Abstract
Key message
A total of 33 additive stem rot QTLs were identified in peanut genome with nine of them consistently detected in multiple years or locations. And 12 pairs of epistatic QTLs were firstly reported for peanut stem rot disease.
Abstract
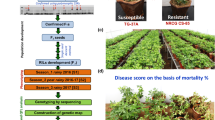
Stem rot in peanut (Arachis hypogaea) is caused by the Sclerotium rolfsii and can result in great economic loss during production. In this study, a recombinant inbred line population from the cross between NC 3033 (stem rot resistant) and Tifrunner (stem rot susceptible) that consists of 156 lines was genotyped by using 58 K peanut single nucleotide polymorphism (SNP) array and phenotyped for stem rot resistance at multiple locations and in multiple years. A linkage map consisting of 1451 SNPs and 73 simple sequence repeat (SSR) markers was constructed. A total of 33 additive quantitative trait loci (QTLs) for stem rot resistance were detected, and six of them with phenotypic variance explained of over 10% (qSR.A01-2, qSR.A01-5, qSR.A05/B05-1, qSR.A05/B05-2, qSR.A07/B07-1 and qSR.B05-1) can be consistently detected in multiple years or locations. Besides, 12 pairs of QTLs with epistatic (additive × additive) interaction were identified. An additive QTL qSR.A01-2 also with an epistatic effect interacted with a novel locus qSR.B07_1-1 to affect the percentage of asymptomatic plants in a row. A total of 193 candidate genes within 38 stem rot QTLs intervals were annotated with functions of biotic stress resistance such as chitinase, ethylene-responsive transcription factors and pathogenesis-related proteins. The identified stem rot resistance QTLs, candidate genes, along with the associated SNP markers in this study, will benefit peanut molecular breeding programs for improving stem rot resistance.


Similar content being viewed by others
References
Adrian M, Jeandet P (2012) Effects of resveratrol on the ultrastructure of Botrytis cinerea conidia and biological significance in plant/pathogen interactions. Fitoterapia 83:1345–1350
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Anco D (2017) Peanut disease management. South carolina pest management handbook for field crops. South Carolina State Library, Columbia, pp 190–201
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 57:289–300
Bera SK, Kamdar JH, Kasundra SV, Ajay BC (2016) A novel QTL governing resistance to stem rot disease caused by Sclerotium rolfsii in peanut. Aust Plant Pathol 45:637–644
Bertioli DJ, Cannon SB, Froenicke L, Huang G, Farmer AD, Cannon EK, Liu X, Gao D, Clevenger J, Dash S (2016) The genome sequences of Arachis duranensis and Arachis ipaensis, the diploid ancestors of cultivated peanut. Nat Genet 47:438
Bertioli DJ, Jenkins J, Clevenger J, Dudchenko O, Gao D, Seijo G, Leal-Bertioli SCM, Ren L, Farmer AD, Pandey MK, Samoluk SS, Abernathy B, Agarwal G, Ballén-Taborda C, Cameron C, Campbell J, Chavarro C, Chitikineni A, Chu Y, Dash S, El Baidouri M, Guo B, Huang W, Kim KD, Korani W, Lanciano S, Lui CG, Mirouze M, Moretzsohn MC, Pham M, Shin JH, Shirasawa K, Sinharoy S, Sreedasyam A, Weeks NT, Zhang X, Zheng Z, Sun Z, Froenicke L, Aiden EL, Michelmore R, Varshney RK, Holbrook CC, Cannon EKS, Scheffler BE, Grimwood J, Ozias-Akins P, Cannon SB, Jackson SA, Schmutz J (2019) The genome sequence of segmental allotetraploid peanut Arachis hypogaea. Nat Genet 51:877–884
Beute M, Wynne J, Emery DJCS (1976) Registration of NC 3033 Peanut Germplasm1 (Reg. No. GP 9) 16:887-887
Bocianowski J (2013) Epistasis interaction of QTL effects as a genetic parameter influencing estimation of the genetic additive effect. Genet Mol Biol 36:93–100
Chen W, Jiao Y, Cheng L, Huang L, Liao B, Tang M, Ren X, Zhou X, Chen Y, Jiang H (2016) Quantitative trait locus analysis for pod- and kernel-related traits in the cultivated peanut (Arachis hypogaea L.). BMC Genet 17:25
Chu Y, Chee P, Culbreath A, Isleib TG, Holbrook CC, Ozias-Akins P (2019) Major QTLs for resistance to early and late leaf spot diseases are identified on chromosomes 3 and 5 in peanut (Arachis hypogaea). Front Plant Sci 10:883
