Abstract
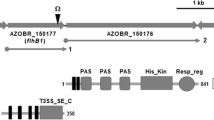
We report, here, the characterization of a mutant strain of Azospirillum brasilense Sp7 impaired in surface motility and chemotactic response. Presence of flagella in the mutant strain was confirmed by western blot analysis, using antisera raised against the polar and lateral flagellins, and by electron microscopy. Genetic complementation and nucleotide sequencing led to the identification of a new gene, named chsA. The deduced translation product, ChsA protein, contained a PAS sensory domain and an EAL domain. As ChsA displayed characteristic signaling protein architecture, it is thought that this protein is a component of the signaling pathway controlling chemotaxis in Azospirillum.




Similar content being viewed by others
References
Alexandre G, Greer SE, Zhulin IB (2000) Energy taxis is the dominant behavior in Azospirillum brasilense. J Bacteriol 182:6042–6048
Ausubel FM, Brent R, Kingston RE, Moore DD, Seidman JG, Smith JA, Struhl K (1999) Short protocols in molecular biology. Wiley, New York
Barak R, Nur I, Okon Y (1983) Detection of chemotaxis in Azospirillum brasilense. J Appl Bacteriol 53:399–403
Bible AN, Stephens BB, Ortega DR, Xie Z, Alexandre G (2008) Function of a chemotaxis-like signal transduction pathway in modulating motility, cell clumping and cell length in the alpha-proteobacterium Azospirillum brasilense. J Bacteriol 190:6365–6367
Carreño-López R, Campos Reales N, Elmerich C, Baca BE (2000) Physiological evidence for differently regulated tryptophan-dependent pathways for indole-3-acetic acid synthesis in Azospirillum brasilense Sp7. Mol Gen Genet 264:521–530
Carreño-López R, Pereg-Gerk L, Baca BE, Elmerich C (2002) Characterization of genes that play a role in swarming motility of Azospirillum brasilense. In: Finan TM, O’Brian MR, Layzell DB, Vessey JK, Newton WE (eds) Nitrogen fixation global perspectives. CABI Publishing, London, p 474
Carreño-López R, Sanchez A, Camargo N, Elmerich C, Baca BE (2008) Characterization of chsA, a new gene controlling the chemotactic response in Azospirillum brasilense. In: Dakora FD, Chimphango SBM, Valentine AJ, Elmerich C, Newton WE (eds) Biological nitrogen fixation: towards poverty alleviation through sustainable agriculture. Springer Science+Business Media BV, Dordrecht, pp 317–318
Croes L, Moens S, van Bastelaere E, Vanderleyden J, Michiels KW (1993) The polar flagellum mediates Azospirillum brasilense adsorption to wheat roots. J Gen Microbiol 139:2261–2269
Dobbelaere S, Croonenborghs A, Thys A, Ptacek D, Vanderleyden J, Dutto P, Labandera-Gonzalez C, Caballero-Mellado J, Aguirre JF, Kapulnik Y, Brener S, Burdman S, Kadouri D, Sarig S, Okon Y (2001) Responses of agronomically important crops to inoculation with Azospirillum. Aust J Plant Physiol 28:871–879
Elmerich C, de Zamaroczy M, Arsène F, Pereg L, Paquelin A, Kaminski A (1997) Regulation of nif gene expression and nitrogen metabolism in Azospirillum. Soil Biol Biochem 29:847–852
Galperin MY (2004) Bacterial signal transduction network in a genomic perspective. Environ Microbiol 6:552–567
Greer-Phillips SE, Alexandre G, Taylor BL, Zhulin IB (2003) Aer and Tsr guide Escherichia coli in spatial gradients of oxidizable substrates. Microbiology 149:2661–2667
Greer-Phillips SG, Stephens BB, Alexandre G (2004) An energy taxis transducer promotes root colonization by Azospirillum brasilense. J Bacteriol 186:6595–6604
Hall PG, Krieg NR (1983) Swarming of Azospirillum brasilense on solid media. Can J Microbiol 29:1592–1594
