Abstract
Lung cancer is one of the cancers with highest morbidity and mortality rates and the metastasis of lung cancer is a leading cause of death. Mechanisms of lung cancer metastasis are yet to be fully understood. Herein, we demonstrate that mice deficient for REGγ, a proteasome activator, exhibited a significant reduction in tumor size, numbers, and metastatic rate with prolonged survival in a conditional Kras/p53 mutant lung cancer model. REGγ enhanced the TGFβ-Smad signaling pathway by ubiquitin-ATP-independent degradation of Smad7, an inhibitor of the TGFβ pathway. Activated TGFβ signaling in REGγ-positive lung cancer cells led to diminished expression of E-cadherin, a biomarker of epithelial–mesenchymal transitions (EMT), and elevated mesenchymal markers compared with REGγ-deficient lung cancer cells. REGγ overexpression was found in lung cancer patients with metastasis, correlating with the reduction of E-Cadherin/Smad7 and a poor prognosis. Overall, our study indicates that REGγ promotes lung cancer metastasis by activating TGF-β signaling via degradation of Smad7. Thus, REGγ may serve as a novel therapeutic target for lung cancers with poor prognosis.
Similar content being viewed by others
Introduction
Lung cancer is the most common cancer worldwide and has a high mortality. Approximately 85% of lung cancers are non-small cell lung cancers (NSCLC) in both smokers and nonsmokers [1]. A number of genetic alterations including KRAS (Kristen Rat Sarcoma viral oncogene) have been found in NSCLC [2, 3]. KRAS mutations represent the most common molecular change in NSCLC [4]. Mutation of p53 also has been frequently reported in lung cancer (50–75%) [5]. Concomitant KrasLSL-G12D (hereafter called KrasG12D) [6] and Trp53f/f[7] expression in NSCLC mouse models leads to a histologically and invasively more “humanized” version of lung cancer [8]. Effective therapies against KRAS or p53 mutations have yet to be developed [9].
It is known that cellular epithelial–mesenchymal transition (EMT) is associated with cancer metastasis, and activation of EMT signaling (loss of E-cadherin) contributes to therapeutic resistance [10]. EMT signaling can be activated by various factors including transforming growth factor-β (TGFβ), Notch, Wnt, or EMT-inducing transcription factors (Snail, Slug, Twist, Zeb1, and Zeb2) [11, 12] TGFβ can act to suppress tumorigenesis, but it induces tumorigenesis by promoting EMT once cancers are established [13]. Smads are downstream mediators activated by TGF-β [14] and Smad7, a critical component in the TGFβ pathway, negatively regulates TGFβ signaling in tumor progression [15].
REGγ (also known as PSME3, PA28γ, or Ki antigen) belongs to the REG or 11S family of proteasome activators that have been shown to bind and activate the 20S proteasome based on their ability to promote the degradation of model peptide substrates. It can promote the degradation of SRC3, p21Waf1/Cip1, p16Ink4A (cyclin-dependent kinase inhibitor 2A), and p19ARF proteins in an ubiquitin- and ATP-independent manner [16,17,18]. REGγ is overexpressed in numerous tumors [19,20,21,22], with a marked increase in late-stage lung cancer and liver cancer [23]. Evidence suggests that REGγ is critical in tumor initiation and progression, however its role in lung cancer development, particularly for cancer metastasis, is unknown.
In this study, we used adenoviral CRE (adeno-Cre) controlled activation of genetically engineered KrasG12D and inactivation of p53 (p53f/f) to induce lung cancers in mice with REGγ-WT (wild type) or REGγ-null background. REGγ deletion attenuated lung cancer progression and metastasis in activated p53f/f;KrasG12D mutant (PK) mice. Molecularly, REGγ suppressed E-cadherin expression by enhancing TGFβ signaling via degradation of Smad7. Clinical implication of REGγ in the EMT signal pathway has been demonstrated with correlation to lung cancer prognosis. Taken together, we conclude that REGγ promotes tumorigenesis and metastasis in lung cancer.
Materials and methods
Reagents
Antibodies were purchased from Invitrogen (REGγ/PA28γ), Abcam (Smad7/MADH7), Sigma (β-actin, Flag, HA), and Cell Signaling Technology (E-cadherin, EMT Antibody Sampler Kit, P-Smad2/3, Smad2/3, Snai1, Slug, Vimentin, IR DyeR 680 goat anti-Mouse, IR DyeR 800 goat anti-Mouse, IR DyeR 800 goat anti-Rabbit). Cycloheximide was purchased from Amresco (Solon, OH). Mg132 (CAS:133407-82-6) was purchased from MedChemExpress. SMAD7-set siRNA/shRNA/RNAi Lentivector (Human) was purchased from Applied Biological Materials (abm) Inc.
Cell culture and expression constructs
A549 and PC9 (ATCC) cell lines were purchased from ATCC. PK and RPK cell lines were generated from PK and RPK mouse lung tumors. All lung cancer cell types were grown in 1640 (Invitrogen), 10% FBS (Invitrogen), and penicillin/streptomycin (Invitrogen). The 293-REGγ-inducible cell lines were previously generated [24] and cultured in DMEM (Invitrogen) supplemented with 10% fetal bovine serum. pSG5-HA-REGγ plasmid was previously constructed. The pCDNA-Flag-hSmad7 plasmid was constructed in this study The primer of Smad7 was designed in accordance with pcDNA3.1 vector sequence and syntyetized by Shanghai BioSune Biological Technology Co., Ltd. The Smad7 sequences with double cleavage sites were amplified by PCR and connected to the pcDNA3.1 vector after double enzyme cutting. The plasmid that expressed Smad7 successfully was screened and identified. All the constructs were verified by DNA sequencing.
