Abstract
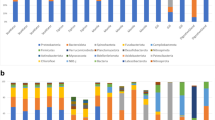
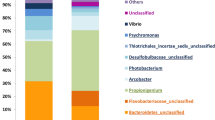
Larvae of the biofouling tubeworm Hydroides elegans (Polychaeta) must contact a bacterial biofilm to induce settlement. To further understand the relationship between particular bacteria and the worm, this study analyzed associated microbiomes at multiple life-history stages with high-throughput 16S rRNA amplicon sequencing. Worms were collected from Pearl Harbor Hawai’i (21°21′27.2″N 157°57′34.8″W), and microbial communities were analyzed from adults, their tubes, spawned eggs, 6-day old nectochaete larvae, and filtered seawater from the larval cultures that included cells of the microalga Isochrysis galbana fed to the larvae. Microbiomes of females sampled at two different times were significantly different: Cellvibrionales dominated the microbiome of adult females in August 2016, whereas Oceanospirillales were dominant in females collected in April 2017. Additionally, in 2017, the microbiomes across all life stages were significantly different from each other; only one OTU in the genus Endozoicomonas was shared between females and their nectochaetes at a relative abundance > 1%. Because of the apparent absence of a consistent microbiome between stages, female worms, their eggs and both trochophore and nectochaete larvae were stained to reveal the presence of bacteria and analyzed with scanning confocal microscopy. The eggs were almost devoid of bacteria, and bacteria seen on larvae were almost all confined to the feeding apparatus. The evidence of inconsistent microbiomes across life stages and lack of bacteria on eggs demonstrates that H. elegans lacks a vertically transmitted microbiome, and bacteria found at each stage came from a varying environment.





Similar content being viewed by others
References
Ainsworth TD, Krause L, Bridge T, Torda G, Raina JB, Zakrzewski M, Gates RD, Padilla-Gamiño JL, Spalding HL, Smith C, Woolsey ES (2015) The coral core microbiome identifies rare bacterial taxa as ubiquitous endosymbionts. ISME J 9:2261. https://doi.org/10.1038/ismej.2015.39
Apprill A, Marlow HQ, Martindale MQ, Rappé MS (2009) The onset of microbial associations in the coral Pocillopora meandrina. ISME J 3:685–699. https://doi.org/10.1038/ismej.2009.3
Bordenstein SR, Theis KR (2015) Host biology in light of the microbiome: ten principles of holobionts and hologenomes. PLoS Biol 13:e1002226. https://doi.org/10.1371/journal.pbio.1002226
Bright M, Bulgheresi S (2010) A complex journey: transmission of microbial symbionts. Nat Rev Microbiol 8:218–230. https://doi.org/10.1038/nrmicro2262
Carpizo-Ituarte E, Hadfield M (1998) Stimulation of metamorphosis in the polychaete Hydroides elegans Haswell (Serpulidae). Biol Bull 194:14–24. https://doi.org/10.2307/1542509
Donaldson GP, Lee SM, Mazmanian SK (2015) Gut biogeography of the bacterial microbiota. Nat Rev Microbiol 14:20–32. https://doi.org/10.1038/nrmicro3552
Dubilier N, Bergin C, Lott C (2008) Symbiotic diversity in marine animals: the art of harnessing chemosynthesis. Nat Rev Microbiol 6:725–740. https://doi.org/10.1038/nrmicro1992
Dufour SC, Laurich JR, Batstone RT, McCuaig B, Elliott A (2014) Magnetosome-containing bacteria living as symbionts of bivalves. ISME J 8:2453–2462. https://doi.org/10.1038/ismej.2014.93
Eberl G (2018) The microbiota, a necessary element of immunity. C R Biol 341:281–283. https://doi.org/10.1016/j.crvi.2018.03.003
