Abstract
Purpose
Retinitis pigmentosa (RP) is a genetically heterogeneous inherited retinal dystrophy. To date, over 80 genes have been implicated in RP. However, the disease demonstrates significant locus and allelic heterogeneity not entirely captured by current testing platforms. The purpose of the present study was to characterize the underlying mutation in a patient with RP without a molecular diagnosis after initial genetic testing.
Methods
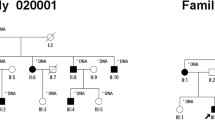
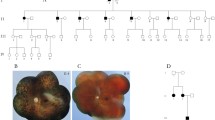
Whole-exome sequencing of the affected proband was performed. Candidate gene mutations were selected based on adherence to expected genetic inheritance pattern and predicted pathogenicity. Sanger sequencing of MERTK was completed on the patient’s unaffected mother, affected brother, and unaffected sister to determine genetic phase.
Results
Eight sequence variants were identified in the proband in known RP-associated genes. Sequence analysis revealed that the proband was a compound heterozygote with two independent mutations in MERTK, a novel nonsense mutation (c.2179C > T) and a previously reported missense variant (c.2530C > T). The proband’s affected brother also had both mutations. Predicted phase was confirmed in unaffected family members.
Conclusion
Our study identifies a novel nonsense mutation in MERTK in a family with RP and no prior molecular diagnosis. The present study also demonstrates the clinical value of exome sequencing in determining the genetic basis of Mendelian diseases when standard genetic testing is unsuccessful.




Similar content being viewed by others
References
Evans RJ, Schwarz N, Nagel-Wolfrum K, Wolfrum U, Hardcastle AJ, Cheetham ME (2010) The retinitis pigmentosa protein RP2 links pericentriolar vesicle transport between the Golgi and the primary cilium. Hum Mol Gen 19:1358–1367. doi:10.1093/hmg/ddq012
Busskamp V, Duebel J, Balya D et al (2010) Genetic reactivation of cone photoreceptors restores visual responses in retinitis pigmentosa. Science 329:413–417. doi:10.1126/science.1190897
Hartong DT, Berson EL, Dryja TP (2006) Retinitis pigmentosa. Lancet 368:1795–1809. doi:10.1016/S0140-6736(06)69740-7
Farrar G, Kenna PF, Humphries P (2002) New EMBO member’s review: on the genetics of retinitis pigmentosa and on mutation-independent approaches to therapeutic intervention. EMBO J 21:857–864. doi:10.1093/emboj/21.5.857
Ferrari S, Di Iorio E, Barbaro V, Ponzin D, Sorrentino FS, Parmeggiani F (2011) Retinitis pigmentosa: genes and disease mechanisms. Curr Genomics 12:238–249. doi:10.2174/138920211795860107
Fernandez-San Jose P, Blanco-Kelly F, Corton M, Trujillo-Tiebas M-J, Gimenez A, Avila-Fernandez A, Garcia-Sandoval B, Lopez-Molina M-I, Hernan I, Carballo M, Riveiro-Alvarez R, Ayuso C (2015) Prevalence of rhodopsin mutations in autosomal dominant retinitis pigmentosa in spain: clinical and analytical review in 200 families. Acta Ophthalmol 93:e38–e44. doi:10.1111/aos.12486
Daiger SP, Sullivan LS, Bowne SJ (2013) Genes and mutations causing retinitis pigmentosa. Clin Genet 84 10.1111/cge.12203
Wang NH-H, Chen S-J, Yang C-F, Chen H-W, Chuang H-P, Lu Y-H, Chen C-H, Wu J-Y, Niu D-M, Chen Y-T (2016) Homozygosity mapping and whole-genome sequencing links a missense mutation in POMGNT1 to autosomal recessive retinitis pigmentosa missense mutation in POMGNT1 and autosomal recessive RP. Invest Ophthalmol Vis Sci 57:3601–3609. doi:10.1167/iovs.16-19463
Anasagasti A, Irigoyen C, Barandika O, López de Munain A, Ruiz-Ederra J (2012) Current mutation discovery approaches in retinitis Pigmentosa. Vis Res 75:117–129. doi:10.1016/j.visres.2012.09.012
Boye SE, Boye SL, Lewin AS, Hauswirth WW (2013) A comprehensive review of retinal gene therapy. Mol Ther 21:509–519. doi:10.1038/mt.2012.280
Tsou W-I, Nguyen K-QN, Calarese DA, Garforth SJ, Antes AL, Smirnov SV, Almo SC, Birge RB, Kotenko SV (2014) Receptor tyrosine kinases, TYRO3, AXL, and MER, demonstrate distinct patterns and complex regulation of ligand-induced activation. J Biol Chem 289:25750–25763. doi:10.1074/jbc.M114.569020
Zagorska A, Traves PG, Lew ED, Dransfield I, Lemke G (2014) Diversification of TAM receptor tyrosine kinase function. Nat Immunol 15:920–928. doi:10.1038/ni.2986
Chen C, Li Q, Darrow AL, Wang Y, Derian CK, Yang J, de Garavilla L, Andrade-Gordon P, Damiano BP (2004) Mer receptor tyrosine kinase signaling participates in platelet function. Arterioscler Thromb Vasc Biol 24:1118–1123. doi:10.1161/01.atv.0000130662.30537.08
Linger RMA, Keating AK, Earp HS, Graham DK (2008) TAM receptor tyrosine kinases: biologic functions, signaling, and potential therapeutic targeting in human cancer. Adv Cancer Res 100:35–83. doi:10.1016/S0065-230X(08)00002-X
