Summary
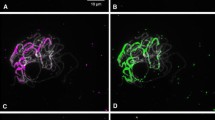
We have employed a combination of techniques to examine the organization of pea 5S rRNA genes. These include the analysis of length variant interspersion patterns in cosmid clones, sequence analysis, Southern analysis of both conventional gels and field inversion gels and in situ hybridization. From these analyses we conclude that the 5S rRNA genes of pea are arranged in three major tandem arrays which are represented by three large EcoRI fragments and that these correspond to the three sites of in situ hybridization in the haploid pea complement
Similar content being viewed by others
References
Appels R, Gerlach WL, Dennis ES, Swift H, Peacock WJ (1980) Molecular and chromosomal organization of DNA sequences coding for the ribosomal RNAs in cereals. Chromosoma 78:293–311
Barsacchi-Pilone G, Nardi I, Batistoni R, Aadronico F, Beccari E (1974) Chromosome location of the genes for 28S 18S and 5S ribosomal RNA in Triturus marmoratus (Amphibia Urodela). Chromosoma 49:135–153
Bäumlein H, Wobus U (1976) Chromosomal localization of ribosomal 5 S RNA genes in Chironomus thummi by in situ hybridization of iodonated 5S RNA. Chromosoma 57:199–204
Bennett MD, Smith JB (1976) Nuclear DNA amounts in angiosperms. Philos Trans R Soc Lond [Biol] 274:227–274
Carle GF, Olsen MV (1984) Separation of chromosomal DNA molecules from yeast by orthogonal-field-alternation gel electrophoresis. Nucleic Acids Res 12:5647–5664
Carle GF, Franke M, Olsen MV (1986) Electrophoretic separation of large DNA molecules by periodic inversion of the electric field. Science 232:65–68
Chu G, Vollrath D, Davis RW (1986) Separation of large DNA molecules by contour clamped homogeneous electric fields. Science 234:1582–1585
Denhardt DT (1966) A membrane-filter technique for the detection of complementary DNA. Biochem Biophys Res Commun 23:641–646
Duggleby RG (1981) A nonlinear regression program for small computers. Anal Biochem 110:9–18
Ellis THN, Davies DR, Casleton JA, Bedford ID (1984) The organization and genetics of rDNA length variants in peas. Chromosoma 91:74–81
Ellis THN, Cleary WG, Burcham KWG, Bowen BA (1987) Ramped field inversion gel electrophoresis: a cautionary note. Nucleic Acid Res 15:5489
Erdmann VA, Wolters J (1986) Collection of published 5S, 5.8S and 4.5S RNA sequences. Nucleic Acids Res 14:r1-r59
Feinberg A, Vogelstein B (1983) A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem 132:6–13
Garson JA, van den Berghe JA, Kemshead JT (1987) Novel nonisotopic in situ hybridization technique detects small (1 kb) unique sequences in routinely G-banded human chromosomes: fine mapping of N-myc and β-NGF genes. Nucleic Acids Res 15:4761–4770
Goldsbrough PB, Ellis THN, Lomonossoff GP (1982) Sequence variation and methylation of the flax 5S RNA genes. Nucleic Acids Res 10:4501–4514
Gruenbaum Y, Naveh-Many T, Cedar H, Razin A (1981) Sequence specificity of methylation in higher plant DNA. Nature 292:860–862
Hemleben V, Werts D (1988) Sequence organization and putative regulatory elements in the 5S rRNA genes of two higher plants (Vigna radiata and Matthiola incana). Gene 62:165–169
Henderson AS, Atwood KC, Yu MT, Warburton D (1976) The site of 5S RNA genes in primates I: The great apes. Chromosoma 56:29–32
Hörz W, Zachau HG (1977) Characterization of distinet segments in mouse satellite DNA by restriction nucleases. Eur J Biochem 73:383–392
Hutchison N, Pardue ML (1976) The mitotic chromosomes of Notophthalmus (=Triturus) viridescens: localization of C banding regions and DNA sequences complementary to 18S 28S and 5S ribosomal RNA. Chromosoma 53:51–69
Ish-Horowitz D, Burke JF (1981) Rapid and efficient cosmid cloning. Nucleic Acids Res 9:2989–2998
Junakovic N (1980) Variability in the molecular organization of the 5 S RNA genes among strains of Drosophila melanogaster. Nucleic Acids Res 8:3611–3622
Kessler C, Höltke H-J (1986) Specificity of restriction endonucleases and methylases—a review (Edition 2). Gene 47:1–153