Churchill GA, Doerge RW (1994) Empirical threshold values for quantitative trait mapping. Genetics 138:963–971
Clevenger J, Chu Y, Chavarro C, Agarwal G, Bertioli DJ, Leal-Bertioli SCM, Pandey MK, Vaughn J, Abernathy B, Barkley NA, Hovav R, Burow M, Nayak SN, Chitikineni A, Isleib TG, Holbrook CC, Jackson SA, Varshney RK, Ozias-Akins P (2017) Genome-wide SNP genotyping resolves signatures of selection and tetrasomic recombination in peanut. Mol Plant 10:309–322
Core Team R (2015) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Culbreath AK, Brenneman TB, Bondari K, Reynolds KL, McLean HS (1995) Late leaf spot, southern stem rot, and peanut yield responses to rates of cyproconazole and chlorothalonil applied alone and in combination. Plant Dis 79:1121–1125
De Oliveira Y, Sosnowski O, Charcosset A, Joets J (2014) BioMercator 4: a complete framework to integrate QTL, meta-QTL, and genome annotation. In: European conference on computational biology, Strasbourg
Dodia S, Rathnakumar A, Mishra G, Radhakrishnan T, Binal J, Thirumalaisamy P, Narendra K, Sumitra C, Dobaria J, Abhay K (2016) Phenotyping and molecular marker analysis for stem-rot disease resistance using F2 mapping population in groundnut. Int J Trop Agric 34:1135–1139
Dodia SM, Joshi B, Gangurde SS, Thirumalaisamy PP, Mishra GP, Narandrakumar D, Soni P, Rathnakumar AL, Dobaria JR, Sangh C, Chitikineni A (2019) Genotyping-by-sequencing based genetic mapping reveals large number of epistatic interactions for stem rot resistance in groundnut. Theor Appl Genet 132(4):1001–1016
Dunn OJ (1964) Multiple comparisons using rank sums. Technometrics 6:241–252
Fujita M, Fujita Y, Noutoshi Y, Takahashi F, Narusaka Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Crosstalk between abiotic and biotic stress responses: a current view from the points of convergence in the stress signaling networks. Curr Opin Plant Biol 9:436–442
Goffinet B, Gerber S (2000) Quantitative trait loci: a meta-analysis. Genetics 155:463–473
Gu Z, Gu L, Eils R, Schlesner M, Brors B (2014) circlize implements and enhances circular visualization in R. Bioinformatics 30(19):2811–2812
Guo Y, Khanal S, Tang S, Bowers JE, Heesacker AF, Khalilian N, Nagy ED, Zhang D, Taylor CA, Stalker HT, Ozias-Akins P, Knapp SJ (2012) Comparative mapping in intraspecific populations uncovers a high degree of macrosynteny between A- and B-genome diploid species of peanut. BMC Genom 13:608
Herselman L, Thwaites R, Kimmins F, Courtois B, Van Der Merwe P, Seal SE (2004) Identification and mapping of AFLP markers linked to peanut (Arachis hypogaea L.) resistance to the aphid vector of groundnut rosette disease. Theor Appl Genet 109:1426–1433
Holbrook CC, Culbreath AK (2007) Registration of ‘Tifrunner’peanut. Plant Regist 1(124):10–3198
Holbrook C, Isleib T, Ozias-Akins P, Chu Y, Knapp S, Tillman B, Guo B, Gill R, Burow MJPS (2013) Development and phenotyping of recombinant inbred line (RIL) populations for peanut (Arachis hypogaea). Peanut Sci 40:89–94
Hu X, Zhang S, Miao H, Cui F, Shen Y, Yang W, Xu T, Chen N, Chi X, Zhang Z (2018) High-density genetic map construction and identification of QTLs controlling oleic and linoleic acid in peanut using SLAF-seq and SSRs. Sci Rep 8:5479
Huang L, He H, Chen W, Ren X, Chen Y, Zhou X, Xia Y, Wang X, Jiang X, Liao B (2015) Quantitative trait locus analysis of agronomic and quality-related traits in cultivated peanut (Arachis hypogaea L.). Theor Appl Genet 128:1103–1115
Jogi A, Kerry JW, Brenneman TB, Leebens-Mack JH, Gold SE (2016) Identification of genes differentially expressed during early interactions between the stem rot fungus (Sclerotium rolfsii) and peanut (Arachis hypogaea) cultivars with increasing disease resistance levels. Microbiol Res 184:1–12
Johnson NL, Kemp AW, Kotz S (2005) Univariate discrete distributions. Wiley, New York