Isolation of primary lung cancer cells from mouse tumors
Fresh lung cancer tumor tissues and appropriate amount of PBS were put in petri dish, the blood (blood clots), fat, necrotic tissue, and connective tissue on the tissue use were removed by ophthalmic tweezers and ophthalmic scissors, and collected the tumor cell rich area, and washed it twice with PBS. Put the tumor tissues into a new petri dish with RPMI 1640 medium, and cut them into fragments about 1 mm3 by ophthalmic scissors. These fragments should be transfered into a 15 ml centrifuge tube, and rinsed with PBS for several times. The tissue blocks were transferred into culture bottles with 5 ml collagenase(I and IV, 1 mg ml−1) and double antibiotics (diluted to 2×), and the fragments of tissues were blown away on a constant temperature shaking bed at 37 °C and 150 r min−1 rotate speed. When the tissue mass had good light transmittance under the microscope and presented a flocculent state, it was transferred to a 15 ml centrifuge tube, which was centrifuged at 1300 r min−1 for 5 min, and the supernatant was discarded. Washed them by PBS for several times and screened the single cells by cell sieve. And then these single cells should be transferred into the culture dish with RPMI 1640 complete culture medium (including 1× double antibiotics). The isolated primary cells need to be subcultured many times before immortalized cells can be obtained.
Genetic mouse models
REGγ knockout (KO) mouse with C57BL/6 genetic background was kindly provided by Dr John J. Monaco at University of Cincinnati and was backcrossed at our facility for more than ten generations. Loxp-Stop-Loxp KrasG12D/FVB/129 p53loxp/loxp mouse (PK mouse) was previously generated [7, 25]. We crossed PK mice with REGγ-null mice to get the REGγ−/−; p53flox/flox; KrasG12D mice (RPK). The 6–8 weeks old PK or RPK mouse was inoculated with 1 × 106 pfu adenoviral Cre (adeno-Cre) by intranasal inhalation to activate oncogenic P53 and KrasG12D in the lungs. Animal studies were conducted under specific pathogen free conditions and experimental mice were handled according to the ethical and scientific standards by the Animal Center at the institute following procedures approved by the Institutional Animal Care and Use Committee. All surgeries were performed under Avertin anesthesia to minimize suffering. After euthanasia, organs, including heart, liver, spleen, kidney, stomach, intestine, spine, brain, breast, skin, and testis or ovary, were undergone gross inspection for metastases and verification under microscope. Metastatic lesions were harvested and validated by histological examination by a pathologist.
Xenograft animal model and in vivo imaging system analysis
After stable transfection with pGL4.17 luciferase vector, A549 shN, A549 shR, and A549 shR siSmad7 cells were delivered to 4–6 weeks old BALB/c nude mice by left ventricular injection of 1 × 105 cells in 100 μl per spot. Four weeks after injection, these nude mice were scanned with an in vivo imaging system (IVIS) and sacrificed for the determination of tumor numbers. Tumor metastasis was visualized by IVIS.
RNA interference and real-time quantitative RT-PCR
The sequence of siRNA against REGγ is 5ʹ-CAGAAGACTTGGTGGCAAA-3ʹ. The siRNA for Smad7 1# is 5ʹ-CCTCCTGCTGTGCAAAGTGTTCAGGTGGC-3ʹ and for Smad7 #2 is 5ʹ-GGTGGCATACTGGGAGGAGAAGACGAGAG-3ʹ. Tissues and cells were homogenized in 1 ml RNAisoTM Plus lysis buffer (TAKARA). Total RNA was extracted and 2 mg RNA was transcribed into cDNA with M-MLV reverse transcriptase (Invitrogen) following the manufacturer’s instruction. The gene-specific primers are as follows: REGγ sense primer, 5ʹ-ACAAGTGAGGCAGAAGAC-3ʹ; REGγ antisense primer, 5ʹ-ATCATGGCTATTGGTGAG-3ʹ; E-cadherin sense primer, 5ʹ-ATTTTTCCCTCGACACCCGAT-3ʹ; E-cadherin antisense primer, 5ʹ-TCCCAGGCGTAGACCAAGA-3ʹ; β-actin sense primer, 5ʹ- CGTCATACTCCTGCTTGCTG-3ʹ; β-actin antisense primer, 5ʹ-GTACGCCAACACAGTGCTG-3ʹ.
Immunohistochemistry and H&E staining
Unnamed human lung cancer (including metastatic cancer originated from lung) or mouse lung cancer tissues were fixed with 4% paraformaldehyde for 12 h, transferred into gradient ethanol, rolled, processed, and embedded into paraffin. Tumors were cut in 4 mm sections with a microtome (Leica, Germany) followed by immunohistochemistry or H&E staining as described [21].
Analysis of human lung cancer samples
Acquisition of human samples was approved by the independent ethics committee at the Changzheng hospital of Shanghai. All primary lung tumor, metastatic cancer, or control (routine analysis) samples were from surgically removed specimen. All clinical samples were devoid of personal information. The sections were stained with specific antibodies and counterstained with haematoxylin. The staining intensity by specific antibodies was evaluated as negative (−), weak staining (+), moderate/strong staining ( + + ) and very strong staining ( + + + ) and quantified as described [23].
Western blotting analysis
The cells or tissues were resuspended in Lysis buffer (50 mM Tris-HCl [pH 7.5], 150 mM NaCl, 1 mM EDTA, 0.5% NP-40) and resolved in 8–12% gradient SDS gels. Separated proteins were transferred to nitrocellulose membranes and immunoblotted with primary antibodies specific for REGγ, Smad7, E-cadherin, N-cadherin, claudin-1, Smad2/3, p-Smad2/3, Vimentin, Snail, Slug, or β-actin (1:1000–1:2000 dilutions) overnight at 4 °C. After incubation with a fluorescent-labeled secondary antibody (1:5000 dilutions), specific signals for proteins were visualized by the LI-COR Odyssey Infrared Imaging System.
Immunoprecipitation
Cells were transfected with constructs or treated as described in the figure legends. Cells were then scraped into ice-cold PBS and lysed with CHAPS lysis buffer containing 20 mM PIPEs (pH 7.2), 5 mM EDTA, 10 mM sodium glycerophosphate, 10 mM sodium pyrophosphate, 0.3% CHAPS, and protease inhibitors for 10–20 min on ice-bath-ultrasound; then centrifuged for 10 min at 12,000 r.p.m. Specific proteins were immunoprecipitated by anti-Flag M2 Affinity Gel or HA beads, followed by three washes with wash buffer (50 mM Tris-HCl, pH 7.5, 1 mM EDTA, 150 mM NaCl, 10% glycerol, and protease inhibitors). The pellet was suspended in SDS sample buffer and resolved in 10% gradient SDS gels for Western blotting.
