Abstract
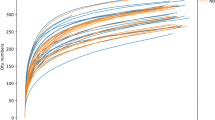
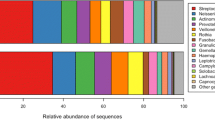
Severe early childhood caries are a prevalent public health problem among preschool children throughout the world. However, little is known about the microbiota found in association with severe early childhood caries. Our study aimed to explore the bacterial microbiota of dental plaques to study the etiology of severe early childhood caries through pyrosequencing analysis based on 16S rRNA gene V1–V3 hypervariable regions. Forty participants were enrolled in the study, and we obtained twenty samples of supragingival plaque from caries-free subjects and twenty samples from subjects with severe early childhood caries. A total of 175,918 reads met the quality control standards, and the bacteria found belonged to fourteen phyla and sixty-three genera. Our results show the overall structure and microbial composition of oral bacterial communities, and they suggest that these bacteria may present a core microbiome in the dental plaque microbiota. Three genera, Streptococcus, Granulicatella, and Actinomyces, were increased significantly in children with severe dental cavities. These data may facilitate improvements in the prevention and treatment of severe early childhood caries.



Similar content being viewed by others
References
Aas JA, Griffen AL, Dardis SR, Lee AM, Olsen I, Dewhirst FE, Leys EJ, Paster BJ (2008) Bacteria of dental caries in primary and permanent teeth in children and young adults. J Clin Microbiol 46:1407–1417
Amarante E, Raadal M, Espelid I (1998) Impact of diagnostic criteria on the prevalence of dental caries in Norwegian children aged 5, 12 and 18 years. Community Dent Oral Epidemiol 26:87–94
Arora A, Scott J, Bhole S, Do L, Schwarz E, Blinkhorn A (2011) Early childhood feeding practices and dental caries in preschool children: a multi-centre birth cohort study. BMC Public Health 11:28
Caporaso JG, Lauber CL, Costello EK, Berg-Lyons D, Gonzalez A, Stombaugh J, Knights D, Gajer P, Ravel J, Fierer N (2011) Moving pictures of the human microbiome. Genome Biol 12:R50
Chao A, Bunge J (2004) Estimating the number of species in a stochastic abundance model. Biometrics 58:531–539
Chao A, Lee S, Jeng S (1992) Estimating population size for capture-recapture data when capture probabilities vary by time and individual animal. Biometrics 48(1):201–216
Cisar J, Kolenbrander P, McIntire F (1979) Specificity of coaggregation reactions between human oral streptococci and strains of Actinomyces viscosus or Actinomyces naeslundii. Infect Immun 24:742–752
Gross EL, Beall CJ, Kutsch SR, Firestone ND, Leys EJ, Griffen AL (2012) Beyond Streptococcus mutans: dental caries onset linked to multiple species by 16S rRNA community analysis. PLoS One 7:e47722
Hamady M, Walker JJ, Harris JK, Gold NJ, Knight R (2008) Error-correcting barcoded primers for pyrosequencing hundreds of samples in multiplex. Nat Methods 5:235–237
Hogg SD (1992) Lactic acid bacteria: the lactic acid bacteria in health and disease. 1:115–132
Hong-Ying W, Petersen PE, Jin-You B, Bo-Xue Z (2011) The second national survey of oral health status of children and adults in China. Int Dent J 52:283–290
Huse SM, Ye Y, Zhou Y, Fodor AA (2012) A core human microbiome as viewed through 16S rRNA sequence clusters. PLoS One 7:e34242
Kanasi E, Dewhirst F, Chalmers N, Kent R Jr, Moore A, Hughes C, Pradhan N, Loo C, Tanner A (2010) Clonal analysis of the microbiota of severe early childhood caries. Caries Res 44:485–497
Keijser BJ, Zaura E, Huse SM, van der Vossen JM, Schuren FH, Montijn RC, ten Cate JM, Crielaard W (2008) Pyrosequencing analysis of the oral microflora of healthy adults. J Dent Res 87:1016–1020
Lazarevic V, Whiteson K, Hernandez D, Francois P, Schrenzel J (2010) Study of inter- and intra-individual variations in the salivary microbiota. BMC Genomics 11:523
Lazarevic V, Whiteson K, Huse S, Hernandez D, Farinelli L, Osteras M, Schrenzel J, Francois P (2009) Metagenomic study of the oral microbiota by Illumina high-throughput sequencing. J Microbiol Methods 79:266–271
Ling Z, Kong J, Jia P, Wei C, Wang Y, Pan Z, Huang W, Li L, Chen H, Xiang C (2010) Analysis of oral microbiota in children with dental caries by PCR-DGGE and barcoded pyrosequencing. Microb Ecol 60:677–690
Marsh PD (2010) Microbiology of dental plaque biofilms and their role in oral health and caries. Dent Clin North Am 54:441
Messer LB (2000) Assessing caries risk in children. Aust Dent J 45:10–16
Peterson SN, Snesrud E, Schork NJ, Bretz WA (2011) Dental caries pathogenicity: a genomic and metagenomic perspective. Int Dent J 61(Suppl 1):11–22
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J, Glockner FO (2007) SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35:7188–7196
Quevedo B, Giertsen E, Zijnge V, Lüthi-Schaller H, Guggenheim B, Thurnheer T, Gmür R (2011) Phylogenetic group-and species-specific oligonucleotide probes for single-cell detection of lactic acid bacteria in oral biofilms. BMC Microbiol 11:14
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ (2009) Introducing Mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Shen S, Samaranayake LP, Yip HK (2005) Coaggregation profiles of the microflora from root surface caries lesions. Arch Oral Biol 50:23–32
Siqueira JF Jr., Fouad AF, Rocas IN (2012) Pyrosequencing as a tool for better understanding of human microbiomes. J Oral Microbiol 4. doi:10.3402/jom.v4i0.10743
Zaura E, Keijser BJF, Huse SM, Crielaard W (2009) Defining the healthy “core microbiome” of oral microbial communities. BMC Microbiol 9:259. doi:10.1186/1471-2180-9-259
Acknowledgments
We acknowledge Dr. Jinhua FANG and colleagues from Affiliated Hospital of Stomatology, Zhejiang University for their clinical assistances. The study was founded by grants 2011 China State key clinical department grants.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jiang, W., Zhang, J. & Chen, H. Pyrosequencing Analysis of Oral Microbiota in Children with Severe Early Childhood Dental Caries. Curr Microbiol 67, 537–542 (2013). https://doi.org/10.1007/s00284-013-0393-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-013-0393-7