Edgar R (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Fieth RA, Gauthier MA, Bayes J, Green KM, Degnan SM (2016) Ontogenetic changes in the bacterial symbiont community of the tropical demosponge Amphimedon queenslandica: metamorphosis is a new beginning. Front Mar Sci 15:228. https://doi.org/10.3389/fmars.2016.00228
Freckelton ML, Nedved BT, Hadfield MG (2017) Induction of invertebrate larval settlement; different bacteria, different mechanisms? Sci Rep 7:42557. https://doi.org/10.1038/srep42557
Goffredi SK, Johnson SB, Vrijenhoek RC (2007) Genetic diversity and potential function of microbial symbionts associated with newly discovered species of Osedax polychaete worms. Appl Environ Microbiol 73:2314–2323. https://doi.org/10.1128/AEM.01986-06
Hadfield MG (2011) Biofilms and marine invertebrate larvae: what bacteria produce that larvae use to choose settlement sites. Annu Rev Mar Sci 3:453–470. https://doi.org/10.1146/annurev-marine-120709-142753
Hadfield MG, Unabia C, Smith CM, Michael TM (1994) Settlement preferences of the ubiquitous fouler Hydroides elegans. In: M. Fingerman, R. Nagabhushanam, R. Sarojini (eds) Recent developments in biofouling control. Oxford & IBH, New Delhi, pp. 65-74
Hadfield MG, Unabia C, Smith CM, Michael TM (1994b) Settlement preferences of the ubiquitous fouler Hydroides elegans. In: Fingerman M, Nagabhushanam R, Sarojini R (eds) Recent developments in biofouling control. Oxford & IBH, New Delhi, pp 65–74
Hadfield MG, Nedved BT, Wilbur S, Koehl MAR (2014) Biofilm cue for larval settlement in Hydroides elegans (Polychaeta): is contact necessary? Mar Biol 161:2577–2587. https://doi.org/10.1007/s00227-014-2529-0
Hardoim CC, Costa R (2014) Temporal dynamics of prokaryotic communities in the marine sponge Sarcotragus spinosulus. Mol Ecol 23:3097–3112. https://doi.org/10.1111/mec.12789
Hentschel U, Piel J, Degnan SM, Taylor MW (2012) Genomic insights into the marine sponge microbiome. Nat Rev Microbiol 10:641–654. https://doi.org/10.1038/nrmicro2839
Hernandez-Agreda A, Gates RD, Ainsworth TD (2017) Defining the core microbiome in corals’ microbial soup. Trends Microbiol 25:125–140. https://doi.org/10.1016/j.tim.2016.11.003
Hooper LV, Littman DR, Macpherson AJ (2012) Interactions between the microbiota and the immune system. Science 336:1268–1273. https://doi.org/10.1126/science.1223490
Huang S, Hadfield MG (2003) Composition and density of bacterial biofilms affect metamorphosis of the polychaete Hydroides elegans. Mar Ecol Prog Ser 260:161–172. https://doi.org/10.3354/meps260161
Huggett MJ, Nedved BT, Hadfield MG (2009) Effects of initial surface wettability on biofilm formation and subsequent settlement of Hydroides elegans. Biofouling 25:387–399
Jones SE, Lennon JT (2010) Dormancy contributes to the maintenance of microbial diversity. Proc Nat Acad Sci 107:5881–5886. https://doi.org/10.1073/pnas.0912765107
Kimes NE, Johnson WR, Torralba M, Nelson KE, Weil E, Morris PJ (2013) The Montastraea faveolata microbiome: ecological and temporal influences on a Caribbean reef-building coral in decline. Environ Microb 15:2082–2094. https://doi.org/10.1111/1462-2920.12130
Klose J, Aistleitner K, Horn M, Krenn L, Dirsch V, Zehl M, Bright M (2016) Trophosome of the deep-sea tubeworm Riftia pachyptila inhibits bacterial growth. PLoS ONE 11:e0146446. https://doi.org/10.1371/journal.pone.0146446