In vitro proteolytic analysis
The in vitro degradation system contained 5 μl Smad7 protein, which was translated in vitro by TNT® T7 Quick Coupled Transcription/Translation System (Promega), 1 μg purified REGγ heptamers, 0.25 μg purified 20S proteasome indicated in 50 μl reaction volume at 30 °C for various time. An aliquot of the reaction was analyzed by western blotting.
MTT assay, transwell assay, and wound-healing assay
MTT assay was performed by seeding 2.5 × 103 cells in a 96-well plate and cultured for 0, 24, 48, 72, 96 h. Then cells were incubated with MTT detection solution at 37 °C for 4 h and terminated by DMSO. Absorbance (490 nm) was measured and analyzed. Transwell assay was performed in 24-well PET inserts (Millipore, 8.0-μm pore size) for cell migration. 5 × 104 cells were plated with serum-free media in the upper chamber of transwell inserts (two replicas for each sample). The inserts were then placed into 10% serum media for 6 h of migration. Cells in the upper chambers were removed with a cotton swab, and migrated cells were fixed in paraformaldehyde 4% and stained with crystal violet 0.5%. Filters were photographed, and the total migrated cells were quantitated.
For wound-healing assay, cells cultured in the 12-well plate were scratched by a small pipette tip to produce a “wound”, and monitor the “healing” after 18 h. Every experiment was repeated independently three times.
F-actin staining
The cover slides were treated by 0.01% Poly-L-Lysine (Sigma) for 30 min and 10 μg/ml Fibronectin (Corning) for 12 h, placed in a 24-well plate, and inoculated with 5 × 104 cells per well. The cells were switched to nonserum medium for 24 h, followed by 10% serum medium for 30, 60, and 120 min. The cells were washed with PBS at 37°C, fixed by 4% paraformaldehyde, permeated by 0.25% Triton X-100, stained by rhodamine-phalloidin (Invitrogen) and DAPI (avoid light). The cover slips were then mounted on slides for microscopic visualization.
Statistical analysis
Prism software (GraphPad Software) was used for statistical analyses. The intensity of the western blot results was analyzed by densitometry using ImageJ software. Values were shown as mean ± s.e.m. Statistical significance between two samples was determined with two-tailed Student’s t tests.
Study approval
All mice studies were approved and used in accordance with institutional guidelines (East China Normal University). Human specimens used in this study have been approved by the Ethics Committee of Shanghai Changzheng Hospital.
Results
REGγ abrogation retards lung cancer progression and metastasis in PK mice
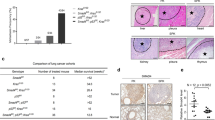
To investigate the contribution of REGγ to lung cancer progression and metastasis using a conditional mouse lung cancer model, we crossed PK mice bearing p53f/f;KrasG12D alleles with REGγ-null mice to generate a cohort differing only in REGγ activity. Approximately 16 weeks after nasal delivery of Cre recombinase, PK mice developed significant numbers of lung adenocarcinoma as described [7, 25] accompanied with metastatic lesions to bone, liver, thorax, kidney, and peritoneum (Fig. 1a, b, Fig. S1A–1E). However, the REGγ−/−;p53f/f;KrasG12D (RPK) mice exhibited fewer and smaller tumors than those in PK mice, with significantly reduced metastases (Fig. 1a, b, Fig. S1A–1E). Statistical analysis showed 29.2% metastases in PK vs. 11.1% in RPK mice (Fig. 1b). In addition, RPK mice had prolonged median survival compared with PK mice (Fig. 1c, d). These data suggest that REGγ functions as an oncogenic promoter for lung cancer formation and metastasis in vivo.
REGγ abrogation retards lung cancer progression and metastasis in PK mice. a Representative images of H&E stained lung cancer metastatic tissues, including liver, thorax, kidney, lymph node, and heart from PK and thorax from RPK mouse. Eight-week-old PK and RPK mouse were all treated 1 × 106 pfu adeno-Cre by nose-dripping for 16–24 weeks. Asterisk indicates the area of metastatic tumors. Scale bar, 50 μm (magnification, ×20). b Statistical analyses of lung cancer metastases in PK and RPK mouse. PK mice had more lung cancer metastasis sites than RPK mice. n = 48 PK mouse group; n = 45 RPK mouse group. *p < 0.05, as determined by CHITEST. c, d Survival statistics of PK and RPK mice with lung cancer. n = 43 in both PK and RPK mouse groups. Median latency refers to the survival time on average (in weeks) after adeno-Cre treatment, estimated by Kaplan–Meier analysis. RPK mice lived longer than PK mice. *P < 0.05. The survival range for all the mice was between 15 and 30 weeks post adeno-Cre infection
REGγ promotes lung cancer cell migration and invasion
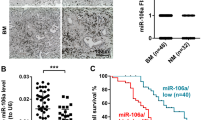
To define the impact of REGγ on lung cancer cell migration and invasion, we generated A549 cell lines stably expressing a control shRNA (shN) or a REGγ-specific shRNA (shR). Within 24 h, REGγ-silenced cells had minimal growth defect compared to A549-shN cells (Fig. S2A), similar to a different lung cancer cell line, PC9, with REGγ silencing, only displaying growth difference at day 4 (Fig. S2B). Using Transwell Matrigel assays to measure migration and invasion, we found higher metastatic potential in A549-shN cells compared with A549-shR cell (Fig. 2a). Also, transient knockdown of REGγ in PC9 cells attenuated invasion/migration capabilities in transwell invasion and wound-healing assays (Fig. S2C–D). Among the indicators for more invasive cells is the formation of lamellipodia, which plays a major role in cell migration. Following serum stimulation, A549-shN cells revealed more F-actin stained leading-edge structures (lamellipodia) than were seen in A549-shR cells (Fig. 2b, Fig. S2E–F). Wound-healing assays consistently validated metastatic indices in A549-shN cells compared with A549-shR cells (Fig. 2c).
