Abstract
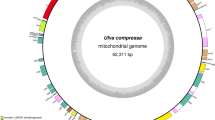
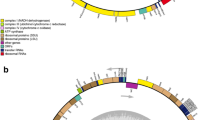
Turbinaria ornata (Turner) J. Agardh is a perennial brown alga native to coral reef ecosystems of tropical areas of the Pacific and Indian Ocean. Very little is known about its organellar genome structure. In the present work, the complete mitochondrial genome sequence of T. ornata was determined and compared with other reported brown algal mtDNAs. The circular mitogenome of 34,981 bp contains a basic set of 65 mitochondrial genes. The structure and organization of T. ornata mitogenome is very similar to Sargassum species. Turbinaria ornata genes overlap by a total of 164 bp in 12 different locations from 1 to 66 bp, and the non-coding sequences are 1872 bp, constituting approximate 5.35 % of the genome. The total spacer size has positive correlation with the brown algal mitogenome size with the correlation coefficient of 0.7972. Several regions displaying greater inconsistency (rnl-trnK spacer, cox2 gene, cox3-atp6 spacer, rps14-rns middle region and trnP-rnl spacer) have been identified in brown algal mtDNAs. The observed uncertainty regarding the position and support values of some branches might be closely associated with the heterogeneity of evolutionary rate.





Similar content being viewed by others
References
Alikhan N, Petty NK, Zakour NLB, Beatson SA (2011) BLAST Ring Image Generator (BRIG): simple prokaryote genome comparisons. BMC Genom 12:402
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Bittick SJ, Bilotti ND, Peterson HA, Stewart HL (2010) Turbinaria ornata as an herbivory refuge for associate algae. Mar Biol 157:317–323
Charrier B, Bail AL, de Reviers B (2012) Plant Proteus: brown algal morphological plasticity and underlying developmental mechanisms. Trends Plant Sci 17(8):468–477
Draisma SGA, Ballesteros E, Rousseau F, Thibaut T (2010) DNA sequence data demonstrate the polyphyly of the genus Cystoseira and other Sargassaceae genera (Phaeophyceae). J Phycol 46:1329–1345
Gómez-Valero L, Latorre A, Silva FJ (2004) The evolutionary fate of nonfunctional DNA in the bacterial endosymbiont Buchnera aphidicola. Mol Biol Evol 21(11):2172–2181
Guiry MD, Guiry GM (2014) AlgaeBase. World-wide electronic publication, National University of Ireland, Galway. http://www.algaebase.org. Accessed on 18 December 2014
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acid Symp Ser 41:95–98
Kimura M (1980) A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kogame K, Uwai S, Shimada S, Masuda M (2005) A study of sexual and asexual populations of Scytosiphon lomentaria (Scytosiphonaceae, Phaeophyceae) in Hokkaido, northern Japan, using molecular marker. Eur J Phycol 40:313–322
Le Lann K, Kervarec N, Payri C, Deslandes E, Stiger Pouvreau V (2008) Discrimination of allied species within the genus Turbinaria (Fucales, Phaeophyceae) using HRMAS NMR spectroscopy. Talanta 74:1079–1083
Li TY, Qu JQ, Feng YJ, Liu C, Chi S, Liu T (2014) Complete mitochondrial genome of Undaria pinnatifida (Alariaceae, Laminariales, Phaeophyceae). Mitochondrial DNA. doi:10.3109/19401736.2013.865172
Liu F, Pang SJ (2014a) Complete mitochondrial genome of the brown alga Scytosiphon lomentaria (Scytosiphonaceae, Phaeophyceae). Mitochondrial DNA. doi:10.3109/19401736.2014.953108
Liu F, Pang SJ (2014b) Complete mitochondrial genome of the invasive brown alga Sargassum muticum (Sargassaceae, Phaeophyceae). Mitochondrial DNA. doi:10.3109/19401736.2014.933333
Liu F, Pang SJ (2014c) Mitochondrial phylogenomics reveals a close relationship between Petalonia fascia (Scytosiphonaceae, Phaeophyceae) and Ectocarpus siliculosus. J Appl Phycol. doi:10.1007/s10811-014-0386-3
Liu F, Pang SJ (2015) Mitochondrial genome of Sargassum thunbergii: conservation and variability of mitogenomes within the subgenus Bactrophycus. Mitochondrial DNA. doi:10.3109/19401736.2015.1007328
Liu F, Pang SJ, Chen WZ (2014a) Complete mitochondrial genome of the brown alga Sargassum hemiphyllum (Sargassaceae, Phaeophyceae): comparative analyses. Mitochondrial DNA. doi:10.3109/19401736.2014.953096
Liu F, Pang SJ, Li J, Li X (2014b) Complete mitochondrial genome of the brown alga Colpomenia peregrina (Scytosiphonaceae, Phaeophyceae): genome characterization and comparative analyses. Mitochondrial DNA. doi:10.3109/19401736.2014.958688
Liu F, Pang SJ, Luo MB (2014c) Complete mitochondrial genome of the brown alga Sargassum fusiforme (Sargassaceae, Phaeophyceae): genome architecture and taxonomic consideration. Mitochondrial DNA. doi:10.3109/19401736.2014.936417
Liu F, Pang SJ, Li X, Li J (2015) Complete mitochondrial genome of the brown alga Sargassum horneri (Sargassaceae, Phaeophyceae): genome organization and phylogenetic analyses. J Appl Phycol 27:469–478
Lohse M, Drechsel O, Kahlau S, Bock R (2013) OrganellarGenomeDRAW—a suite of tools for generating physical maps of plastid and mitochondrial genomes and visualizing expression data sets. Nucl Acids Res. doi:10.1093/nar/gkt289