Hauwaerts D, Alexandre G, Das SK, Vanderleyden J, Zhulin IB (2002) A major chemotaxis gene cluster in Azospirillum brasilense and relationships between chemotaxis operons in α-proteobacteria. FEMS Microbiol Lett 208:61–67
Jenal U, Malone J (2006) Mechanisms of cyclic-di-GMP signaling in bacteria. Annu Rev Genet 40:385–407
Katupitiya S, Millet J, Vesk M, Viccars L, Zeman A, Lidong Z, Elmerich C, Kennedy IR (1995) A mutant of Azospirillum brasilense Sp7 impaired in flocculation with a modified colonization pattern and superior nitrogen fixation in association with wheat. Appl Environ Microbiol 61:1987–1995
Knauf VC, Nester EW (1982) Wide host range cloning vectors: a cosmid clone bank of an Agrobacterium Ti plasmid. Plasmid 8:45–54
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Milcamps A, van Dommelen A, Stigler J, Vanderleyden J, de Bruijn FJ (1996) The Azospirillum brasilense rpoN gene is involved in nitrogen fixation nitrate assimilation ammonium uptake and flagellar biosynthesis. Can J Microbiol 42:3205–3218
Moens S, Michiels K, Keijers V, van Leuven F, Vanderleyden J (1995) Cloning sequencing and phenotype analysis of laf1 encoding the flagellin of the lateral flagella of Azospirillum brasilense Sp7. J Bacteriol 177:5419–5426
Pereg-Gerk L, Paquelin A, Gounon P, Kennedy IR, Elmerich C (1998) A Transcriptional regulator of the LuxR-UhpA family FlcA controls flocculation and wheat root surface colonization by Azospirillum brasilense Sp7. Mol Plant Microbe Interact 11:177–187
Pereg-Gerk L, Gilchrist K, Kennedy IR (2000) Mutants with enhanced nitrogenase activity in hydroponic Azospirillum brasilense-wheat associations. Appl Environ Microbiol 66:2175–2184
Potrich DP, Bressel TA, Schrank IS, Passaglia LMP (2001) Sequencing and promoter analysis of the nifENXorf3orf5fdxAnifQ operon from Azospirillum brasilense Sp7. Brazilian J Med Biol Res 34:1379–1395
Römling U, Gomelsky M, Galperin MY (2005) C-di-GMPc: the dawning of a novel signaling system. Mol Microbiol 57:629–639
Schmidt AJ, Ryenkov DA, Gomelsky M (2005) The ubiquitous protein domain EAL is a cyclic diguanylate-specific phosphodiesterase: enzymatically active and inactive EAL domains. J Bacteriol 187:4774–4781
Simm R, Morr M, Kader A, Nimtz M, Römling U (2004) GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol Microbiol 53:1123–1134
Simon R, Priefer U, Pühler A (1983) A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in Gram negative bacteria. Bio/Technology 1:784–791
Stephens BB, Loar SN, Alexandre G (2006) Role of CheB and CheR in the complex chemotactic and aerotactic pathway of Azospirillum brasilense. J Bacteriol 188:4759–4768
Tamayo R, Tischler AD, Camilli A (2005) The EAL domain protein VieA is a cyclic diguanylate cyclase phosphodiesterase. J Biol Chem 280:33324–33330
Taylor B, Zhulin I (1999) PAS domains: internal sensors of oxygen redox potential and light. Microbiol Mol Biol Rev 63:479–506
Vande Broek A, Lambrecht M, Vanderleyden J (1998) Bacterial chemotactic motility is important for the initiation of wheat root colonization by Azospirillum brasilense. Microbiology 144:2599–2606
Acknowledgments
The authors wish to thank Ms N. Desnoues and Ma. L. Xiqui for skillful technical assistance and Ms Jerri Bram for improving the language. This work was partially supported by a grant of VIEP-SEP.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Ursula Priefer.
Rights and permissions
About this article
Cite this article
Carreño-López, R., Sánchez, A., Camargo, N. et al. Characterization of chsA, a new gene controlling the chemotactic response in Azospirillum brasilense Sp7. Arch Microbiol 191, 501–507 (2009). https://doi.org/10.1007/s00203-009-0475-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-009-0475-x