REGγ promotes lung cancer cell migration and invasion. a REGγ is important for lung cancer cell migration and invasion in Transwell Matrigel. 2 × 104 A549(shN/R) cells were seeded on each transwell insert. The migrating cells were fixed and stained with crystal violet; the number of migrated cells was quantified. A549 shN cells migration ability better than A549 shR cells. Data represent mean ± s.e.m.; ***p < 0.001, as determined by Student’s t test. Scale bar, 100 μm (magnification, ×10). b REGγ positively contributes to the formation of lamellipodia. A549(shN/R) cells were serum-starved 24 h before stimulation with serum for 0.25, 0.5, 1, 2 h at 37 °C. Cells were then fixed and stained with rhodamine-phalloidin (Red, F-actin) and DAPI (Blue, nucleus). The speed of cytoskeleton reconstruction, the number and length of cilia and pseudopodium, the change of cell morphology all represent the cell migration activity. The white triangles indicate cilia and pseudopodium. A549(shN)cells can form cilia and pseudopodium more and faster than A549 shR cells. It means that A549(shN) cells had greater athletic ability. Scale bar, 25 μm (magnification, ×40). c REGγ promotes wound healing in cell culture. Cells were made a wound between the two red dashed lines, the area of the two red dashed lines represents the level of wound healing and the cell migration activity. Wound-healing percentage is the ratio of the wound-healing area and the primary wound area. A549(shN) cells’ wound healing ability better than A549(shR) cells’. Scale bar, 100 μm (magnification, ×10). d–f The role of REGγ in murine lung cancer cell migration/invasion. Primary lung cancer cells were isolated from lung tumors in PK and RPK mice and immortalized. Transwell assays (d), rhodamine-phalloidin staining assay (e) and wound-healing (f) experiments were performed in PK and RPK lung tumor cells. These results showed that PK cells were stronger on migration and invition than RPK cells. Data represent means ± s.e.m. *p < 0.05; **p < 0.01; ***p< 0.001, as determined by Student’s t test
To substantiate in vivo action of REGγ on cell migration/invasion at cellular levels, we isolated primary lung cancer cells from PK and RPK mouse tumors to generate immortalized PK and RPK cell lines. Metastatic capacity of the mouse tumor-derived cells was analyzed by transwell invasion, F-actin staining, and wound-healing assays in PK and RPK cells. Consistent with the observation in A549 cells, REGγ depletion in mouse tumor cell significantly reduced invasion/migration in RPK cells (Fig. 2d–f, Fig. S2G–H). Our observations support the conclusion that REGγ enhances lung cancer cell migration/invasion.
REGγ inhibits the expression of E-cadherin in vitro and in vivo
Loss of E-cadherin gene expression causes dysfunction of cell junction system, allowing cancer cell invasion and metastasis [26]. To elucidate the underlying mechanism by which REGγ regulates lung cancer cell invasion and metastasis, we investigated expression of E-cadherin in various lung cancer cell lines by alteration of REGγ levels. Compared with A549-shN cells, A549-shR cells had an increased expression of E-cadherin at mRNA and protein levels (Fig. 3a, b), suggesting a REGγ-dependent regulation of E-cadherin. Using PC9 cells transfected with an siRNA against REGγ, we also observed an increase in E-cadherin (Fig. 3c, d). With the murine lung tumor-derived cells, we detected higher levels of E-cadherin in RPK than in PK cells, also with concurrent increase in mRNA and protein levels of E-cadherin (Fig. 3e, f). In addition, RPK tumors had elevated E-cadherin by western blot analysis compared to tumors from PK mice (Figs. 4c, 3g). These results indicate that REGγ suppresses the expression of E-cadherin in vitro and in vivo.
REGγ inhibits the expression of E-cadherin in vivo and in vitro. a The protein levels of E-cadherin were tested by western blotting analysis in A549 control cells (A549 shN) and A549 REGγ knockdown cells (A549 shR). b The mRNA levels of E-cadherin were examined by real-time quantitative RT-PCR in the A549 shN/shR cells. A549 shR cells expressed more E-cadherin at both protein and mRNA level. c. After transient knockdown of REGγ in PC9 cells, the protein levels of E-cadherin were detected by western blotting in PC9 siN and siR cells. d Quantitative RT-PCR analysis of E-cadherin mRNA expression in PC9 siN and siR cells. The expression of E-cadheirn was inhibited by REGγ in PC9 cells. e–f The expression of E-cadherin was detected in immortalized PK and RPK tumor cells. g Quantitative analysis of the E-cadherin mRNA expression in the PK and RPK mouse lung tumors. Whether in mouse lung tumor cells or in mouse lung tumor tissuses, the expression of E-cadherin was highter in RPK group than PK group. Each spot represents a sample from an individual mouse, n = 10. Data represent means ± s.e.m.; **p < 0.01; ***p < 0.001, as determined by Student’s t test
REGγ promotes TGFβ-induced EMT. a A549 shN and shR cells were starved for 24 h before being treated with or without TGFβ1(5 ng ml−1) for 12 h, and then the cells were analyzed for the expression of EMT-related factors and components in TGFβ/Smads pathway by western blotting. b PK and RPK cells were examined for factors related to EMT and TGFβ/Smad signaling pathway as in a. EMT pathway was stimulated by TGFβ1 but inhibites by REGγ. c The protein expression of E-cadherin and Smad7 in PK and RPK mouse lung tumors. Each lane represents a sample from an individual mouse. d The Smad7 mRNA expression of PK and RPK mouse lung tumors was evaluated by real-time quantitative PCR, the difference of Smad7 mRNA expression is not significant between PK and RPK mouse lung tumors. n = 10; data represent means ± s.e.m.; p > 0.05, no significant correlation, as determined by Student’s t test
REGγ accentuates TGFβ-induced EMT
It is known that Smad2/Smad3 are critical mediators for TGFβ-induced EMT during tumor progression [27,28,29]. To investigate whether REGγ could modulate the expression of E-cadherin through the TGFβ/Smad pathway, we performed a high-throughput proteomic screen of REGγ targets or effectors by antibody array analysis (FullMoon BioSystem) [30]. Among the top ten proteins differentially expressed in REGγ WT and REGγ KO MEFs, we found significantly elevated levels of active Smad3 (phosphor-Ser 204) in WT compared with KO cells [31]. We validated the finding that p-Smad2/3 levels were higher in A549-shN or PK cells than in A549-shR or REGγ-deficient PK cells (while total Smad2/3 levels were unchanged). These differences increased following TGFβ treatment (Fig. 4a, b), suggesting that REGγ may potentiate TGFβ signaling by Smad2/3 activation.
