Abstract
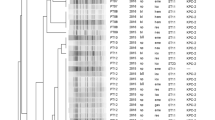
The objectives of this study were to determine the distribution of phylogenetic groups among Klebsiella pneumoniae isolates from Recife, Brazil and to assess the relationship between the groups and the isolation sites and resistance profile. Ninety four isolates of K. pneumoniae from hospital or community infections and from normal microbiota were analyzed by gyrA PCR–RFLP, antibiotic susceptibility, and adonitol fermentation. The results revealed the distinction of three phylogenetic groups, as it has also been reported in Europe, showing that these clusters are highly conserved within K. pneumoniae. Group KpI was dominantly represented by hospital and community isolates while groups KpII and KpIII displayed mainly normal microbiota isolates. The resistance to third generation cephalosporins, aztreonam, imipenem, amoxicillin/clavulanic acid, and streptomycin was only observed in KpI. The percentage of resistance was higher in KpI, followed by KpII and KpIII. The differences in the distribution of K. pneumoniae phylogenetic groups observed in this study suggest distinctive clinical and epidemiological characteristics among the three groups, which is important to understand the epidemiology of infections caused by this organism. This is the first study in Brazil on K. pneumoniae isolates from normal microbiota and community infections regarding the distribution of phylogenetic groups based on the gyrA gene.

Similar content being viewed by others
References
Alves MS, Dias RCS, Castro ACD et al (2006) Identification of clinical isolates of indole-positive and indole-negative Klebsiella spp. J Clin Microbiol 44:3640–3646
Brisse S, Verhoef J (2001) Phylogenetic diversity of Klebsiella pneumoniae and Klebsiella oxytoca clinical isolates revealed by randomly amplified polymorphic DNA, gyrA and parC genes sequencing and automated ribotyping. Int J Syst Evol Microbiol 51:915–924
Brisse S, Van Himbergen T, Kusters K et al (2004) Development of a rapid identification method for Klebsiella pneumoniae phylogenetic groups and analysis of 420 clinical isolates. Clin Microbiol Infect 10:942–945
Castanheira M, Mendes RE, Rhomberg PR et al (2008) Rapid emergence of bla CTX-M among Enterobacteriaceae in U.S. Medical Centers: molecular evaluation from the MYSTIC Program 2007. Microb Drug Resist 14:211–216
Clinical and Laboratory Standards Institute (2006) Performance standards for antimicrobial susceptibility testing. 16th informational supplement. CLSI M100-S17, Wayne, PA, 177 pp
Farmer JJ, Davis BR, Hickman-Brenner FW et al (1985) Biochemical identification of new species and biotypes of Enterobacteriaceae isolated from clinical specimens. J Clin Microbiol 21:46–76
García San Miguel L, Cobo J, Valverde A et al (2007) Clinical variables associated with the isolation of Klebsiella pneumoniae expressing different extended-spectrum beta-lactamases. Clin Microbiol Infect 13:532–538
Haeggman S, Löfdahl S, Paauw A et al (2004) Diversity and evolution of the class a chromosomal beta-lactamase gene in Klebsiella pneumoniae. Antimicrob Agents Chemother 48:2400–2408
Koh TH, Sng LH, Babini GS et al (2001) Carbapenem-resistant Klebsiella pneumoniae in Singapore producing IMP-1 beta-lactamase and lacking an outer membrane protein. Antimicrob Agents Chemother 45:1939–1940
Livermore DM (1995) β-Lactamases in laboratory and clinical resistance. Clin Microbiol Rev 8:557–584
Lopes ACS, Rodrigues JF, Clementino MBM et al (2007) Application of PCR ribotyping and tDNA-PCR for Klebsiella pneumoniae identification. Mem Inst Oswaldo Cruz 102:827–832
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Laboratory Press, New York, pp 368–369
Minarini LA, Gales AC, Palazzo IC et al (2007) Prevalence of community-occurring extended spectrum beta-lactamase-producing Enterobacteriaceae in Brazil. Curr Microbiol 54:335–341
Moland ES, Hanson ND, Herrera VL et al (2003) Plasmid-mediated, carbapenem-hydrolysing beta-lactamase, KPC-2, in Klebsiella pneumoniae isolates. J Antimicrob Chemother 51:711–714
Monteiro J, Santos AF, Asensi MD et al (2009) First report of KPC-2-producing Klebsiella pneumoniae strains in Brazil. Antimicrob Agents Chemother 53:333–334
Paterson DL, Hujer KM, Hujer AM et al (2003) Extended-spectrum beta-lactamases in Klebsiella pneumoniae bloodstream isolates from seven countries: dominance and widespread prevalence of SHV- and CTX-M-type beta-lactamases. Antimicrob Agents Chemother 47:3554–3560
Peirano G, Seki LM, Val Passos VL et al (2009) Carbapenem-hydrolysing beta-lactamase KPC-2 in Klebsiella pneumoniae isolated in Rio de Janeiro, Brazil. J Antimicrob Chemother 63:265–268
Rosenblueth M, Martínez L, Silva J et al (2004) Klebsiella variicola, a novel species with clinical and plant associated isolates. Syst Appl Microbiol 27:27–35
Selden R, Lee S, Wang WL et al (1971) Nosocomial Klebsiella infections: intestinal colonization as a reservoir. Ann Intern Med 74:657–664
Souza Lopes AC, Rodrigues JF, Morais Júnior MA (2005) Molecular typing of Klebsiella pneumoniae isolates from public hospitals in Recife, Brazil. Microbiol Res 160:37–46
Vercauteren E, Descheemaeker P, Ieven M et al (1997) Comparison of screening methods for detection of extended-spectrum β-lactamases and their prevalence among blood isolates of Escherichia coli and Klebsiella spp. in a Belgian teaching hospital. J Clin Microbiol 35:2191–2197
Woodford N, Tierno PM Jr, Young K et al (2004) Outbreak of Klebsiella pneumoniae producing a new carbapenem-hydrolyzing class A beta-lactamase, KPC-3, in a New York Medical Center. Antimicrob Agents Chemother 48:4793–4799
Acknowledgments
This work was financially supported by Fundação de Amparo à Ciência e Tecnologia do Estado de Pernambuco (FACEPE).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
de Melo, M.E.S., Cabral, A.B., Maciel, M.A.V. et al. Phylogenetic Groups Among Klebsiella pneumoniae Isolates from Brazil: Relationship with Antimicrobial Resistance and Origin. Curr Microbiol 62, 1596–1601 (2011). https://doi.org/10.1007/s00284-011-9903-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-011-9903-7