Khatri K, Kunwar S, Barocco R, Dufault NJPS (2017) Monitoring fungicide sensitivity levels and mycelial compatibility groupings of Sclerotium rolfsii Isolates from Florida peanut fields. Peanut Sci 44:83–92
Kokalis-Burelle N, Porter D, Rodriguez-Kabana R, Smith D, Subrahmanyam P (1997) Compendium of peanut diseases. American Phytopathological Society, Saint Paul
Li H, Ye G, Wang J (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175(1):361–374
Li H, Ribaut JM, Li Z, Wang J (2008) Inclusive composite interval mapping (ICIM) for digenic epistasis of quantitative traits in biparental populations. Theor Appl Genet 116(2):243–260
Li S, Wang J, Zhang L (2015) Inclusive composite interval mapping of QTL by environment interactions in biparental populations. PLoS ONE 10(7):e0132414
Little E (2015) Georgia plant disease loss estimates. The University of Georgia Cooperative Extension Bulletin, University of Georgia, Athens
Mamidi S, Miklas PN, Trapp J, Felicetti E, Grimwood J, Schmutz J, Lee R, McClean PE (2016) Sequence-based introgression mapping identifies candidate white mold tolerance genes in common bean. Plant Genome 9:2
Marrs KA (1996) The functions and regulation of glutathione S-transferases in plants. Annu Rev Plant Biol 47:127–158
McHale L, Tan X, Koehl P, Michelmore RW (2006) Plant NBS-LRR proteins: adaptable guards. Genome Biol 7:212
Meng L, Li H, Zhang L, Wang J (2015) QTL IciMapping: integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J 3(3):269–283
Monnahan PJ, Kelly JK (2015) Epistasis is a major determinant of the additive genetic variance in Mimulus guttatus. PLoS Genet 11:e1005201
Morrell PL, Buckler ES, Ross-Ibarra J (2012) Crop genomics: advances and applications. Nat Rev Genet 13:85
Pandey MK, Agarwal G, Kale SM, Clevenger J, Nayak SN, Sriswathi M, Chitikineni A, Chavarro C, Chen X, Upadhyaya HD (2017) Development and evaluation of a high density genotyping ‘Axiom_Arachis’ array with 58K SNPs for accelerating genetics and breeding in groundnut. Sci Rep 7:40577
Punja ZK (1988) Sclerotium (Athelia) Rolfsii, a pathogen of many plant species. In: Sidhu GS (ed) Genetics of plant pathogenic fungi. Advances in plant pathology. Academic Press, Cambridge, pp 523–534
Rajyaguru RH, Thirumalaisamy P, Patel KG, Thumar JT (2017) Biochemical basis of genotypic and bio-agent induced stem rot resistance in groundnut. Legume Res Int J 40:2
Rasheed A, Hao Y, Xia X, Khan A, Xu Y, Varshney RK, He Z (2017) Crop breeding chips and genotyping platforms: progress, challenges, and perspectives. Mol Plant 10:1047–1064
Schlötterer C (2004) The evolution of molecular markers—just a matter of fashion? Nat Rev Genet 5:63
Shapiro SS, Wilk MB (1965) An analysis of variance test for normality. Biometrika 52:591–611
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Sosnowski O, Charcosset A, Joets J (2012) BioMercator V3: an upgrade of genetic map compilation and quantitative trait loci meta-analysis algorithms. Bioinformatics 28:2082–2083
Sujay V, Gowda M, Pandey M, Bhat R, Khedikar Y, Nadaf H, Gautami B, Sarvamangala C, Lingaraju S, Radhakrishan T (2012) Quantitative trait locus analysis and construction of consensus genetic map for foliar disease resistance based on two recombinant inbred line populations in cultivated groundnut (Arachis hypogaea L.). Mol Breed 30:773–788
Van Loon LC, Rep M, Pieterse CM (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44:135–162
Van Ooijen JW (2006) JoinMap 4®, software for the calculation of genetic linkage maps in experimental populations. Kyazma BV, Wageningen
Vasconcellos RC, Oraguzie OB, Soler A, Arkwazee H, Myers JR, Ferreira JJ, Song Q, McClean P, Miklas PN (2017) Meta-QTL for resistance to white mold in common bean. PLoS ONE 12:e0171685
Vuong TD, Diers BW, Hartman GL (2008) Identification of QTL for resistance to Sclerotinia stem rot in soybean plant introduction 194639. Crop Sci 48:2209–2214
Wang C, Rutledge J, Gianola D (1994) Bayesian analysis of mixed linear models via Gibbs sampling with an application to litter size in Iberian pigs. Genet Sel Evol 26:91