León PE (1976) Molecular hybridization of iodinated 4S, 5S and 18S+28S RNA to salamander chromosomes. J Cell Biol 69:287–300
León PE, Kezer J (1978) Location of 5S RNA genes on chromosomes of plethadontid salamanders. Chromosoma 65:213–230
Long EO, Dawid IB (1980) Repeated genes in Eukaryotes. Annu Rev Biochem 49:727–764
Maniatis T, Jeffrey A, Kleid DG (1975) Nucleotide sequence of the rightward operator of phage λ. Proc Natl Acad Sci USA 72:1184–1188
Maniatis T, Fritsch EF, Sambrook J (1982) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor New York
McMahon M, Stamenkovich D, Petes TD (1984) Tandemly arranged variant 5S ribosomal RNA genes in the yeast Saccharomyces cerevisiae. Nucleic Acids Res 12:8001–8016
Murray NE, Brammar WJ, Murray K (1977) Lambdoid phages that simplify the recovery of in vitro recombinants. Mol Gen Genet 150:53–61
Norrander J, Kempe T, Messing J (1983) Construction of improved M13 vectors using oligonucleotide-directed mutagenesis. Gene 26:101–106
Pardue ML, Brown DD, Birnstiel ML (1973) Location of the genes for 5 S ribosomal RNA in Xenopus laevis. Chromosoma 42:191–203
Pukkila PJ (1975) Identification of the lampbrush chromosome loops which transcribe 5S ribosomal RNA in Notophthalamus (triturus) viridescens. Chromosoma 53:71–89
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain terminating inhibitors. Proc Natl Acad Sci USA 74:5463–5467
Scalenghe F, Turco E, Edström JE, Pirrotta V, Melli M (1981) Microdissection and cloning of DNA from a specific region of Drosophila melanogaster polytene chromosomes. Chromosoma 82:205–216
Schwartz DC, Cantor CR (1984) Separation of yeast chromosomesized DNAs by pulsed field gradient gel electrophoresis. Cell 37:67–75
Sealey PG, Southern EM (1982) Gel electrophoresis of DNA. In: Rickwood D, Hames BD (eds) Gel Electrophoresis of nucleic acids: a practical approach. IRL Press, Oxford, England
Simpson PR, Newman MA, Davies DR (1988) Detection of legumin gene DNA sequences in pea by in situ hybridization. Chromosoma 96:454–458
Southern EM (1979a) Gel electrophoresis of restriction fragments. Methods Enzymol 68:152–176
Southern EM (1979b) Measurement of DNA length by gel electrophoresis. Anal Biochem 100:319–323
Southern EM, Anand R, Brown WRA, Fletcher DS (1987) A model for the separation of large DNA molecules by crossed field gel electrophoresis. Nucleic Acids Res 15:5925–5943
Szabo P, Lee MR, Elder FB, Prensky W (1978) Localization of 5S RNA and rRNA genes in the Norway Rat. Chromosoma 65:161–172
Thomas PS (1980) Hybridization of denatured RNA and small DNA fragments transferred to nitrocellulose. Proc Natl Acad Sci USA 77:5201–5205
Tyler-Smith C, Brown WRA (1987) Structure of the major block of alphoid satellite DNA on the human Y chromosome. J Mol Biol 195:457–470
Vandenberghe A, Chen M-W, Dams E, de Bare R, de Roeck E, Huysmans E, de Wachter R (1984) The corrected nucleotide sequences of 5S RNA secondary structure and evolution. FEBS Lett 171:17–23
Wen W-N, León PE, Hague DR (1974) Multiple sites for 5S and 18S+28S RNA on chromosomes of Glyptotendipes barbipes (Staeger). J Cell Biol 62:132–144
Wieslander K, Lambert B, Egyhàzi E (1975) Localization of 5S RNA genes in Chironomus tentans. Chromosoma 51:49–56
Wimber DE, Steffensen DM (1970) Localization of 5S RNA genes on Drosophila chromosomes by RNA-DNA hybridization. Science 170:639–641
Wimber DE, Duffey PA, Steffensen DM, Prensky W (1974) Localization of the 5S RNA genes in Zea mays by RNA-DNA hybridization in situ. Chromosoma 47:353–359
Author information
Authors and Affiliations
Additional information
Communicated by R. Hagemann
Rights and permissions
About this article
Cite this article
Ellis, T.H.N., Lee, D., Thomas, C.M. et al. 5S rRNA genes in Pisum: Sequence, long range and chromosomal organization. Mol Gen Genet 214, 333–342 (1988). https://doi.org/10.1007/BF00337732
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF00337732