Oudot-Le Secq MP, Fontaine JM, Rousvoal S, Kloareg B, Loiseaux-De Goër S (2001) The complete sequence of a brown algal mitochondrial genome, the Ectocarpale Pylaiella littoralis (L.) Kjellm. J Mol Evol 53:80–88
Oudot-Le Secq MP, Kloareg B, Loiseaux-De Goër S (2002) The mitochondrial genome of the brown alga Laminaria digitata: a comparative analysis. Eur J Phycol 37:163–172
Oudot-Le Secq MP, Loiseaux-De Goër S, Stam WT, Olsen JL (2006) Complete mitochondrial genome of the three brown algae (Heterokonta: Phaeophyceae) Dictyota dichotoma, Fucus vesiculosus and Desmarestia viridis. Curr Genet 49:47–58
Phillips N, Kapraun DF, Garreta AG, Siguan MAR, Rull JL, Soler NS, Lewis R, Kawai H (2011) Estimates of nuclear DNA content in 98 species of brown algae (Phaeophyta). AoB Plants. doi:10.1093/aobpla/plr001
Prathep A, Wichachucherd B, Thongroy P (2007) Spatial and temporal variation in density and thallus morphology of Turbinaria ornata in Thailand. Aquat Bot 86:132–138
Qu JQ, Liu C, Wang XM, Zhang ZB, Chi S, Liu T (2014) Complete mitochondrial genome of Costaria costata shows conservative evolution in Laminariales. Mitochondrial DNA. doi:10.3109/19401736.2013.863290
Ravin NV, Galachyants YP, Mardanov AV, Beletsky AV, Petrova DP, Sherbakova TA, Zakharova YR, Likhoshway YV, Skryabin KG, Grachev MA (2010) Complete sequence of the mitochondrial genome of a diatom alga Synedra acus and comparative analysis of diatom mitochondrial genomes. Curr Genet 56:215–223
Rohfritsch A, Payri C, Stiger V, Bonhomme F (2007) Molecular and morphological relationships between two closely related species, Turbinaria ornata and T. conoides (Sargassaceae, Phaeophyceae). Biochem Syst Ecol 35:91–98
Rohfritsch A, Payri C, Stiger V, Bonhomme F (2010) Habitat-related allelic variation revealed by an anonymous DNA locus in reef-dwelling Turbinaria ornata (Fucales, Phaeophyceae). Bot Mar 53:189–192
Rousseau F, Burrowes R, Peters AF, Kuhlenkamp R, de Reviers B (2001) A comprehensive phylogeny of the Phaeophyceae based on nrDNA sequences resolves the earliest divergences. Comptes Rendus de l’Academie des Sciences. Serie III. Sciences de la Vie (Paris) 324:305–319
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:686–689
Silberfeld T, Rousseau F, de Reviers B (2014) An updated classification of brown algae (Ochrophyta, Phaeophyceae). Cryptogamie Algologie 35(2):117–156
Stewart HL (2004) Hydrodynamic consequences of maintaining an upright posture by different magnitudes of stiffness and buoyancy in the tropical alga Turbinaria ornata. J Mar Syst 49:157–167
Stewart HL (2006) Ontogenetic changes in buoyancy, breaking strength, extensibility, and reproductive investment in a drifting macroalga Turbinaria ornata (Phaeophyta). J Phycol 42:43–50
Stiger V, Payri CE (2005) Natural settlement dynamics of a young population of Turbinaria ornata and phonological comparisons with older populations. Aquat Bot 81:225–243
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The ClustalX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882
Whelan S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol 18:691–699
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20(17):3252–3255
Wynne MJ (2002) Turbinaria foliosa sp. nov. (Fucales, Phaeophyceae) from the Sultanate of Oman, with a census of currently recognized species in the genus Turbinaria. Phycol Res 50:283–293
Yotsukura N, Shimizu T, Katayama T, Druehl LD (2010) Mitochondrial DNA sequence variation of four Saccharina species (Laminariales, Phaeophyceae) growing in Japan. J Appl Phycol 22:243–251
Zhang J, Wang XM, Liu C, Jin YM, Liu T (2013) The complete mitochondrial genomes of two brown algae (Laminariales, Phaeophyceae) and phylogenetic analysis within Laminaria. J Appl Phycol 25:1247–1253
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 41206146, 41176135), the Scientific Research Foundation for Outstanding Young Scientists of Shandong Province (No. BS2013HZ004), the Special Fund of the membership of Youth Innovation Promotion Association of Chinese Academy of Sciences (No. 2015164), the 863 Hi-Tech Research and Development Program of China (No. 2012AA10A413), the Science and Technology Service Network Program of Chinese Academy of Sciences (No. KFJ-EW-STS-060), and the Knowledge Innovation Programs of Chinese Academy of Sciences (No. KSCX2-EW-B-14).
Compliance with Ethical standards
The authors declare that they have no conflict of interest. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. This article does not contain any studies with human participants or animals performed by any of the authors. Informed consent was obtained from all individual participants included in the study.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Liu, F., Pang, S. Mitochondrial genome of Turbinaria ornata (Sargassaceae, Phaeophyceae): comparative mitogenomics of brown algae. Curr Genet 61, 621–631 (2015). https://doi.org/10.1007/s00294-015-0488-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-015-0488-8