Wang S, Basten C, Zeng Z (2012) Windows QTL cartographer 2.5. Department of Statistics, North Carolina State University, Raleigh
Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, Berlin
Yang J, Hu C, Hu H, Yu R, Xia Z, Ye X, Zhu J (2008) QTLNetwork: mapping and visualizing genetic architecture of complex traits in experimental populations. Bioinformatics 24:721–723
Zar JH (2005) Spearman rank correlation. Encycl Biostat 7:121
Zhao X, Han Y, Li Y, Liu D, Sun M, Zhao Y, Lv C, Li D, Yang Z, Huang L (2015) Loci and candidate gene identification for resistance to Sclerotinia sclerotiorum in soybean (Glycine max L. Merr.) via association and linkage maps. Plant J 82:245–255
Zhao Y, Zhang C, Chen H, Yuan M, Nipper R, Prakash C, Zhuang W, He G (2016) QTL mapping for bacterial wilt resistance in peanut (Arachis hypogaea L.). Mol Breed 36:13
Zhou X, Xia Y, Ren X, Chen Y, Huang L, Huang S, Liao B, Lei Y, Yan L, Jiang H (2014) Construction of a SNP-based genetic linkage map in cultivated peanut based on large scale marker development using next-generation double-digest restriction-site-associated DNA sequencing (ddRADseq). BMC Genom 15:351
Zhuang W, Chen H, Yang M, Wang J et al (2019) The genome of cultivated peanut provides insight into legume karyotypes, polyploid evolution and crop domestication. Nat Genet 51(5):865
Funding
This study was funded by the Florida Peanut Producers Association, the National Peanut Foundation and USDA National Institute of Food and Agriculture, Hatch Project 1011664.
Author information
Authors and Affiliations
Contributions
JW coordinated the research. YC, TGI, POA and CH developed the RIL population. ND and TB provided the inoculum and advised on the inoculation procedure. BT and JW supervised the phenotyping in Florida. YCT, HZ, ZP, XY, YL and JW conducted the experiments and collected the phenotypic data in Florida. TB and POA supervised the research in Georgia. RC and TB conducted the experiments and collected the phenotypic data in Georgia. CC and POA conducted the linkage map analysis. ZL analyzed the whole data sets and prepared the manuscript draft. All authors revised the manuscript and read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Availability of data and material
Data supporting the results are provided in the additional files and analyzed/organized in tables. The original data and the materials are available upon reasonable request to the corresponding author at wangjp@ufl.edu.
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Communicated by Albrecht E. Melchinger.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
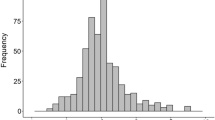
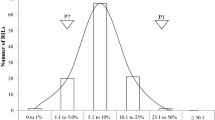
Fig. S1
The distributions of phenotypic data from all datasets (TIFF 44237 kb)
Table S1
Description of experiments in this study (XLSX 10 kb)
Table S2
Kruskal-Wallis ANOVA test for different datasets (XLSX 11 kb)
Table S3
The correlation of phenotypic data from all experiments (XLSX 12 kb)
Table S4
Information of the linkage groups used for stem rot QTL mapping (XLSX 10 kb)
Table S5
The original additive QTLs detected for stem rot resistance by three types of software (XLSX 19 kb)
Table S6
Summary of original additive QTLs’ contribution to phenotype variations (XLSX 10 kb)
Table S7
The consensus additive QTLs detected for stem rot resistance after QTL meta-analysis (XLSX 15 kb)
Table S8
The epistatic QTLs for stem rot resistance detected by QTL IciMapping and QTLNetwork (XLSX 17 kb)
Table S9
The candidate genes identified within all stem rot QTL intervals (XLSX 27 kb)
Rights and permissions
About this article
Cite this article
Luo, Z., Cui, R., Chavarro, C. et al. Mapping quantitative trait loci (QTLs) and estimating the epistasis controlling stem rot resistance in cultivated peanut (Arachis hypogaea). Theor Appl Genet 133, 1201–1212 (2020). https://doi.org/10.1007/s00122-020-03542-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03542-y