl J, Lycett SJ, Worobey M, Pybus OG, Ma SK, Cheung CL, Raghwani J, Bhatt S, Peiris JS, Guan Y, Rambaut A (2009) Origins and evolutionary genomics of the 2009 swine-origin H1N1 influenza A epidemic. Nature 459:1122–1125
Li Y, Li J, Belisle S, Baskin CR, Tumpey TM, Katze MG (2011) Differential miRNA expression and virulence of avian, 1918 reassortant, and reconstructed 1918 influenza A viruses. Virology 421:105–113
Zhao Y, Srivastava D (2007) A developmental view of miRNA function. Trends Biochem Sci 32:189–197
Bartel DP (2004) MiRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
He L, Hannon GJ (2004) MiRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet 5:522–531
Valencia-Sanchez MA, Liu J, Hannon GJ, Parker R (2006) Control of translation and mRNA degradation by miRNAs and siRNAs. Genes Dev 20:515–524
Fontana L, Pelosi E, Greco P, Racanicchi S, Testa U, Liuzzi F, Croce CM, Brunetti E, Grignani F, Peschle C (2007) MiRNAs 17-5p-20a-106a control monocytopoiesis through AML1 targeting and M-CSF receptor upregulation. Nat Cell Biol 9:775–787
Felli N, Fontana L, Pelosi E, Botta R, Bonci D, Facchiano F, Liuzzi F, Lulli V, Morsilli O, Santoro S, Valtieri M, Calin GA, Liu CG, Sorrentino A, Croce CM, Peschle C (2005) MiRNAs 221 and 222 inhibit normal erythropoiesis and erythroleukemic cell growth via kit receptor down-modulation. Proc Natl Acad Sci USA 102:18081–18086
He L, He X, Lowe SW, Hannon GJ (2007) miRNAs join the p53 network–another piece in the tumour-suppression puzzle. Nat Rev Cancer 7:819–822
Song L, Liu H, Gao S, Jiang W, Huang W (2010) Cellular miRNAs inhibit replication of the H1N1 influenza A virus in infected cells. J Virol 84:8849–8860
Loveday EK, Svinti V, Diederich S, Pasick J, Jean F (2012) Temporal- and strain-specific host miRNA molecular signatures associated with swine-origin H1N1 and avian-origin H7N7 influenza A virus infection. J Virol 86:6109–6122
Rogers JV, Price JA, Wendling MQ, Long JP, Bresler HS (2012) Preliminary miRNA analysis in lung tissue to identify potential therapeutic targets against H5N1 infection. Viral Immunol 25:3–11
Skovgaard K, Cirera S, Vasby D, Podolska A, Breum SØ, Dürrwald R, Schlegel M, Heegaard PM (2013) Expression of innate immune genes, proteins and miRNAs in lung tissue of pigs infected experimentally with influenza virus (H1N2). Innate Immun 19:531–544
Wang Y, Brahmakshatriya V, Lupiani B, Reddy SM, Soibam B, Benham AL, Gunaratne P, Liu HC, Trakooljul N, Ing N, Okimoto R, Zhou H (2012) Integrated analysis of miRNA expression and mRNA transcriptome in lungs of avian influenza virus infected broilers. BMC Genomics 13:278
Wang Y, Brahmakshatriya V, Zhu H, Lupiani B, Reddy SM, Yoon BJ, Gunaratne PH, Kim JH, Chen R, Wang J, Zhou H (2009) Identification of differentially expressed miRNAs in chicken lung and trachea with avian influenza virus infection by a deep sequencing approach. BMC Genomics 10:512
Terrier O, Textoris J, Carron C, Marcel V, Bourdon JC, Rosa-Calatrava M (2013) Host miRNA molecular signatures associated with human H1N1 and H3N2 influenza A viruses reveal an unanticipated antiviral activity for miR-146a. J Gen Virol 94:985–995
Barnard DL (2009) Animal models for the study of influenza pathogenesis and therapy. Antiviral Res 82:A110–A122. doi:10.1016/j.antiviral02.190
Wang L, Liu H, Li D, Chen H (2011) Identification and characterization of maize microRNAs involved in the very early stage of seed germination. BMC Genom 12:154