Transcription factors involved in EMT transition, such as Snai1 and Slug, are downstream targets of TGFβ, mediating transcriptional inhibition of E-cadherin [32, 33]. In lung cancer cells with REGγ depletion, we found concurrent reduction in Slug and Snai1 expression and decreased protein levels, comparing the REGγ-positive and REGγ-deficient cells. These differences were increased in the presence of TGFβ. Intriguingly, we discovered a positive correlation between Smad7, an inhibitor of TGFβ signaling [15], and E-cadherin, but an inverse relationship between Smad7 and Snai1/Slug levels (Fig. 4a), suggesting Smad7 as a potential factor bridging the regulation of TGFβ pathways by the REGγ proteasome. We validated our observations that REGγ impinged on expression of E-cadherin, Snai1, and Slug in immortalized PK and RPK cells (Fig. 4b). Furthermore, IHC data demonstrated a positive correlation between REGγ and TGFβ/Smad pathway components, but a negative association of REGγ with E-cadherin and Claudin-1 (a major constituent of the tight junction complexes) in lung tumor samples from PK and RPK mice (Fig. S3A–B).
To gain additional insights into the co-regulation of REGγ and EMT markers, we analyzed the mRNA levels for E-cadherin, Smad7, Snai1, and Slug. REGγ affected both protein and mRNA expression of E-cadherin, Smad7, Snai1, and Slug (Fig. S4A). However, Smad7 was only differentially regulated in REGγ WT vs. REGγ-depleted cells at protein, but not mRNA, levels (Fig. 4c, d), suggesting that REGγ may regulate Smad7 post transcriptionally.
REGγ mediates ubiquitin-independent degradation of Smad7
Given the alteration of Smad7 protein levels, but not mRNA levels, between REGγ WT vs. REGγ-depleted mouse and human lung cancer cells (Fig. 4a, b, Fig. S4B–D), we investigated this phenomenon in another human lung cancer cell line, PC9. Again, we noticed a dramatic increase in Smad7 protein, but not mRNA, levels when REGγ was silenced (Fig. 5a, b). Also, mouse lung tumors from RPK mice showed a higher Smad7 protein level than those from PK mice (Fig. 4c, d). To determine if Smad7 could be a target of the REGγ proteasome, we looked for physical interactions between REGγ and Smad7 in reciprocal co-immunoprecipitation assays (Fig. 5c, d). Indeed, Smad7 was dramatically stabilized in REGγ-deficient A549-shR cells in the presence of cycloheximide (Fig. 5e, f), an inhibitor for de novo protein synthesis. Except that, we generated stable 293T cell lines constitutively expressing a REGγ-specific KO or wide type (WT control) using TALENs. The half-life of Smad7 was longer in 293T-KO cells compared with that in 293T-WT cells (Fig. S5A–B). Furthermore, we performed gain-of-function experiments using the previously established doxycycline-inducible 293 cells system [24] to overexpress REGγ in either WT REGγ or the REGγ-N151Y (a dominant-negative mutant) cells,. Induction of REGγ promoted degradation of Smad7, while induced expression of the mutant REGγ failed to do so (Fig. S5C–D). Interestingly, a combination of MG132 and CHX resulted in dramatically stabilized Smad7 levels, suggesting Smad7 decay is dependent on proteasomal activity (Fig. S5E–F).
REGγ mediates ubiquitin-independent degradation of Smad7. a The protein expression of Smad7 in PC9 cells before and after RNAi depletion of REGγ was detected by western blotting. Smad7 was higher expressed in PC9 siR cells than PC9 siN cells at protein level. b The mRNA expression of Smad7 in a was assayed by real-time quantitative RT-PCR. n = 10; data represent means ± s.e.m.; p > 0.05, no significant correlation, as determined by Student’s t test. It showed that REGγ effected Smad7 just at protein level. c, d The interactions between REGγ and Smad7 were determined by reciprocal co-immunoprecipitation and western blot analysis following transient expression of 4 μg of HA-REGγ, 2 μg Flag-hSmad7, or 2 μg of the control Flag-vector in 293 T cells. REGγ and Smad7 can bind with each other. e A549 shN and shR cells were treated with cycloheximide (CHX, 100 μg ml−1) for indicated times followed by western blotting. CHX can inhibit protein synthesis, it showed that REGγ can degrade Smad7 in A549 shN cells. f Quantitated results in e were plotted against indicated time periods to indicate dynamic changes. Data represent means ± s.e.m.; *p < 0.05; **p < 0.01; as determined by Student’s t test. g The in vitro ubiquitin-independent degradation assay needed purified protein of REGγ-WT, REGγ-N151Y, 20S proteasome and in vitro translated protein of Smad7 were incubated at 37 °C for 3 h. An antiactin immunoblot is shown as the loading control. It showed that REGγ can degrade Smad7 independent ubiquitin in vitro. All experiments were repeated at least three times
To determine whether REGγ degrades Smad7 directly, we used cell-free proteolysis [21, 34]. Incubation of purified Smad7 with a combination of REGγ and 20S proteasome exhibited significant degradation of Smad7. However, the REGγ-N151Y-20S proteasome showed no significant difference (Fig. 5g). Taken together, our results demonstrate that REGγ can interact with Smad7 and promote its degradation in vitro and in lung cancer cells.
REGγ elicits Smad7-dependent regulation of E-cadherin
To determine whether REGγ-abrogation induced a Smad7-dependent increase in E-cadherin, we depleted Smad7 in A549-shR cells using lentivirus stably expressing Smad7-siRNA or control-siRNA [35]. Silencing Smad7 significantly reduced expression of E-cadherin and enhanced Snai1 and Slug in A549-shR cells (Fig. 6a, Fig. S6A), indicating that ablation of Smad7 successfully “compensated” loss of REGγ function. A similar conversion of EMT marker expression profile was observed in REGγ-abrogated PK cells with Smad7 RNAi (Fig. 6b). In addition, RNAi depletion of Smad7 in A549-shR cells markedly increased transwell migration and invasion compared with A549-shR cells treated with a control siRNA (Fig. 6c, e). A similar result for the Smad7 RNAi effect on cell migration/invasion was observed in PK cells with double knockdown of REGγ and Smad7 (Fig. 6d, f).