Kuczynski J, Stombaugh J, Walters WA, Gonzalez A, Caporaso G, Knight R (2012) Using QIIME to analyze 16S rRNA gene sequences from microbial communities. Curr Protoc Microbiol 27:1E–5. https://doi.org/10.1002/9780471729259.mc01e05s27
Lau SC, Mak KK, Chen F, Qian PY (2002) Bioactivity of bacterial strains isolated from marine biofilms in Hong Kong waters for the induction of larval settlement in the marine polychaete Hydroides elegans. Mar Ecol Prog Ser 226:301–310. https://doi.org/10.3354/meps226301
Lee KWK, Periasamy S, Mukherjee M, Xie C, Kjelleberg S, Rice SA (2014) Biofilm development and enhanced stress resistance of a model, mixed-species community biofilm. ISME J 8:894–907. https://doi.org/10.1038/ismej.2013.194
Lema KA, Bourne DG, Willis BL (2014) Onset and establishment of diazotrophs and other bacterial associates in the early life history stages of the coral Acropora millepora. Mol Ecol 23:4682–4695. https://doi.org/10.1111/mec.12899
Lema KA, Constancias F, Rice SA, Hadfield MG (2019) High bacterial diversity in near-shore and oceanic biofilms and their influence on larval settlement by Hydroides elegans (Polychaeta) (submitted)
Li M, Yang H, Gu JD (2009) Phylogenetic diversity and axial distribution of microbes in the intestinal tract of the polychaete Neanthes glandicincta. Micro Ecol 58:892–902
Lozupone C, Knight R (2005) UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235. https://doi.org/10.1128/AEM.71.12.8228-8235.2005
McFall-Ngai M, Hadfield MG, Bosch TCG, Carey HV, Domazet-Loso T, Douglas AE et al (2013) Animals in a bacterial world, a new imperative for the life sciences. Proc Natl Acad Sci 110:3229–3236. https://doi.org/10.1073/pnas.1218525110
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Moisander PH, Sexton AD, Daley MC (2015) Stable associations masked by temporal variability in the marine copepod microbiome. PLoS ONE 10:e0138967. https://doi.org/10.1371/journal.pone.0138967
Mortzfeld BM, Urbanski S, Reitzel AM, Künzel S, Technau U, Fraune S (2016) Response of bacterial colonization in Nematostella vectensis to development, environment and biogeography. Environ Microbiol 18:1764–1781. https://doi.org/10.1111/1462-2920.12926
Nakagawa S, Shimamura S, Takaki Y, Suzuki Y, Murakami SI, Watanabe T (2014) Allying with armored snails: the complete genome of gammaproteobacterial endosymbiont. ISME J 8:40–51. https://doi.org/10.1038/ismej.2013.131
Neave MJ, Streten-Joyce C, Glasby CJ, McGuinness KA, Parry DL, Gibb KS (2012) The bacterial community associated with the marine polychaete Ophelina sp. 1 (Annelida: Opheliidae) is altered by copper and zinc contamination in sediments. Micro Ecol 63:639–650. https://doi.org/10.1007/s00248-011-9966-9
Neave MJ, Apprill A, Ferrier-Pages C, Voolstra C (2016) Diversity and function of prevalent symbiotic marine bacteria in the genus Endozoicomonas. Appl Microbiol Biotechnol 100:8315–8324. https://doi.org/10.1007/s0025
Neave MJ, Michell CT, Apprill A, Voolstra CR (2017) Endozoicomonas genomes reveal functional adaptation and plasticity in bacterial strains symbiotically associated with diverse marine hosts. Sci Rep 7:40579. https://doi.org/10.1038/srep40579
Nedved BT, Hadfield M (2009) Hydroides elegans (Annelida: Polychaeta): a model for biofouling research. In: Flemming H-C, Murthy PS, Venkatesan R, Cooksey K (eds) Marine and industrial biofouling. Springer, Berlin, pp 203–217
Nussbaumer AD, Fisher CR, Bright M (2006) Horizontal endosymbiont transmission in hydrothermal vent tubeworms. Nature 441:345–348. https://doi.org/10.1038/nature04793