Kawai T, Akira S (2010) The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat Immunol 11:373–384
Kumar H, Kawai T, Akira S (2009) Pathogen recognition in the innate immune response. Biochem J 420:1–16
Kobayashi T, Walsh MC, Choi Y (2004) The role of TRAF6 in signal transduction and the immune response. Microbes Infect 6:1333–1338
Sun Y, Li Q, Gui H, Xu DP, Yang YL, Su DF, Liu X (2013) MiRNA-124 mediates the cholinergic anti-inflammatory action through inhibiting the production of pro-inflammatory cytokines. Cell Res 23:1270–1283
de Jonge WJ, van der Zanden EP, The FO, Bijlsma MF, van Westerloo DJ, Bennink RJ, Berthoud HR, Uematsu S, Akira S, van den Wijngaard RM, Boeckxstaens GE (2005) Stimulation of the vagus nerve attenuates macrophage activation by activating the Jak2-STAT3 signaling pathway. Nat Immunol 6:844–851
Borovikova LV, Ivanova S, Zhang M, Yang H, Botchkina GI, Watkins LR, Wang H, Abumrad N, Eaton JW, Tracey KJ (2000) Vagus nerve stimulation attenuates the systemic inflammatory response to endotoxin. Nature 405:458–462
Wang H, Yu M, Ochani M, Amella CA, Tanovic M, Susarla S, Li JH, Wang H, Yang H, Ulloa L, Al-Abed Y, Czura CJ, Tracey KJ (2003) Nicotinic acetylcholine receptor alpha7 subunit is an essential regulator of inflammation. Nature 4(21):384–388
Takeuchi K, Ito F (2004) Suppression of adriamycin-induced apoptosis by sustained activation of the phosphatidylinositol-3′-OH kinase-Akt pathway. J Biol Chem 279:892–900
Yu HG, Ai YW, Yu LL, Zhou XD, Liu J, Li JH, Xu XM, Liu S, Chen J, Liu F, Qi YL, Deng Q, Cao J, Liu SQ, Luo HS, Yu JP (2008) Phosphoinositide 3-kinase/Akt pathway plays an important role in chemoresistance of gastric cancer cells against etoposide and doxorubicin induced cell death. Int J Cancer 122:433–443
de Souza Rocha Simonini P, Breiling A, Gupta N, Malekpour M, Youns M, Omranipour R, Malekpour F, Volinia S, Croce CM, Najmabadi H, Diederichs S, Sahin O, Mayer D, Lyko F, Hoheisel JD, Riazalhosseini Y (2010) Epigenetically deregulated miRNA-375 is involved in a positive feedback loop with estrogen receptor alpha in breast cancer cells. Cancer Res 70:9175–9184
Ding L, Xu Y, Zhang W, Deng Y, Si M, Du Y, Yao H, Liu X, Ke Y, Si J, Zhou T (2010) MiR-375 frequently downregulated in gastric cancer inhibits cell proliferation by targeting JAK2. Cell Res 20:784–793
Tsukamoto Y, Nakada C, Noguchi T, Tanigawa M, Nguyen LT, Uchida T, Hijiya N, Matsuura K, Fujioka T, Seto M, Moriyama M (2010) MiRNA-375 is downregulated in gastric carcinomas and regulates cell survival by targeting PDK1 and 14-3-3zeta. Cancer Res 70:2339–2349
Ueda T, Volinia S, Okumura H, Shimizu M, Taccioli C, Rossi S, Alder H, Liu CG, Oue N, Yasui W, Yoshida K, Sasaki H, Nomura S, Seto Y, Kaminishi M, Calin GA, Croce CM (2010) Relation between miRNA expression and progression and prognosis of gastric cancer: a miRNA expression analysis. Lancet Oncol 11:136–146
Acknowledgments
This study was supported by the College of Veterinary Medicine of South China Agricultural University and University-Based Key Laboratory of Preventive Veterinary Medicine of Foshan University in Guangdong Province.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Additional information
L. Huang and J. Ma contributed equally.
Rights and permissions
About this article
Cite this article
Huang, L., Ma, J., Sun, Y. et al. Altered splenic miRNA expression profile in H1N1 swine influenza. Arch Virol 160, 979–985 (2015). https://doi.org/10.1007/s00705-015-2351-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-015-2351-0