REGγ elicits Smad7-dependent regulation of E-cadherin. a, b. The expression of E-cadherin and its transcriptional repressor Snai1 and Slug were examined by western blotting before and after Smad7 knockdown in A549 shR (A549 shR siS) and RPK (RPK siS) cells. Smad7 knockdown can reverse the E-cadheirn high expression and P-Smad2/3 and Snai1 low expression by the lack of REGγ. c, d Cells treated in a–b were analyzed by transwell invasion. The migrated cells were fixed and stained with crystal violet. Scale bar, 100 μm (magnification, ×10). e, f The number of migrated cell (results from c–d) was quantified. Smad7 knockdown can reverse the reduction of cells’ migration and invation ability by the lack of REGγ. Data represent means ± s.e.m.; ***p < 0.001, as determined by Student’s t test. g A549 shN, A549 shR, and A549 shR siS cells with a luciferase reporter gene were injected into 4–6 weeks old BALB/c nude mice and representative IVIS images of tumor metastases were demonstrated. h The nude mice in g were sacrificed to count the lung cancer metastatic tumor numbers for quantification. Smad7 knockdown can reverse the decrease of lung cancer metastasis tumor number by the lack of REGγ. Data represent means ± s.e.m.; ***p < 0.001, as determined by Student’s t test
To determine the in vivo role of REGγ-Smad7 regulation in metastatic potential, A549-shN and A549-shR cells transfected with siSmad7 or control siRNA were injected into nude mice via tail veins. The luminescence images, monitored by a fluorescence microscope following injection of luciferin to the mice, demonstrated a much reduced area of tumor cell metastasis in A549-shR injected mice compared to A549-shN and increased metastasis in A549-shR-injected mice when Smad7 was silenced (Fig. 6g–h). Furthermore, we detected the Smad7 levels in tumors derived from cells silenced Smad7 and found the expression of Smad7 still decreased in the dissected tumors (Fig. S6B). In summary, we demonstrated in vitro and in vivo that Smad7 mediates REGγ-dependent regulation of EMT to promote lung cancer metastasis.
Clinical relevance of REGγ/Smad7-dependent regulation of EMT and lung cancer prognosis
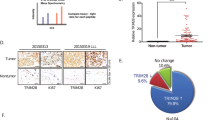
Given the importance of REGγ in EMT and cancer metastasis in mouse models, we explored clinical significance of REGγ in human lung cancers. IHC analysis of human lung cancer samples indicated that REGγ was highly expressed in tumor areas in contrast to paired nontumor tissues (Fig. 7a). Comparative analysis of normal and metastatic tissues showed a negative correlation between REGγ and Smad7 in all tumor samples inspected (Fig. 7b). Quantified IHC analysis of human tumor specimen indicated that REGγ was negatively associated with E-cadherin and Smad7 (Fig. 7c). Bioinformatics analyses using the TCGA database revealed a negative correlation between REGγ and Smad7 (Fig. 7d). Univariate Cox regression analysis disclosed an inverse correlation between REGγ level and patient survival rate, suggesting a poor prognosis associated with higher REGγ expression (Fig. S7A). Taken together, we conclude that overexpression of REGγ in lung cancers could be a marker for poor survival of human lung cancers with metastasis.
REGγ is significantly relevant to Smad7-dependent regulation of EMT and lung cancer prognosis. a Representative H&E and IHC images showing cancer areas and positive REGγ staining in human lung cancer specimen, with adjacent normal lung tissues. REGγ was higher expressed in lung tumor cells than that in normal cells. n = 9; N normal lung tissue, T tumor of lung cancer tissues. Scale bar, 25 μm (magnification, ×40). b Representative IHC images displaying an inverse correlation between REGγ, Smad7, and E-cadherin in normal and metastatic lung cancer tissues. Smad7 and E-cadherin were high expressed in normal lung tissues, but REGγ was high expressed in lung cancer lymph and bone metastasis tissuses. c Summary and statistical analysis of experiments in b with human metastatic lung cancer and normal lung tissues (n = 15), lymph nodes metastases (n = 22), and bone metastases (n = 8) samples. Scale bar, 25 μm (magnification, ×40). d The gene expression information from 1129 cases of lung cancer patients in The Cancer Genome Atlas (TCGA), reflecting the negative correlation between REGγ and Smad7 expression
Discussion
Factors contributing to lung cancer metastasis are far from being clarified. In the present study, we demonstrate the role of REGγ in lung metastasis, in addition to its known oncogenic functions by targeting various tumor suppressors [16,17,18, 31, 34, 36], in a genetically engineered conditional KrasG12D and p53 mutant mouse model. Our data provide the first in vivo evidence that REGγ promotes lung cancer metastasis by potentiating the TGFβ signal pathway. This study not only reveals a new role of REGγ in the development of advanced lung cancer, it also indicates a potentially broad impact of REGγ in the regulation of EMT-related metastasis in multiple tumors.
As a multifunctional mediator acting in regulating cellular proliferation, differentiation, apoptosis, adhesion, and migration [15], TGFβ functions is a double-edged sword for tumor suppression and tumor progression. It also acts to promote tumor invasion and metastasis by inducing EMT in advanced cancers [13, 37]. Interestingly, one of TGFβ’s antitumor functions is achieved by inhibition of REGγ transcription [38], which prevents degradation of important tumor suppressor proteins such as p21/p16. In this study, we find that one of REGγ’s tumor-promoting functions is to positively regulate TGFβ/Smad pathway and, therefore, promote EMT and cell invasion. Perhaps the action of oncogenic REGγ on TGFβ/Smad pathway at early stages of cancer development triggers counterbalance by the tumor suppressive TGFβ/Smad pathway. In advanced or metastatic lung cancer, however, REGγ exacerbates the oncogenic TGFβ/Smad signaling to promote tumor invasion and metastasis.
Among the eight different Smad proteins in vertebrates, Smad6 and Smad7 are inhibitory Smads (I-Smads) to compete with activated type I receptors and R-Smads [39]. Smad7 is ubiquitylated at Lys64 and Lys70 for its degradation and acetylation of these lysines by p300 blocks proteasome-mediated degradation of Smad7 [40]. Thus, acetylation, deacetylation, and ubiquitylation status of Smad7 determine its stability. It has been reported that TGF-β signaling inhibits the acetylation of Smad7, despite additional reports for TGF-β independent action of HDACs [41]. Like many important proteins that are subjected to degradation by both ubiquitin-dependent and ubiquitin-independent proteasome pathways, Smad7 is found in this study as a new target of the REGγ-proteasome pathway although the substrate recognition fashion remains to be elucidated. This finding addresses the molecular mechanism by which REGγ promotes EMT through empowering TGF-β pathway. Indeed, we have carried out multiple studies to substantiate that the REGγ-EMT phenomenon is Smad7-dependent. It would be interesting to investigate what happens to lung cancer metastasis in PK mice upon conditional ablation of both Smad7 and REGγ in mouse lungs.