Oliveros J (2017) Venny. An interactive tool for comparing lists with Venn’s diagrams. http://bioinfogp.cnb.csic.es/tools/venny/index.htm
Reveillaud J, Maignien L, Eren AM, Huber JA, Apprill A, Sogin ML, Vanreusel A (2014) Host-specificity among abundant and rare taxa in the sponge microbiome. ISME J 8:1198–1209
Ruehland C, Blazejak A, Lott C, Loy A, Erséus C, Dubilier N (2008) Multiple bacterial symbionts in two species of co-occurring gutless oligochaete worms from Mediterranean sea grass sediments. Environ Microbiol 10:3404–3416. https://doi.org/10.1111/j.1462-2920.2008.01728.x
Shade A, Handelsman J (2012) Beyond the Venn diagram: the hunt for a core microbiome. Environ Microbiol 14:4–12. https://doi.org/10.1111/j.1462-2920.2011.02585.x
Shikuma NJ, Hadfield MG (2006) Temporal variation of an initial marine biofilm community and effects on larval settlement and metamorphosis of the tubeworm Hydroides elegans. Biofilms 2:231–238
Shikuma NJ, Pilhofer M, Weiss GL, Hadfield MG, Jensen GJ, Newman DK (2014) Marine tubeworm metamorphosis induced by arrays of bacterial phage tail-like structures. Science 343:529–533. https://doi.org/10.1126/science.1246794
Tran C, Hadfield MG (2011) Larvae of Pocillopora damicornis (Anthozoa) settle and metamorphose in response to surface-biofilm bacteria. Mar Ecol Prog Ser 433:85–96
Unabia CRC, Hadfield M (1999) Role of bacteria in larval settlement and metamorphosis of the polychaete Hydroides elegans. Mar Biol 133:55–64. https://doi.org/10.1007/s002270050
Walters W (2016). Improved bacterial 16S rRNA gene (V4 and V4–5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. mSystems 1:e00009–00015. https://doi.org/10.1128/msystems.00009-15
Weiland-Bräuer N, Neulinger SC, Pinnow N, Künzel S, Baines JF, Schmitz RA (2015) Composition of bacterial communities associated with Aurelia aurita changes with compartment, life stage, and population. Appl Environ Microbiol 81:6038–6052. https://doi.org/10.1128/AEM.01601-15
Zardus JD, Nedved BT, Huang Y, Tran C, Hadfield MG (2008) Microbial biofilms facilitate adhesion in biofouling invertebrates. Biol Bull 214:91–98. https://doi.org/10.2307/25066663
Acknowledgements
We are pleased to acknowledge lab assistants Kenneth Choi and Krystel Sarvis for assisting in culturing larvae. Dr. Marnie Freckelton provided very useful commentary and assistance on earlier drafts of this manuscript. The authors are grateful to the reviewers for their constructive comments and recommendations. This is publication number 10576 from the School of Ocean, and Earth Science and Technology and the Kewalo Marine Laboratory at the University of Hawai’i at Mānoa.
Funding
Research reported here was supported by funds from grants to MGH from the Oak Foundation (no. ORIO-13-013) and to MGH and BN from the Gordon and Betty Moore Foundation (no. 5009), and U.S. Office of Naval Research Grant no. N00014-15-1-2658.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interests.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Responsible Editor: O. Vadstein.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Reviewed by S. M. Degnan and an undisclosed expert.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Vijayan, N., Lema, K.A., Nedved, B.T. et al. Microbiomes of the polychaete Hydroides elegans (Polychaeta: Serpulidae) across its life-history stages. Mar Biol 166, 19 (2019). https://doi.org/10.1007/s00227-019-3465-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-019-3465-9