Parinot C, Nandrot FE (2016) A comprehensive review of mutations in the MERTK proto-oncogene. In: Bowes Rickman C, LaVail MM, Anderson ER, Grimm C, Hollyfield J, Ash J (eds) Retinal degenerative diseases: mechanisms and experimental therapy. Springer International Publishing, Cham, pp 259–265
Nandrot EF, Kim Y, Brodie SE, Huang X, Sheppard D, Finnemann SC (2004) Loss of synchronized retinal phagocytosis and age-related blindness in mice lacking αvβ5 integrin. J Exp Med 200:1539–1545. doi:10.1084/jem.20041447
Nandrot EF, Anand M, Almeida D, Atabai K, Sheppard D, Finnemann SC (2007) Essential role for MFG-E8 as ligand for αvβ5 integrin in diurnal retinal phagocytosis. Proc Natl Acad Scie USA 104:12005–12010. doi:10.1073/pnas.0704756104
Qin S (2016) Blockade of MerTK activation by AMPK inhibits RPE cell phagocytosis. In: Bowes Rickman C, LaVail MM, Anderson ER, Grimm C, Hollyfield J, Ash J (eds) Retinal degenerative diseases: mechanisms and experimental therapy. Springer International Publishing, Cham, pp 773–778
International Human Genome Sequencing Consortium (2001) Initial sequencing and analysis of the human genome. Nature 409:860–921. doi:10.1038/35057062
Carnevali P, Baccash J, Halpern AL, Nazarenko I, Nilsen GB, Pant KP, Ebert JC, Brownley A, Morenzoni M, Karpinchyk V, Martin B, Ballinger DG, Drmanac R (2012) Computational techniques for human genome resequencing using mated gapped reads. J Comput Biol 19:279–292. doi:10.1089/cmb.2011.0201
Drmanac R, Sparks AB, Callow MJ et al (2010) Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 327:78–81. doi:10.1126/science.1181498
McHenry CL, Liu Y, Feng W, Nair AR, Feathers KL, Ding X, Gal A, Vollrath D, Sieving PA, Thompson DA (2004) MERTK arginine-844-cysteine in a patient with severe rod–cone dystrophy: loss of mutant protein function in transfected cells. Invest Ophthalmol Vis Sci 45:1456–1463. doi:10.1167/iovs.03-0909
Bourne MC, Campbell DA, Tansley K (1938) Hereditary degeneration of the rat retina. Br J Ophthalmol 22:613–623. doi:10.1136/bjo.22.10.613
D’Cruz PM, Yasumura D, Weir J, Matthes MT, Abderrahim H, LaVail MM, Vollrath D (2000) Mutation of the receptor tyrosine kinase gene Mertk in the retinal dystrophic RCS rat. Hum Mol Gen 9:645–651. doi:10.1093/hmg/9.4.645
Nandrot E, Dufour EM, Provost AC, Péquignot MO, Bonnel S, Kn G, Marchant D, Rouillac C, Sépulchre de Condé B, Bihoreau M-T, Shaver C, Dufier J-L, Marsac C, Lathrop M, Menasche M, Abitbol MM (2000) Homozygous deletion in the coding sequence of the C-MER gene in RCS rats unravels general mechanisms of physiological cell adhesion and apoptosis. Neurobiol Dis 7:586–599. doi:10.1006/nbdi.2000.0328
Bok D, Hall MO (1971) The role of the pigment epithelium in the etiology of inherited retinal dystrophy in the rat. J Cell Biol 49:664–682. doi:10.1083/jcb.49.3.664
Finnemann SC, Nandrot EF (2006) MERTK activation during rpe phagocytosis in vivo requires αvβ5 integrin. Adv Exp Med Biol 572:499–503. doi:10.1007/0-387-32442-9_69
Dowling JE, Sidman RL (1962) Inherited retinal dystrophy in the rat. J Cell Biol 14:73–109. doi:10.1083/jcb.14.1.73
Weisschuh N, Mayer AK, Strom TM et al (2016) Mutation detection in patients with retinal dystrophies using targeted next generation sequencing. PLoS One 11:e0145951. doi:10.1371/journal.pone.0145951
Acknowledgements
The authors extend our deep gratitude to the proband and his family. We also thank Vaiva Liakaite at the University of Illinois at Chicago’s DNA Services Facility and the sequencing center at BGI for assisting with the present study. Lastly, the authors thank Darshana Patel for assisting with the coordination of this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This report was supported by funding from Search for Vision, R01EY023644, NEI core grant EY001792, and Research to Prevent Blindness (departmental support). The sponsor had no role in the design or conduct of this research.
Conflict of interest
All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest (such as honoraria; educational grants; participation in speakers’ bureaus; membership, employment, consultancies, stock ownership, or other equity interest; and expert testimony or patent-licensing arrangements), or non-financial interest (such as personal or professional relationships, affiliations, knowledge or beliefs) in the subject matter or materials discussed in this manuscript.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Al-khersan, H., Shah, K.P., Jung, S.C. et al. A novel MERTK mutation causing retinitis pigmentosa. Graefes Arch Clin Exp Ophthalmol 255, 1613–1619 (2017). https://doi.org/10.1007/s00417-017-3679-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00417-017-3679-9