In summary, this study identifies the REGγ as a new factor in the regulation of lung cancer metastasis. REGγ enhances TGFβ-induced EMT signaling in vitro and in vivo to promote lung cancer cell migration and invasion via degradation of Smad7 in a proteasome-dependent and ubiquitin-independent manner (Fig. 8). Combination of experimental, clinical, and bioinformatics analyses have revealed biological significance of the REGγ-Smad7 regulation in lung cancer progression and metastasis. Our study indicates that the REGγ-Smad7 pathway could serve as prognostic markers and potential therapeutic targets for metastatic lung cancers.
A proposed model for the role of REGγ in lung cancer metastasis. Elevated REGγ enhances degradation of Smad7 protein in an ubiquitin-independent manner, neutralizing the inhibitory effect of Smad7 on TGFβ receptor-I phosphorylation of Smad2/3, thus leading to Smad2/3 association with Smad4 in the nucleus and enhanced gene expression, including Snail, transcriptional repressors of E-cadherin. Finally, reduction in E-cadherin promotes the epithelial–mesenchymal transitions (EMT) and causes lung cancer metastasis
Figure S1 shows PK or RPK mouse lung tumors. Figure S2 shows wound healing and the cell migration activity results of different cells. Figure S3 shows H&E and IHC images in PK and RPK mouse lung tumors. Figure S4 shows the protein and mRNA expression of Smad7 in different cells. Figure S5 demonstrated REGγ degraded Smad7 in 293T-WT/KO and 293 doxycycline-inducible cells. Figure S6 shows the expression of EMT-related factors in A549 shN, A549 shR, A549 shR siN and A549 shR siS cells and the expression of Smad7 in nude mouse lung tumor tissues. Figure S7 shows whether the REGγ expression in TCGA is correlated with the survival of patients.
References
Wang L, Yue W, Zhang L, Zhao X, Wang Y, Xu S. mTOR and PTEN expression in non-small cell lung cancer: analysis by real-time fluorescence quantitative polymerase chain reaction and immunohistochemistry. Surg Today. 2012;42:419–25.
Maemondo M, Inoue A, Kobayashi K, Sugawara S, Oizumi S, Isobe H, et al. Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR. N. Engl J Med. 2010;362:2380–8.
Lee T, Lee B, Choi YL, Han J, Ahn MJ, Um SW. Non-small cell lung cancer with concomitant EGFR, KRAS, and ALK Mutation: clinicopathologic features of 12 cases. J Pathol Transl Med. 2016;50:197–203.
Roberts PJ, Stinchcombe TE. KRAS mutation: should we test for it, and does it matter? J Clin Oncol. 2013;31:1112–21.
Takahashi T, Nau MM, Chiba I, Birrer MJ, Rosenberg RK, Vinocour M, et al. p53: a frequent target for genetic abnormalities in lung cancer. Science. 1989;246:491–4.
Tuveson DA, Shaw AT, Willis NA, Silver DP, Jackson EL, Chang S, et al. Endogenous oncogenic K-ras(G12D) stimulates proliferation and widespread neoplastic and developmental defects. Cancer Cell. 2004;5:375–87.
Jonkers J, Meuwissen R, van der Gulden H, Peterse H, van der Valk M, Berns A. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nat Genet. 2001;29:418–25.
Ji H, Ramsey MR, Hayes DN, Fan C, McNamara K, Kozlowski P, et al. LKB1 modulates lung cancer differentiation and metastasis. Nature. 2007;448:807–10.
Meuwissen R, Berns A. Mouse models for human lung cancer. Genes Dev. 2005;19:643–64.
Otsuki Y, Saya H, Arima Y. Prospects for new lung cancer treatments that target EMT signaling. Dev Dyn. 2018;247:462–72.
Andrew DJ, Ewald AJ. Morphogenesis of epithelial tubes: Insights into tube formation, elongation, and elaboration. Dev Biol. 2010;341:34–55.
Timmerman LA, Grego-Bessa J, Raya A, Bertran E, Perez-Pomares JM, Diez J, et al. Notch promotes epithelial-mesenchymal transition during cardiac development and oncogenic transformation. Genes Dev. 2004;18:99–115.
Heldin CH, Vanlandewijck M, Moustakas A. Regulation of EMT by TGFbeta in cancer. FEBS Lett. 2012;586:1959–70.
Hu HH, Chen DQ, Wang YN, Feng YL, Cao G, Vaziri ND, et al. New insights into TGF-beta/Smad signaling in tissue fibrosis. Chem Biol Interact. 2018;292:76–83.
Hayashi H, Abdollah S, Qiu Y, Cai J, Xu YY, Grinnell BW, et al. The MAD-related protein Smad7 associates with the TGFbeta receptor and functions as an antagonist of TGFbeta signaling. Cell. 1997;89:1165–73.
Li X, Lonard DM, Jung SY, Malovannaya A, Feng Q, Qin J, et al. The SRC-3/AIB1 coactivator is degraded in a ubiquitin- and ATP-independent manner by the REGgamma proteasome. Cell. 2006;124:381–92.
Li X, Amazit L, Long W, Lonard DM, Monaco JJ, O’Malley BW. Ubiquitin- and ATP-independent proteolytic turnover of p21 by the REGgamma-proteasome pathway. Mol Cell. 2007;26:831–42.
Chen X, Barton LF, Chi Y, Clurman BE, Roberts JM. Ubiquitin-independent degradation of cell-cycle inhibitors by the REGgamma proteasome. Mol Cell. 2007;26:843–52.
Okamura T, Taniguchi S, Ohkura T, Yoshida A, Shimizu H, Sakai M, et al. Abnormally high expression of proteasome activator-gamma in thyroid neoplasm. J Clin Endocrinol Metab. 2003;88:1374–83.
Wang X, Tu S, Tan J, Tian T, Ran L, Rodier JF, et al. REG gamma: a potential marker in breast cancer and effect on cell cycle and proliferation of breast cancer cell. Med Oncol. 2011;28:31–41.
Li L, Dang Y, Zhang J, Yan W, Zhai W, Chen H, et al. REGgamma is critical for skin carcinogenesis by modulating the Wnt/beta-catenin pathway. Nat Commun. 2015;6:6875.
Wang Q, Gao X, Yu T, Yuan L, Dai J, Wang W, et al. REGgamma controls Hippo signaling and reciprocal NF-kappaB-YAP regulation to promote colon cancer. Clin Cancer Res. 2018;24:2015–25.
He J, Cui L, Zeng Y, Wang G, Zhou P, Yang Y, et al. REGgamma is associated with multiple oncogenic pathways in human cancers. BMC Cancer. 2012;12:75.
Liu J, Wang Y, Li L, Zhou L, Wei H, Zhou Q, et al. Site-specific acetylation of the proteasome activator REGgamma directs its heptameric structure and functions. J Biol Chem. 2013;288:16567–78.
Jackson EL, Willis N, Mercer K, Bronson RT, Crowley D, Montoya R, et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 2001;15:3243–8.
Liu YN, Lee WW, Wang CY, Chao TH, Chen Y, Chen JH. Regulatory mechanisms controlling human E-cadherin gene expression. Oncogene. 2005;24:8277–90.
Tang YN, Ding WQ, Guo XJ, Yuan XW, Wang DM, Song JG. Epigenetic regulation of Smad2 and Smad3 by profilin-2 promotes lung cancer growth and metastasis. Nat Commun. 2015;6:8230.
Kalluri R, Weinberg RA. The basics of epithelial-mesenchymal transition. J Clin Investig. 2009;119:1420–8.
Wendt MK, Smith JA, Schiemann WP. Transforming growth factor-beta-induced epithelial-mesenchymal transition facilitates epidermal growth factor-dependent breast cancer progression. Oncogene. 2010;29:6485–98.
Dong S, Jia C, Zhang S, Fan G, Li Y, Shan P, et al. The REGgamma proteasome regulates hepatic lipid metabolism through inhibition of autophagy. Cell Metab. 2013;18:380–91.
Xu J, Zhou L, Ji L, Chen F, Fortmann K, Zhang K, et al. The REGgamma-proteasome forms a regulatory circuit with IkappaBvarepsilon and NFkappaB in experimental colitis. Nat Commun. 2016;7:10761.
Mallini P, Lennard T, Kirby J, Meeson A. Epithelial-to-mesenchymal transition: what is the impact on breast cancer stem cells and drug resistance. Cancer Treat Rev. 2014;40:341–8.
Puisieux A, Brabletz T, Caramel J. Oncogenic roles of EMT-inducing transcription factors. Nat Cell Biol. 2014;16:488–94.
Li L, Zhao D, Wei H, Yao L, Dang Y, Amjad A, et al. REGgamma deficiency promotes premature aging via the casein kinase 1 pathway. Proc Natl Acad Sci USA. 2013;110:11005–10.
Wang J, Lu Y, Koch AE, Zhang J, Taichman RS. CXCR6 induces prostate cancer progression by the AKT/mammalian target of rapamycin signaling pathway. Cancer Res. 2008;68:10367–76.
Liu J, Yu G, Zhao Y, Zhao D, Wang Y, Wang L, et al. REGgamma modulates p53 activity by regulating its cellular localization. J Cell Sci. 2010;123:4076–84.
David CJ, Huang YH, Chen M, Su J, Zou Y, Bardeesy N, et al. TGF-beta tumor suppression through a lethal EMT. Cell. 2016;164:1015–30.
Ali A, Wang Z, Fu J, Ji L, Liu J, Li L, et al. Differential regulation of the REGgamma-proteasome pathway by p53/TGF-beta signalling and mutant p53 in cancer cells. Nat Commun. 2013;4:2667.
Miyazawa K, Miyazono K. Regulation of TGF-beta Family Signaling by Inhibitory Smads. Cold Spring Harb Perspect Biol. 2017;9:1–25.
Gronroos E, Hellman U, Heldin CH, Ericsson J. Control of Smad7 stability by competition between acetylation and ubiquitination. Mol Cell. 2002;10:483–93.
Simonsson M, Heldin CH, Ericsson J, Gronroos E. The balance between acetylation and deacetylation controls Smad7 stability. J Biol Chem. 2005;280:21797–803.
Acknowledgements
This work was supported by the National Basic Research Program of China (2016YFC0902102, 2015CB910402). This study was also funded by the National Basic Research Program (2011CB504200, 2015CB910403). This work was also supported in part by grants from National Natural Science Foundation of China (81401837, 81471066, 81672887, 81261120555, 31200878, 31071875, 81271742, 31401012, 31730017), the Science and Technology Commission of Shanghai Municipality (19140900400, 14430712100), Shanghai Rising-Star Program (16QA1401500), Shanghai natural science foundation (17ZR1407900, 12ZR1409300, 14ZR1411400) and project funded by China Postdoctoral Science Foundation(2019M651434). The authors especially thank Prof Longying Tang for her helpful comments on this manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: LL, SS, TW, LP, PZ, GC, TH, KL, QL, SX. Supervision: LT, QH, and JF. Writing—original draft: LL, XY, and XL. Writing—review and editing: LL, REM [5], and XL.
Corresponding authors
Ethics declarations
Ethics animal experimentation
Animal experiments were approved by the East China Normal University of Institutional Animal Care and Use Committee.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Piacentini
Supplementary information
Rights and permissions
About this article
Cite this article
Tong, L., Shen, S., Huang, Q. et al. Proteasome-dependent degradation of Smad7 is critical for lung cancer metastasis. Cell Death Differ 27, 1795–1806 (2020). https://doi.org/10.1038/s41418-019-0459-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0459-6
This article is cited by
-
Clinically relevant stratification of lung squamous carcinoma patients based on ubiquitinated proteasome genes for 3P medical approach
EPMA Journal (2024)
-
PHF14 enhances DNA methylation of SMAD7 gene to promote TGF-β-driven lung adenocarcinoma metastasis
Cell Discovery (2023)
-
Role of mitochondrial stress and the NLRP3 inflammasome in lung diseases
Inflammation Research (2023)
-
Identification and characterization of colorectal-cancer-associated SNPs on the SMAD7 locus
Journal of Cancer Research and Clinical Oncology (2023)
-
Loss of deubiquitylase USP2 triggers development of glioblastoma via TGF-β signaling
Oncogene (2022)